- The paper presents a novel hybrid CNN-GNN architecture for node-wise uncertainty quantification in CXR landmark segmentation.
- It rigorously validates uncertainty metrics through occlusion, noise, and OOD experiments, achieving ROC AUC up to 0.98.
- CheXmask-U offers a public dataset with 657,566 annotated CXR segmentations, enabling refined clinical decision support.
Uncertainty Quantification in Landmark-Based Anatomical Segmentation for Chest X-rays
Introduction
Quantifying uncertainty in anatomical segmentation has become essential for trustworthy deployment of deep learning-based systems in medical imaging, particularly under clinically relevant conditions such as occlusion, corruption, and out-of-distribution (OOD) samples. Unlike dense pixel-based approaches, landmark-based segmentation provides topo-anatomical guarantees, but until now, fine-grained uncertainty estimation at the node (landmark) level has remained largely understudied. This paper presents a systematic framework for node-wise uncertainty estimation within landmark-based segmentation, focusing on chest X-ray (CXR) analysis using a variational hybrid CNN-GNN architecture.
HybridGNet Architecture for Landmark Segmentation
HybridGNet integrates a CNN-based encoder with a GCNN-based decoder, operating in a probabilistic VAE latent space. The primary innovation lies in using this latent space to provide two orthogonal uncertainty metrics: (i) model (epistemic) uncertainty via the learned latent variance, and (ii) predictive uncertainty via variance over Monte Carlo-sampled landmark predictions.
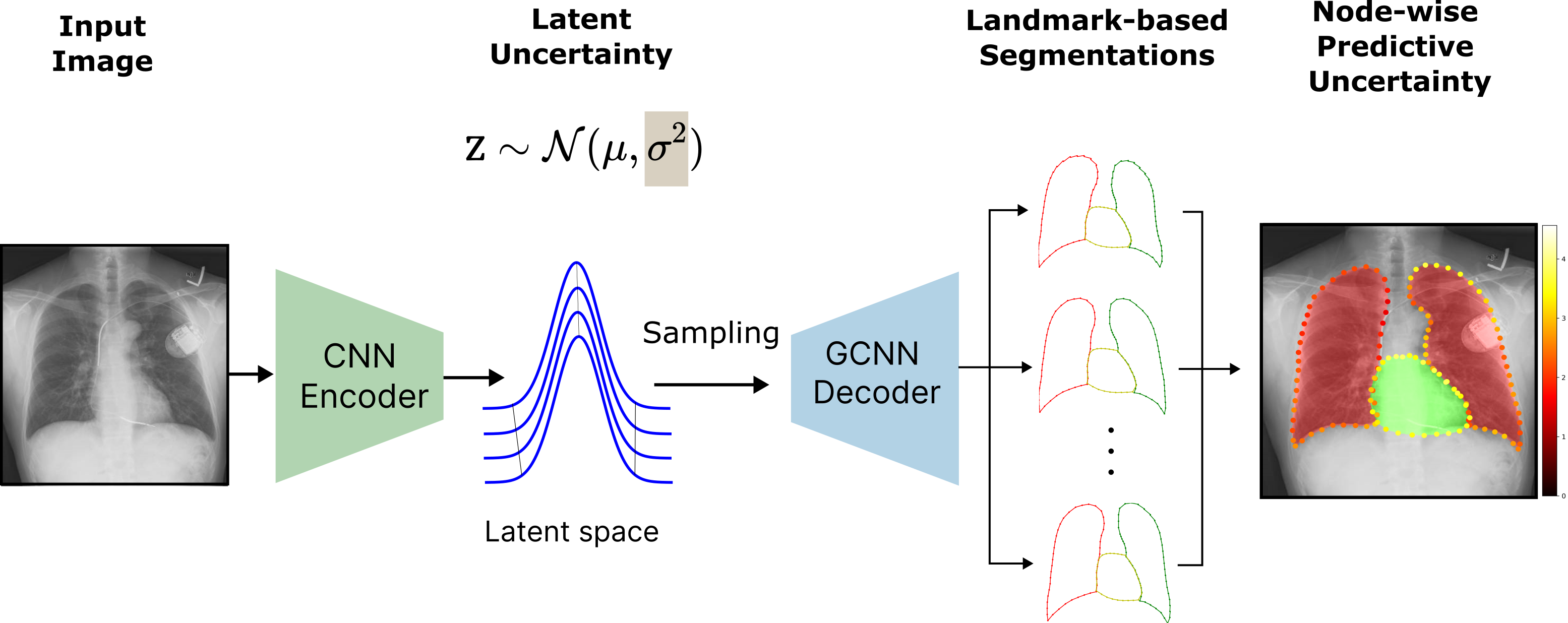
Figure 1: Block diagram of HybridGNet showing the extraction of a latent uncertainty σ2, followed by sampling-based node-wise predictive uncertainty estimation.
