- The paper introduces NNCME-2, a neural framework that uses autoregressive density estimators, natural gradient, and TDVP to solve the chemical master equation.
- It demonstrates a 5–22x speedup over previous methods while accurately capturing both dominant statistics and rare-event dynamics in complex reaction networks.
- The approach scales to high-dimensional and spatially extended systems, enabling efficient exploration of rare transitions and metastable states in stochastic kinetics.
Tracking Large Chemical Reaction Networks and Rare Events with Neural Networks
Introduction: Overcoming the Limitations of Traditional CME Solvers
Solving the chemical master equation (CME) for complex reaction networks is central to modeling stochastic chemical kinetics, systems biology, and spatially embedded processes. The CME’s exponential state space makes direct numerical integration infeasible beyond low system sizes, particularly when dealing with spatially extended networks or rare-event statistics. Traditional methods such as finite state projection, buffered-space truncation, and tensor-network contractions offer partial relief but break down as the number of species, states, or network dimensionality increases. Moreover, conventional simulation techniques (e.g., SSA/Gillespie) become computationally prohibitive for accessing rare transitions and metastable events, as their scaling with system size is typically exponential.
This paper proposes a substantial advancement in the neural-network-based approach to CME solutions, introducing the NNCME-2 framework. The methodology leverages advanced neural autoregressive architectures, efficient second-order optimizers (notably natural gradient and TDVP), and enhanced-sampling techniques, allowing both tractable scaling to large/high-dimensional systems and accurate estimation of rare-event probabilities and transitions.
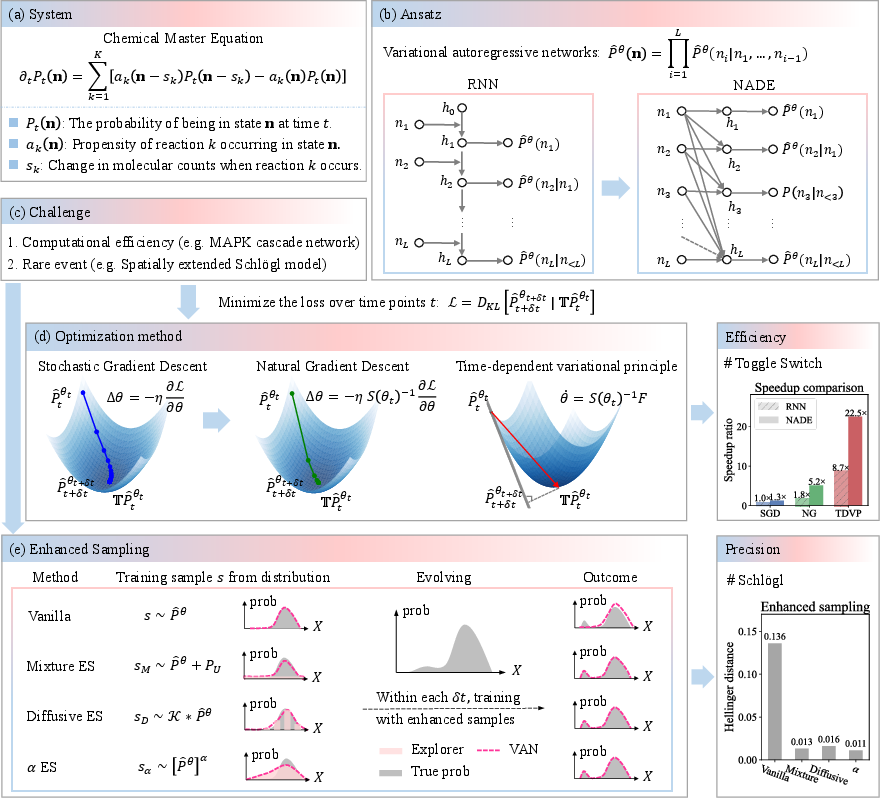
Figure 1: Schematic overview of NNCME-2 combining VAN representations, efficient optimization, and enhanced-sampling for both scalable CME tracking and rare event capture.
Neural Autoregressive Architectures for Stochastic Dynamics
NNCME-2 parameterizes the CME probability distribution via neural-autoregressive density estimators (NADE). The VAN factorizes the joint probability into conditionals, significantly improving sampling and normalization over traditional sequential RNNs while retaining the flexibility to model arbitrary dependency structures. The main practical implementation is the NADE, due to its favorable trade-off between parameter efficiency and expressivity. The authors additionally demonstrate that transformer-based VANs, while more computationally demanding, can further extend the expressive range relevant for complex or highly multimodal distributions.
The feedforward NADE approach allows efficient, scalable training and sampling over high-dimensional species configurations. Importantly, this architecture can be adopted with very minor modification for spatially structured and complex/topologically rich networks, as required for realistic reaction-diffusion modeling or dense biochemical networks.
