Graph AI generates neurological hypotheses validated in molecular, organoid, and clinical systems
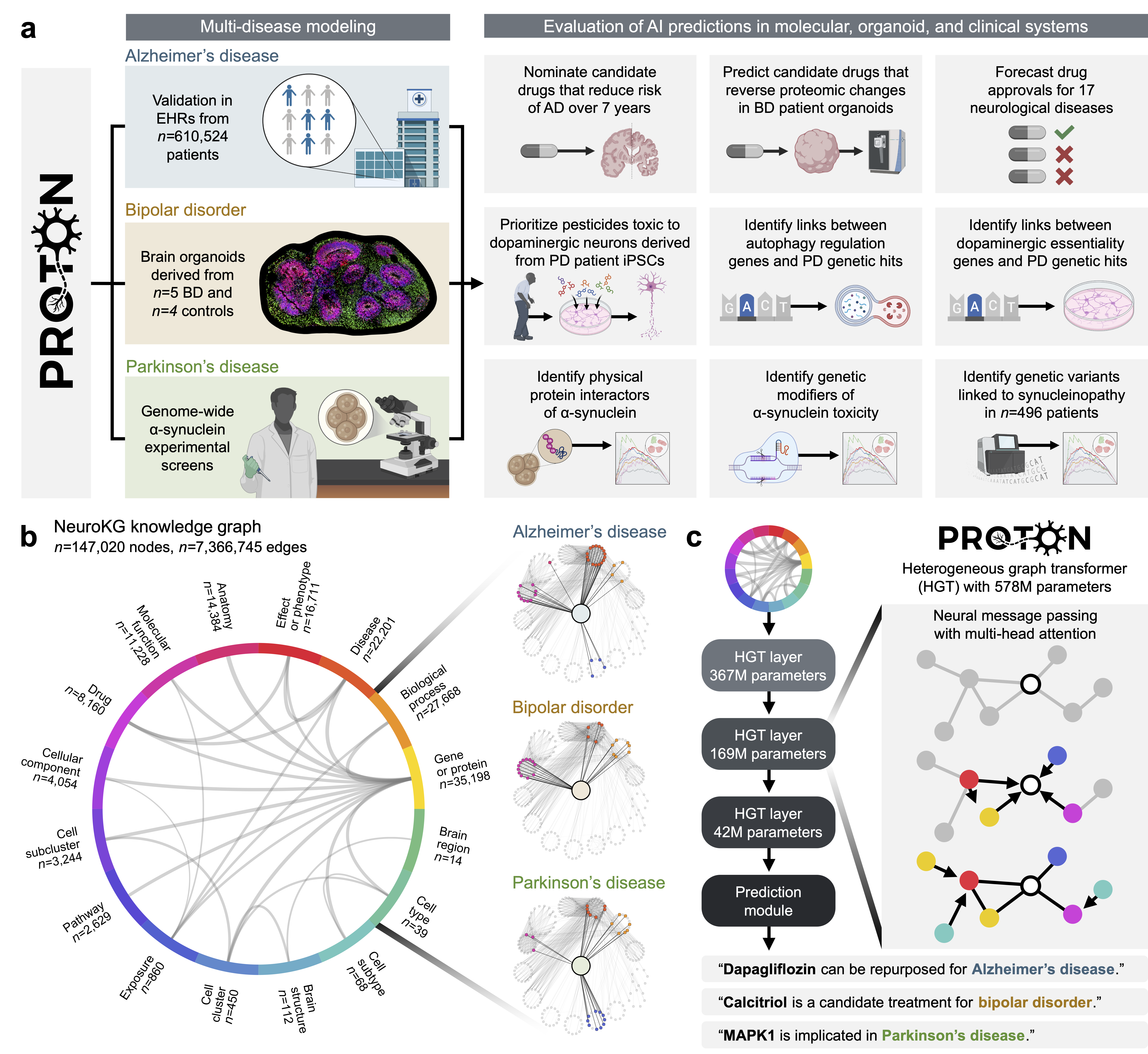
Abstract: Neurological diseases are the leading global cause of disability, yet most lack disease-modifying treatments. We present PROTON, a heterogeneous graph transformer that generates testable hypotheses across molecular, organoid, and clinical systems. To evaluate PROTON, we apply it to Parkinson's disease (PD), bipolar disorder (BD), and Alzheimer's disease (AD). In PD, PROTON linked genetic risk loci to genes essential for dopaminergic neuron survival and predicted pesticides toxic to patient-derived neurons, including the insecticide endosulfan, which ranked within the top 1.29% of predictions. In silico screens performed by PROTON reproduced six genome-wide $α$-synuclein experiments, including a split-ubiquitin yeast two-hybrid system (normalized enrichment score [NES] = 2.30, FDR-adjusted $p < 1 \times 10{-4}$), an ascorbate peroxidase proximity labeling assay (NES = 2.16, FDR $< 1 \times 10{-4}$), and a high-depth targeted exome sequencing study in 496 synucleinopathy patients (NES = 2.13, FDR $< 1 \times 10{-4}$). In BD, PROTON predicted calcitriol as a candidate drug that reversed proteomic alterations observed in cortical organoids derived from BD patients. In AD, we evaluated PROTON predictions in health records from $n = 610,524$ patients at Mass General Brigham, confirming that five PROTON-predicted drugs were associated with reduced seven-year dementia risk (minimum hazard ratio = 0.63, 95% CI: 0.53-0.75, $p < 1 \times 10{-7}$). PROTON generated neurological hypotheses that were evaluated across molecular, organoid, and clinical systems, defining a path for AI-driven discovery in neurological disease.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
Overview
This paper introduces Proton, a powerful AI model designed to help scientists discover new ideas (“hypotheses”) about brain diseases. The team built a huge “map” of biology related to the human brain—connecting genes, proteins, cell types, brain regions, environmental exposures, symptoms, and drugs—and trained Proton to find useful patterns across this map. They then tested Proton’s predictions in three real situations: Parkinson’s disease (PD), bipolar disorder (BD), and Alzheimer’s disease (AD).
What questions did the researchers ask?
To make the work easy to follow, here are the main questions the team wanted to answer:
- Can an AI model connect the dots between many kinds of brain data (genes, cells, drugs, and patient health records) to suggest useful ideas for treatment or risk?
- For Parkinson’s disease, can the AI point to important genes and harmful environmental chemicals (like pesticides) that affect the neurons lost in PD?