Landmarks are graph-represented anatomical points with a shared adjacency structure, ensuring anatomical feasibility and facilitating structured predictive modeling. The network supports skip connections from the encoder to the decoder, which were analyzed for their impact on uncertainty interpretability, especially under corruption.
Uncertainty Estimation Methodology
Latent-space uncertainty is quantified as the mean latent variance vector σ2—reflecting epistemic model uncertainty not explained by the training data. Predictive uncertainty is estimated by batch-wise decoding of N latent samples through the GCNN, yielding landmark sets whose per-node sample variance serves as the uncertainty metric.
This node-wise uncertainty is highly informative, spatially resolving model confidence and allowing differential trust in various anatomical regions.
Experimental Validation
Extensive corruption studies investigated the robustness and interpretability of both uncertainty metrics:
- Occlusion Experiments: Artificial black-square masks were used to create localized perturbations. Per-node uncertainty systematically increased under occluded regions, confirming strong spatial specificity.
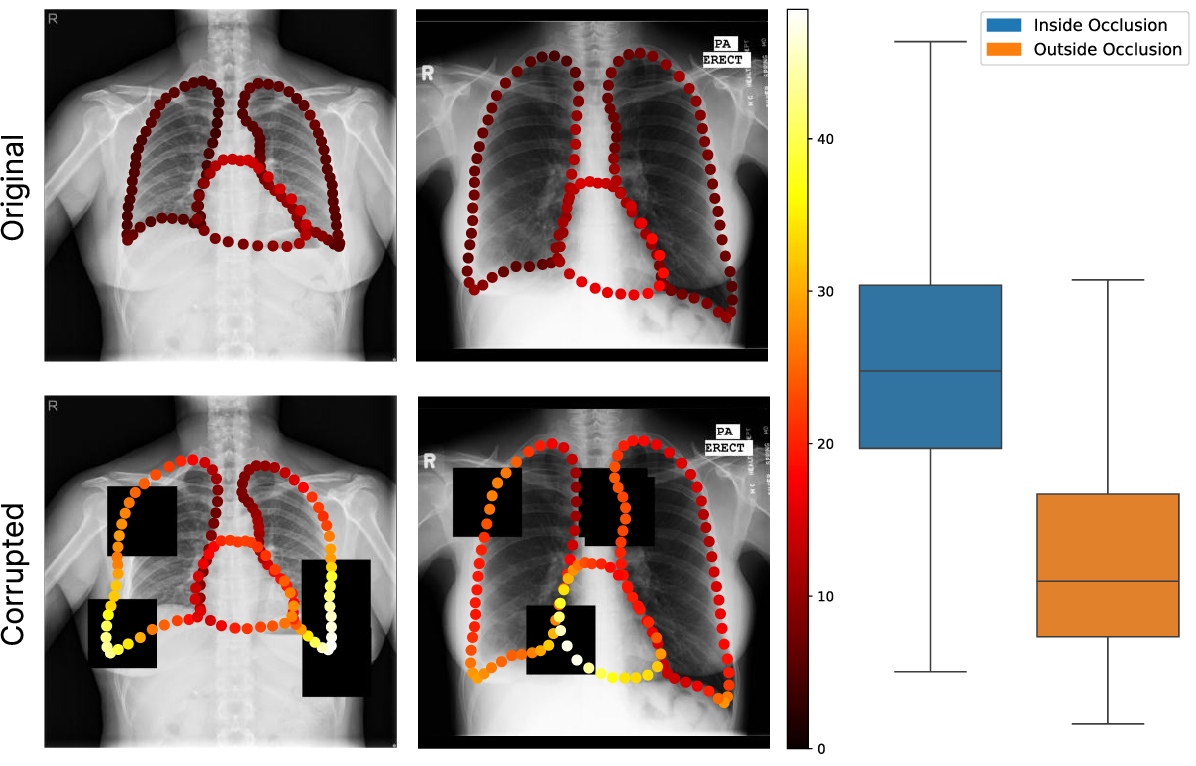
Figure 2: Visualization of uncertainty maps under occlusion, with clear uncertainty elevation in masked zones; boxplots confirm statistical significance.