Second-Order Optimization: Natural Gradient Descent and TDVP
A major limitation of previous neural CME approaches has been slow convergence, suboptimal accuracy, and high computational cost when using standard SGD, particularly for stiff/highly multimodal distributions. NNCME-2 systematically replaces SGD with second-order methods:
- Natural Gradient (NG) rescales updates using the Fisher information metric, yielding parameter updates along the steepest descent in distribution space, respecting underlying statistical geometry.
- Time-Dependent Variational Principle (TDVP) projects the evolution onto the tangent space of the variational manifold, yielding continuous-time parameter updates with typically just one iteration per time step needed for accurate tracking.
The stochastic low-rank approximation to the FIM (discussed in detail and benchmarked empirically) allows efficient NG application even for large parameter sets by exploiting matrix identities and sample-based traces.
Benchmarking on the genetic toggle switch demonstrates that NNCME-2 achieves a 5–22x speedup versus previous RNN-based/SKD approaches, with nearly ideal scaling and no loss of accuracy (Fig. 2). These improvements are critical for tractability on larger reaction networks, where standard approaches become infeasible.
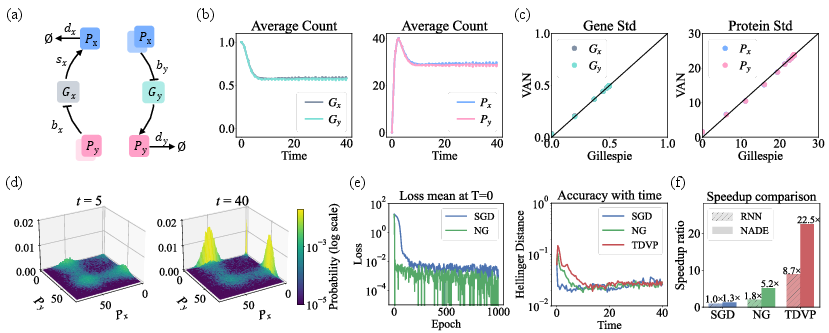
Figure 2: Genetic toggle switch results showing 5x+ acceleration versus NNCME-1, matching the accuracy of Gillespie and recovering multimodal/joint statistics.
Enhanced Sampling for Rare Event Characterization
Rare events in metastable and spatial systems determine macroscopic kinetics, phase transitions, and stochastic activation phenomena. Direct training of VANs (or any sampler) from high-probability regions generally fails to populate rare-event states, resulting in poor or biased predictions of transition rates and steady-state weights. NNCME-2 incorporates several enhanced-sampling procedures:
- Mixture ES: Augments training batches with uniformly random samples, ensuring low-density regions are reached and mode collapse is mitigated.
- Diffusive ES: Convolves the proposal distribution with a smoothing kernel, broadening exploration and facilitating saddle-crossing in the configuration landscape.
- Alpha ES: Samples from an over-dispersed power-reweighted (α<1) version of the current probability, effectively “annealing” and flattening sharp dominant basins.
These strategies are rigorously benchmarked on the Schlögl bistability model and in large spatially extended 1D and 2D lattices.
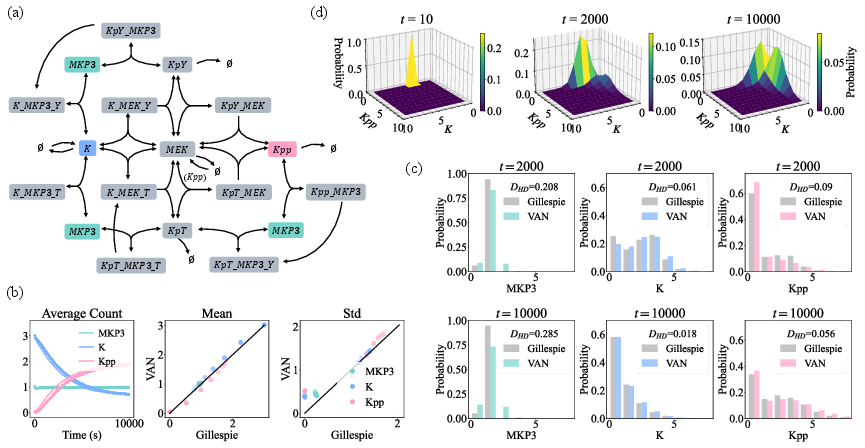
Figure 3: MAPK cascade, a 16-species, 35-reaction network, tackled by NNCME-2 with high efficiency and close agreement to Gillespie.