- For bipolar disorder, can the AI find drugs that might fix the molecular “protein patterns” seen in patient-derived mini-brain models?
- For Alzheimer’s disease, can the AI predict existing drugs (used for other conditions) that are linked to lower dementia risk in real-world patients?
How did they do the research?
Building an AI “brain map”
- The team created NeuroKG, a big knowledge graph (think of it like a city map) where:
- “Cities” are biological items (genes, proteins, brain cell types, drugs, diseases).
- “Roads” are the connections between them (e.g., which gene affects which protein, which drug targets which gene).
- Proton is a graph transformer model. Instead of reading sentences, it “reads” this map and learns how different parts relate. It’s trained to guess missing links (like predicting a road that should connect two cities).
- The model learns “embeddings,” which are like GPS coordinates for each item so that related items end up near each other.
Testing in Parkinson’s disease
- PD mainly harms dopamine-producing neurons (cells important for movement).
- The team compared Proton’s predictions to multiple large lab experiments focused on a protein called alpha‑synuclein (which builds up in PD).
- They also checked whether Proton can link genetic risk regions for PD (from GWAS—big studies of many people’s DNA) to genes that are essential for dopamine neuron survival.
- Finally, they fine-tuned Proton on a small dataset about pesticides and tested if it could spot which chemicals are both linked to PD and toxic to patient-derived neurons in dishes.
Testing in bipolar disorder with “mini-brains”
- They used human brain organoids—tiny, lab-grown 3D brain-like structures made from patient cells. These don’t capture everything about BD but keep patient-specific biology.
- Proton ranked drugs that might help BD. The team chose calcitriol (the active form of vitamin D) among the top suggestions and treated BD organoids.
- They measured proteomics—basically, an inventory of proteins to see how treatment changes the molecular patterns.
Testing in Alzheimer’s disease with health records
- Proton predicted drugs (approved for other diseases) that might be linked to lower dementia risk.
- The team checked this using real-world electronic health records (EHR) from 610,524 patients over several years.
- They used survival analysis (a statistical method) to compare the chance of developing dementia in patients who took the candidate drugs versus those who didn’t, while balancing groups by age and sex.
- A “hazard ratio” less than 1 means the drug group had a lower risk during the follow-up period.
What did they find?