- Noise Corruption: Gaussian noise of increasing magnitude was applied to the images. Both metrics increased monotonically with corruption—except for non-monotonicity in the skip-connection model's predictive uncertainty at severe corruptions, attributed to direct high-resolution feature flow bypassing the variational bottleneck.
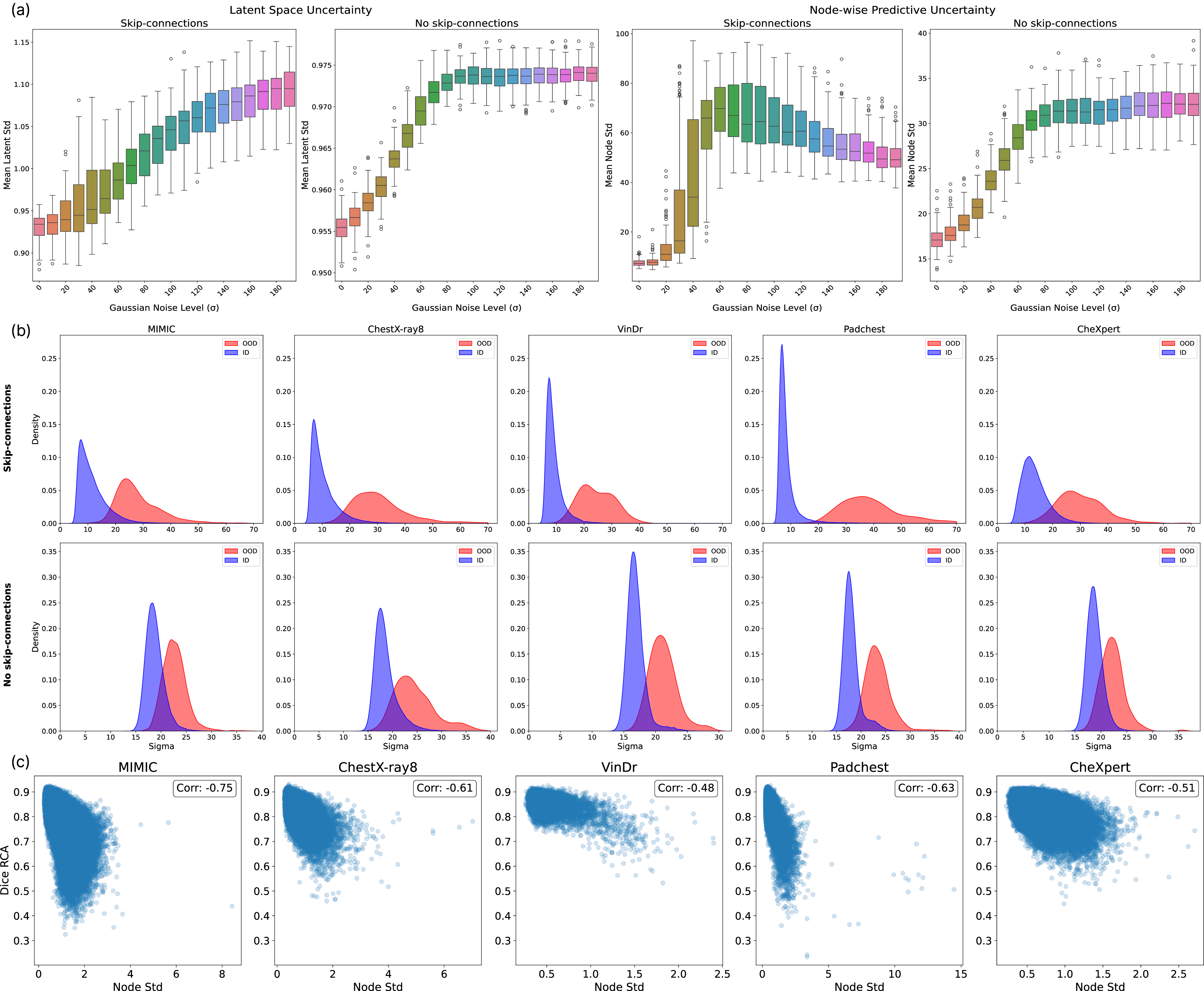
Figure 3: (a) Both uncertainty measures rise with Gaussian noise, plateauing at high levels; (b) KDEs for OOD detection with strong ID/OOD separation; (c) Strong anti-correlation between average uncertainty and RCA-estimated Dice, evidencing uncertainty's reliability as a quality surrogate.