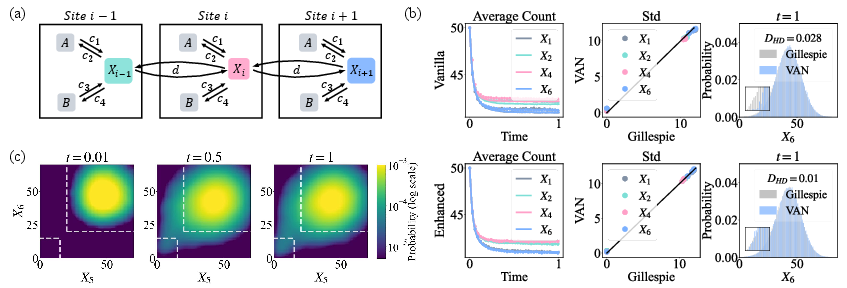
Figure 4: Comparison of rare-event sampling in a 6-site Schlögl system with and without enhanced-sampling; only enhanced-sampling captures low-probability peaks and transitions.
High-Dimensional and Spatially Extended Reaction Networks
NNCME-2 is demonstrated on:
- The MAPK cascade (16 species, 35 reactions): Previously intractable for previous neural CME methods, NNCME-2 attains accurate statistics, matching Gillespie simulations with tractable cost.
- Spatially extended Schlögl lattices: Both 1D (up to 8 sites) and 2D (2×4) lattices were handled, exceeding the limits of tensor-network approaches (MPS/PEPS) in two dimensions. Rare-event probability scaling and transition rate estimation remain accurate, with computational cost scaling far more favorably than classical simulation even as the probability mass of critical configurations becomes exponentially suppressed.
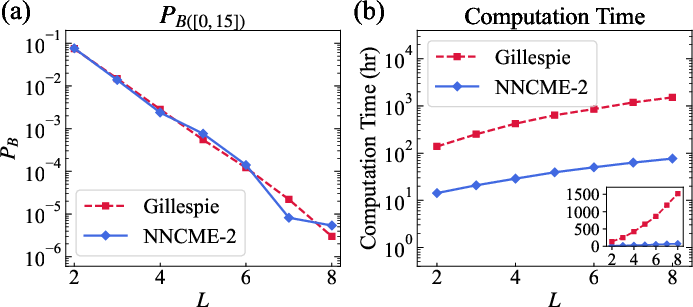
Figure 5: Scaling of rare probability mass in the 1D Schlögl model and corresponding computational cost; NNCME-2 outpaces Gillespie when growing system size and rare-event suppression.
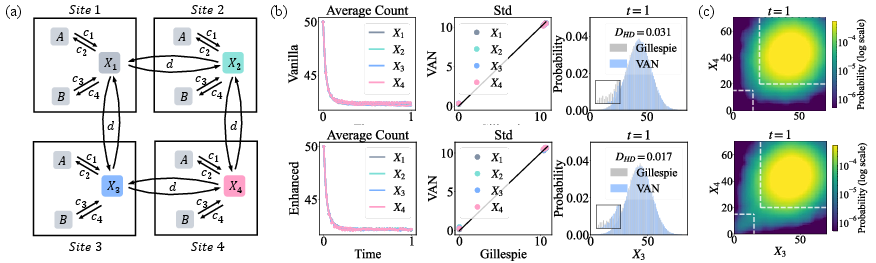
Figure 6: 2D lattice Schlögl results: NNCME-2 tracks mean/variance and captures rare-event structure (marginal, joint distributions) intractable to conventional simulators.
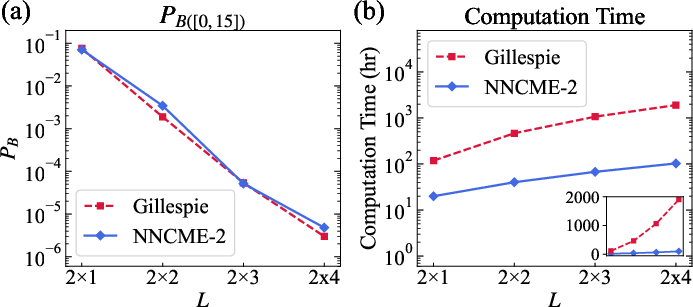
Figure 7: Cost scaling and rare-event probability accuracy for 2D spatially extended models—NNCME-2 displays favorable system size dependence.
Theoretical Claims and Numerical Validation
- Consistent recovery of rare-event dynamics in both 1D and 2D reaction-diffusion lattices, outperforming unaugmented VAN/traditional neural solutions.
- Robust stability of NADE-based architectures with natural gradient/TDVP optimization even in late-time, highly multimodal regimes.
- Superior or competitive empirical cost scaling versus SSA and tensor-network approaches on large/high-dimensional systems, even with substantially suppressed rare-event mass.