Parkinson’s disease (PD)
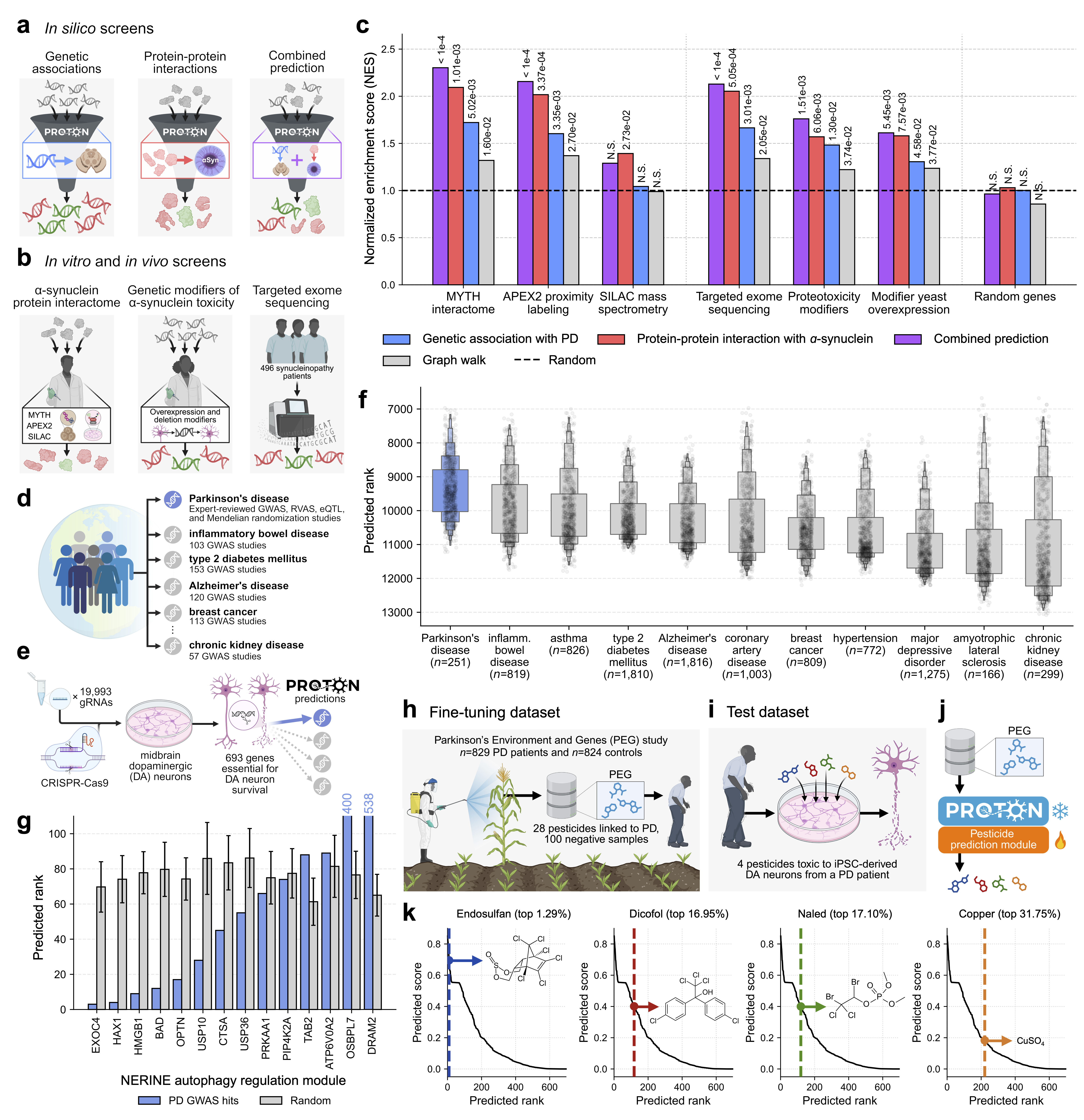
- Proton’s predictions matched the results of several large experiments studying alpha‑synuclein. This means Proton can act like a “virtual screen,” pointing to genes and proteins worth testing.
- It linked PD genetic risk regions to genes necessary for dopamine neuron survival more strongly than for other diseases.
- After fine-tuning, Proton correctly ranked pesticides that were toxic to patient-derived dopamine neurons. Notably, the insecticide endosulfan was ranked in the top 1.29% of all predictions.
Why this is important: It shows Proton can connect genetics, proteins, and environmental risks to the vulnerable brain cells affected in PD.