- OOD Detection: On CheXMask, predictive uncertainty delivered excellent ID/OOD discrimination, with ROC AUC up to 0.98 using skip-connections. Latent-space-based anomaly scores (Isolation Forest) also performed well.
- Error Prediction: Cross-image and cross-node correlations between node-wise uncertainty and error (relative to multi-expert annotation) were strong, evidencing the practical predictive value of per-node UQ.
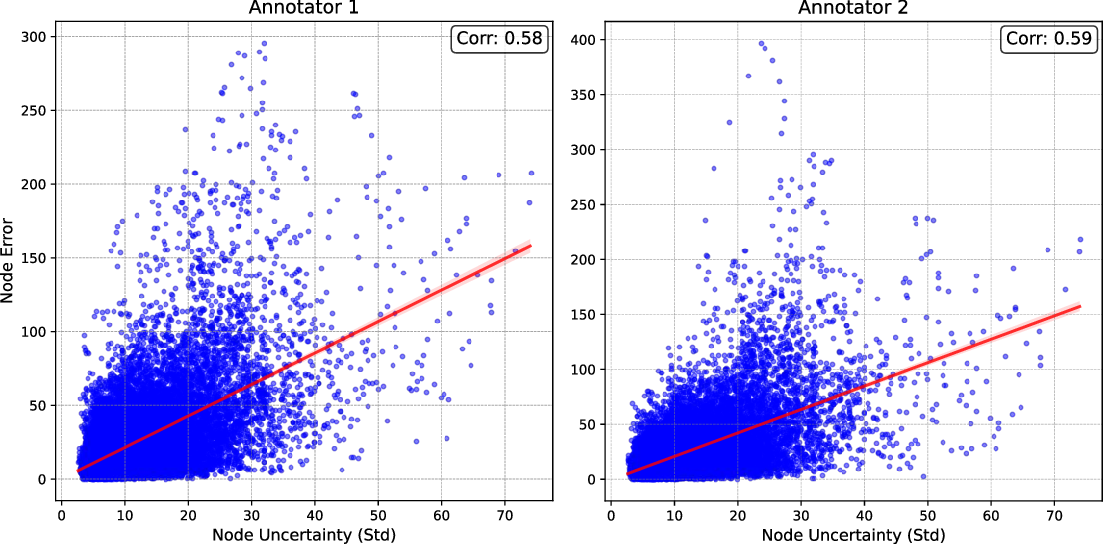
Figure 4: High correlation between predictive uncertainty and node-level annotation errors, confirming practical validity of uncertainty estimates.
CheXmask-U: Dataset Release
CheXmask-U provides 657,566 chest X-ray segmentations, each annotated with per-node uncertainty for direct community use. Uncertainty metrics were computed via 50 Monte Carlo landmark samples per image, using pre-trained HybridGNet weights. Beyond simply supplementing the anatomical coordinate sets, this resource enables slicing, filtering, or region-focused confidence weighting in downstream CXR tasks, without requiring consumers to re-run computationally intensive UQ techniques.
Node-level validation, involving comparison with expert multiple annotations, demonstrates that CheXmask-U uncertainty estimates reliably reflect region-specific reliability, surpassing the original image-level RCA-Dice score in granularity.
Implications and Future Directions
This node-level UQ framework has direct implications for image triage, automated reporting, and selective use of segmentation outputs in context-aware clinical decision support. The ability to spatially map uncertainty supports safe automation, targeted human review, and can guide adaptive acquisition or annotation strategies.
In future work, adaptations to multi-organ, multi-modality, or 3D contexts are clear next steps, as well as integration with more sophisticated epistemic/aleatoric disentanglement methods and application-specific thresholds for automatic acceptance/rejection or weighting schemes.
Conclusion
This paper proposes and validates the first comprehensive node-level uncertainty quantification scheme for landmark-based anatomical segmentation in CXR, underpinned by systematic experiments across occlusion, noise, OOD, and error correlation axes. CheXmask-U constitutes a public, fine-grained resource for advancing anatomically aware, uncertainty-calibrated segmentation in medical imaging.