Implications and Future Directions
NNCME-2 establishes a robust, scalable, and general approach for learning time-evolving distributions in stochastic kinetics. It is not system-design dependent and works for arbitrary reaction topologies, unlike buffer-based or tensor contraction methods. The approach is extensible to:
- More complex geometries, e.g., disordered, nonlocal or network-based reaction systems
- Nonequilibrium phase transitions and spatially inhomogeneous phenomena (e.g., front propagation, nucleation)
- Integration with tensor-network sampling to further bridge rare-event coverage and model expressivity
Notably, the methodology enables systematic exploration of configuration-space regions that are crucial for functional predictions of gene regulatory networks, enzyme cascades, cellular signaling, or phase transitions—areas where rare events are mechanistic drivers.
The combination of advanced autoregressive architectures, information-geometric optimization, and adaptive sampling provides a blueprint for future methods in stochastic simulation, nonequilibrium statistical mechanics, and potentially for operator learning in related high-dimensional dynamical systems.
Conclusion
NNCME-2 represents a highly efficient and scalable framework for solving the CME in large, high-dimensional, or rare-event-dominated regimes. By leveraging advanced neural parameterizations, geometry-aware optimization, and principled enhanced-sampling, it achieves drastic reductions in computational cost, accurate recovery of both dominant and rare probability regions across diverse systems, and enables investigation of reaction networks previously inaccessible to direct simulation. The approach is generalizable to a wide range of future applications in chemical, biological, and soft matter modeling, and opens avenues for further integration of machine learning in nonequilibrium statistical dynamics.
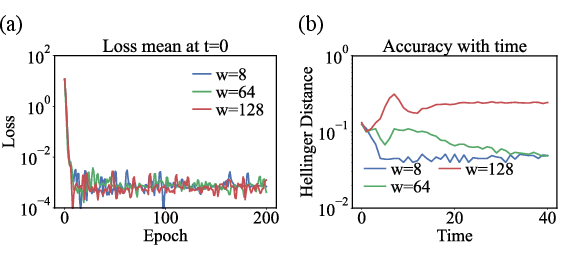
Figure 8: Effect of network width on model capacity, convergence, and downstream accuracy—critical for balancing runtime and representation power.
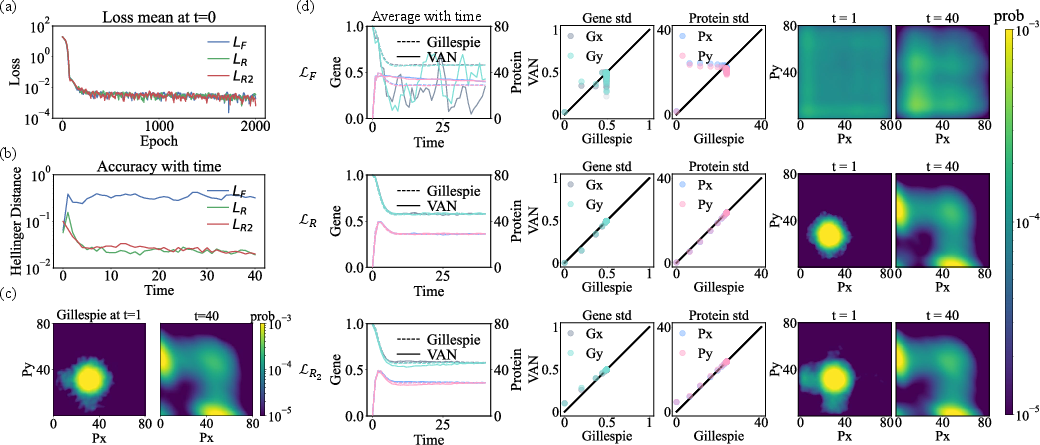
Figure 9: Evaluation of KL-divergence objectives for learning CME dynamics, illustrating Hellinger distances and joint statistics fidelity across variants.
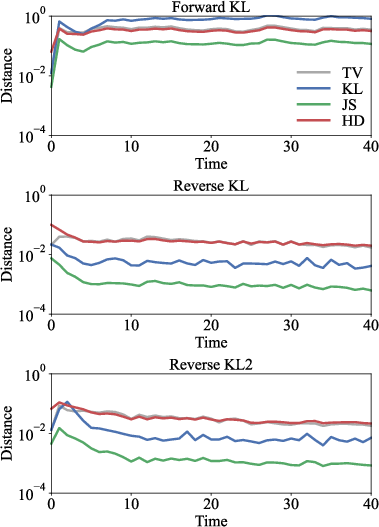
Figure 10: Hellinger distance proves to be a representative and robust evaluation metric for distribution matching across training schemes.