Bipolar disorder (BD)
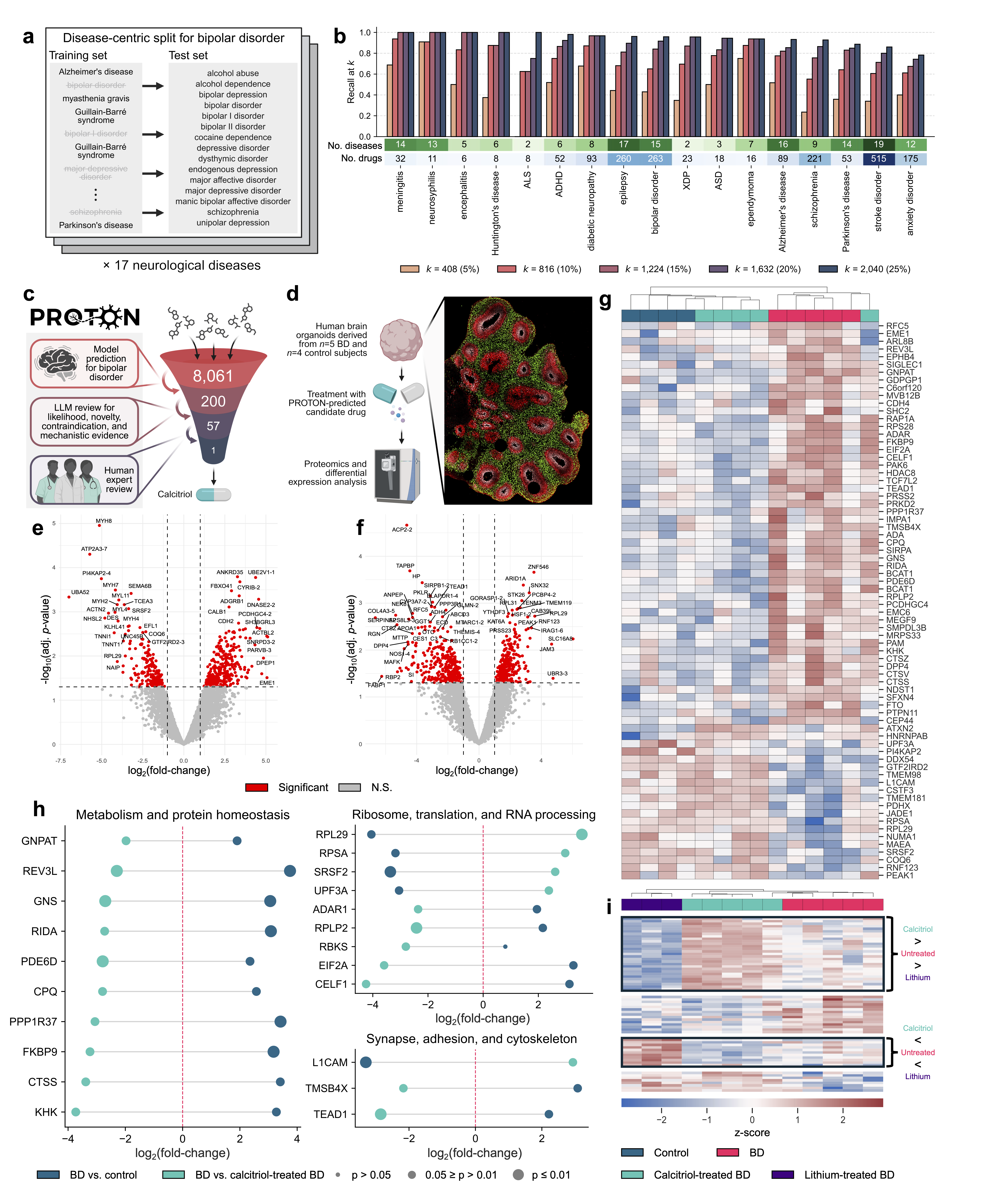
- Proton suggested calcitriol (active vitamin D) as a candidate drug.
- When BD organoids were treated with calcitriol, the protein patterns shifted closer to healthy controls, suggesting partial “normalization.”
- The changes involved processes like RNA editing (how cells fine-tune messages), metabolism, and synaptic function (how brain cells connect and communicate).
- Calcitriol’s effects were partly different from lithium, a common BD treatment, hinting at a complementary mechanism.
Why this is important: Proton helped pick a promising drug that improved disease-related molecular changes in a human-relevant system.
Alzheimer’s disease (AD) and related dementias
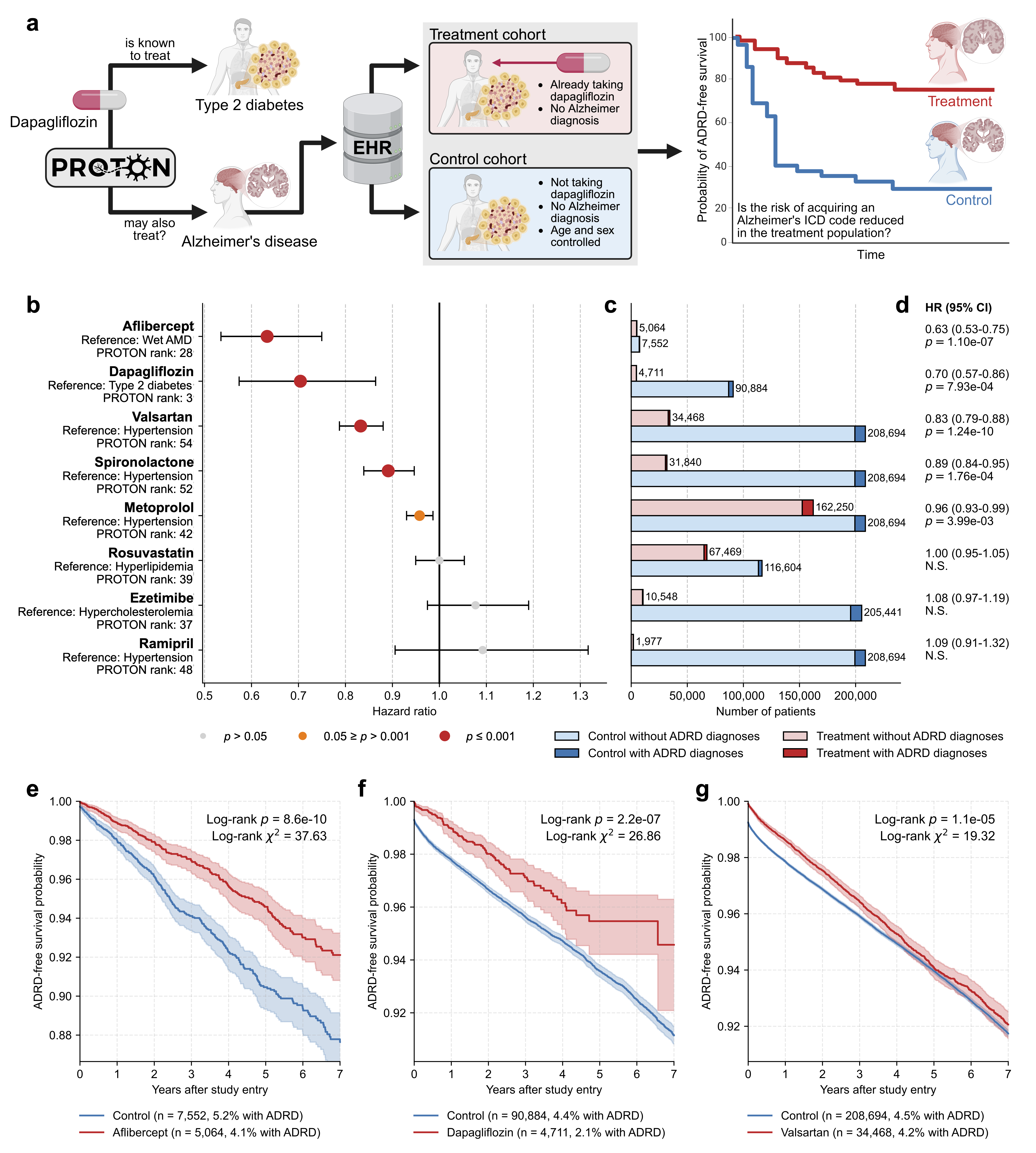
- In analyses of 610,524 patients’ health records, 5 out of 8 Proton-predicted drugs were linked to a lower seven-year dementia risk.
- Examples include:
- Aflibercept (used for eye disease): about 37% lower risk.
- Dapagliflozin (a diabetes drug): about 30% lower risk.
- Valsartan, spironolactone, and metoprolol (blood pressure/heart drugs) also showed protective associations.
- Not all drugs showed benefit, and these findings are associations, not proof of prevention.
Why this is important: Proton’s ideas matched real-world patterns, pointing to medicines worth further clinical testing.
What does this mean for the future?
This work suggests a practical way to use AI in brain research:
- Build comprehensive maps that connect molecular biology, cells, drugs, and patient data.
- Use AI to propose testable ideas.
- Check those ideas in lab experiments (like organoids) and in real-world patient data.
Potential impact:
- Faster discovery of drug repurposing opportunities for brain diseases.
- Better understanding of how genes, environment (like pesticides), and cell biology come together in disease.
- A blueprint for “AI–experiment loops,” where AI proposes, experiments test, and the results feed back to improve the AI.
Important note:
- These results are promising, but they’re not the final word. Some predictions need larger studies and clinical trials. AI models can reflect biases in the data they learn from, and not all diseases have enough data yet. Still, Proton shows how AI can make useful, testable suggestions across multiple scales—from molecules to patients.
Knowledge Gaps
Below is a concise, actionable list of the paper’s unresolved knowledge gaps, limitations, and open questions that future work could address.
- Coverage gaps in NeuroKG: limited representation of rare and understudied neurological diseases, developmental stages, and disease progression dynamics; unclear impact of missing modalities (e.g., proteo-genomics, longitudinal cell-state trajectories, metabolomics).
- Knowledge graph bias: uneven literature attention inflates connectivity for well-studied genes/diseases; residual degree bias in transformer attention may favor hubs despite upweighting of rare edge types.
- Validation of KG edges: 16% of edges in disease neighborhoods lack PaperQA2 support; reliance on an LLM-based validator risks literature hallucinations or selective citation; need for independent, human-curated audits and negative evidence incorporation.
- Information leakage risk in repurposing evaluation: disease-centric splits remove drug–disease edges but may still leak signals via phenotypes, clinical trials, side effects, or related nodes not fully ablated; formal leakage audits and stricter relation-type/temporal splits are needed.
- Temporal generalization not tested: no time-split (pre-/post-year) evaluation to assess genuine forecasting of future approvals or clinical evidence.
- Limited baselines: beyond RWR for PD, no comparisons to strong knowledge-graph and link-prediction baselines (e.g., TransE/RotatE/ComplEx, GraIL, R-GCN, HGT variants, DRKG-based models) or to non-graph repurposing methods.
- Ablation studies missing: no systematic quantification of contribution from each node/edge type, snRNA-seq contextualization, or relation class weighting; unclear which components drive performance.
- Interpretability: lack of path-level rationales/attribution (e.g., subgraph explanations, attention rollout) hinders mechanistic trust and experimental prioritization.
- Uncertainty quantification absent: no calibrated confidence/credibility intervals for rankings; uncertainty estimates needed to prioritize higher-confidence, novel hypotheses.
- Out-of-distribution generalization: no tests on unseen nodes/entities (new drugs, newly discovered genes) or OOD disease spaces; inductive capacity unproven.
- Scalability and maintenance: unclear strategies for continual learning, KG updating, versioning, and preventing catastrophic forgetting as new neurological data arrive.
- PD in silico screens: significant enrichments but few experimentally validated novel targets; need prospective wet-lab validation (e.g., CRISPR perturbations) of top-ranked, previously untested genes/proteins in DA neurons.
- PD GWAS-to-mechanism mapping: predictions linking DA-essential genes to PD loci lack causal validation (eQTL colocalization, CRISPR perturbation, perturb-seq) and tissue/cell-type specificity tests across vulnerable and resilient neuronal/glial subtypes.
- PD pesticide predictions: fine-tuning used only 28 positives and 100 negatives; held-out test of n=4 pesticides is underpowered; requires broader external validation, dose–response assays, structure-activity controls, and new in vitro/in vivo toxicity tests.
- Environmental exposure modeling: no accounting for mixtures, exposure timing, metabolism, gene–environment interactions, or real-world exposure levels; mechanistic links to PD pathobiology remain speculative.
- BD organoid experiments: small cohort (n=5 BD, n=4 control), short treatment (1 week), and cortical-only model; lacks replication, cross-line/batch controls, and evaluation in additional brain regions/cell types (e.g., inhibitory neurons, astrocytes, microglia).
- BD functional phenotyping: proteomic normalization not linked to electrophysiology, synaptic function, network activity, or cellular stress assays; unclear if proteomic shifts translate to functional rescue.
- BD calcitriol mechanism: no tests for VDR dependency (antagonists/knockdown), downstream signaling, or causal role of RNA editing (ADAR activity, A-to-I editing readouts); mechanism-of-action remains inferential.
- BD comparators: used lithium orotate (non-standard clinical formulation) instead of lithium carbonate; no dose–response or combination/synergy studies with standard-of-care medications.
- AD EHR analysis — confounding: IPTW/Cox adjusted only for age and sex; no adjustment for race/ethnicity, socioeconomic status, comorbidities (e.g., vascular/metabolic), baseline cognition, polypharmacy, healthcare utilization, or disease severity.
- AD EHR analysis — design biases: potential immortal time bias (different index dates for treated vs controls), misclassification from ICD-based ADRD case definitions, untested proportional hazards assumption, lack of competing-risk (death) modeling, and no falsification endpoints.
- AD EHR analysis — exposure definition: no dose, duration, adherence, time-varying exposure/covariates, or lag analyses; unclear robustness to medication switching and concomitant therapy.
- AD EHR analysis — selection and multiplicity: only 8 drugs evaluated from one health system; selection based on rank and prevalence may introduce bias; multiple testing adjustments and external replication cohorts are needed.
- Generalizability: findings from a single healthcare system (MGB) and specific organoid models may not transfer to broader, diverse populations or other laboratories; external, multi-institutional replication is lacking.
- Safety and translation: calcitriol dosing and safety (e.g., hypercalcemia risk) not addressed; no preclinical efficacy/safety studies or clinical trial planning for BD or AD candidates.
- Model fairness and equity: limited assessment of performance and benefits across demographics (sex, ancestry, socioeconomic status); need bias audits and subgroup analyses.
- Negative evidence integration: KG does not explicitly encode failed trials or negative mechanistic findings; incorporating negatives could reduce false positives and improve causal plausibility.
- Prospective validation loop: no prospective, preregistered predictions with blinded validation across molecular, organoid, and clinical systems; establishing an AI–experiment–clinic discovery loop remains an open challenge.
Practical Applications
Immediate Applications
Below are actionable use cases that can be deployed now, based on the paper’s validated findings, released code/models, and demonstrated workflows.
- AI-assisted drug repurposing triage in neurology
- Sector: Healthcare, pharma/biotech, software
- What: Use Proton’s in silico screening to rank approved drugs and narrow candidates for Alzheimer’s disease and related dementias (ADRD), bipolar disorder (BD), Parkinson’s disease (PD), and other neurological indications.
- Tools/workflows: Proton model weights (HuggingFace), code (GitHub), NeuroKG dataset; run indication ranking per disease; shortlist top-k for downstream validation (organoids, EHR, literature).
- Assumptions/dependencies: Predictions reflect associations, not causality; clinical benefit must be tested prospectively; performance depends on NeuroKG coverage and biases.
- Real-world evidence (RWE) screening for neuroprotective associations
- Sector: Healthcare systems, payers, pharmacoepidemiology
- What: Reproduce the ADRD pipeline: pair Proton-ranked candidates with EHR-based retrospective survival analyses to screen for protective associations before committing to trials.
- Tools/workflows: Disease-specific ICD-10 cohorts, inverse probability treatment weighting, doubly robust Cox models; Proton for candidate down-selection.
- Assumptions/dependencies: Confounding by indication remains; requires IRB/ethics approvals, fit-for-purpose EHR data quality, and statistical expertise.
- Organoid-guided validation of AI-nominated candidates
- Sector: Biotech, academic labs, CROs
- What: Use cortical or dopaminergic neuron organoids to test Proton-selected compounds for reversal of disease proteomic programs (e.g., calcitriol in BD organoids).
- Tools/workflows: Patient-derived iPSC organoids, deep proteomics (Orbitrap), differential expression and pathway analysis; compare with standard-of-care (e.g., lithium) for orthogonal mechanism assessment.
- Assumptions/dependencies: Organoids are disease-relevant but not complete disease models; dosing/PK/PD may not translate directly to patients.
- Target and mechanism discovery from multi-scale genetics
- Sector: Pharma/biotech R&D, academia
- What: Use Proton to connect GWAS signals to cell-type-essential genes and protein interactors (e.g., PD GWAS ↔ dopaminergic neuron essential genes; α-synuclein interactome).
- Tools/workflows: In silico PPI screens, gene set enrichment analysis (GSEA), module-level rare variant corroboration (e.g., NERINE).
- Assumptions/dependencies: Dependent on interactome completeness and single-cell annotations; potential degree bias in graph learning.
- Environmental toxicant prioritization for neurotoxicity testing
- Sector: Public health, environmental safety, regulatory science
- What: Few-shot fine-tune Proton on PWAS hits to prioritize pesticides for in vitro neurotoxicity testing (e.g., endosulfan ranked top 1.29% and shown toxic to patient-derived neurons).
- Tools/workflows: Curate PWAS positives/negatives; fine-tune Proton; test top-ranked compounds in dopaminergic neuron models.
- Assumptions/dependencies: Requires small labeled sets; translational relevance depends on exposure levels, mixtures, and chronicity; some compounds (e.g., endosulfan) are already banned but remain as legacy contaminants.
- Best-practice ML evaluation using disease-centric data splits
- Sector: Academia, software/ML tooling, industry AI teams
- What: Adopt the paper’s disease-centric data split methodology to prevent information leakage when evaluating drug-disease link prediction on KGs.
- Tools/workflows: Remove all disease-related drug links pre-training; evaluate recall@k on held-out indications and off-label uses; release of split scripts.
- Assumptions/dependencies: Requires rigorous ontology mapping of “related diseases”; still subject to indirect leakage via correlated edges.
- Knowledge-graph maintenance with AI-augmented literature curation
- Sector: Academia, publishers, data providers, software
- What: Use PaperQA2 or similar agents to validate KG edges and prioritize curation gaps (83.87% neighborhood support achieved for major diseases).
- Tools/workflows: Periodic KG refresh, agent-assisted evidence retrieval, human-in-the-loop curation dashboards.
- Assumptions/dependencies: LLM retrieval quality and literature coverage; ongoing curator oversight is essential.
- Portfolio prioritization for neuroscience pipelines
- Sector: Pharma/biotech strategy, venture investment
- What: Integrate Proton rankings with target feasibility, IP, and competitive intelligence to prioritize assets for in-licensing or internal development.
- Tools/workflows: Multi-criteria decision analyses combining Proton scores with chemistry, safety, and commercial filters.
- Assumptions/dependencies: Downstream success still depends on target druggability and clinical tractability.
- Clinician–scientist hypothesis generation
- Sector: Academic medicine, translational research
- What: Query Proton for cell-type-specific disease mechanisms (e.g., DA neuron vulnerability in PD) to guide design of CRISPR screens, PPI assays, or biomarker panels.
- Tools/workflows: In silico screen → prioritized experiments → wet-lab validation loops.
- Assumptions/dependencies: Requires local assay capacity; iterative cycles improve value.
- Public and occupational health risk assessment triage
- Sector: Policy/public health, NGOs
- What: Use Proton-based shortlists to triage compounds or exposure contexts for deeper epidemiological or toxicological investigation in communities with elevated PD/AD risks.
- Tools/workflows: PWAS-style designs, exposure registries, environmental biomonitoring; align with existing surveillance programs.
- Assumptions/dependencies: Data availability on exposures; ethical and regulatory frameworks for community studies.
Long-Term Applications
These applications are promising but need further research, scaling, or regulatory/operational development before broad deployment.
- AI-driven adaptive platform trials in neurology
- Sector: Healthcare, pharma/biotech, regulators
- What: Use Proton to seed multi-arm, adaptive trials for ADRD, BD, PD; update arms as EHR/organoid evidence accumulates.
- Tools/workflows: Bayesian adaptive designs, RWE-informed prior elicitation, platform governance.
- Assumptions/dependencies: Regulatory acceptance of AI-guided arm selection; robust safety monitoring; interoperable data infrastructure.
- Personalized therapeutic matching via patient-derived organoids + Proton
- Sector: Precision medicine, biotech, reference labs
- What: Combine Proton shortlists with patient-specific organoid response profiles to select individualized treatments.
- Tools/workflows: Standardized iPSC derivation, automated organoid phenotyping, multi-omic readouts, decision-support integration.
- Assumptions/dependencies: Turnaround time, cost, and standardization hurdles; payer coverage and clinical utility evidence required.
- Mechanism-of-action deconvolution and target discovery at scale
- Sector: Pharma/biotech discovery
- What: Use Proton to bridge rare/common variants and PPIs across cell types to identify convergent pathways and new targets (e.g., autophagy module in PD).
- Tools/workflows: Network module mining, causal inference over KGs, CRISPR perturb-seq for validation.
- Assumptions/dependencies: KG completeness, bias mitigation, and causal validation technologies.
- Regulatory science for chemical safety and post-market surveillance
- Sector: Policy, regulatory agencies (EPA, EMA, FDA)
- What: Incorporate Proton-like models into chemical review workflows to prioritize testing, re-evaluations, and monitoring of legacy pollutants with suspected neurotoxicity.
- Tools/workflows: Model qualification, transparent audit trails, standardized reporting.
- Assumptions/dependencies: Policy frameworks for AI evidence acceptance; multidisciplinary review panels.
- Clinical decision support (CDS) for neuroprotective prescribing
- Sector: Healthcare IT, EHR vendors
- What: In the future, suggest medications with favorable dementia-risk profiles among clinically equivalent options (e.g., within antihypertensive class) using Proton-informed evidence.
- Tools/workflows: CDS hooks, patient-level risk adjustment, shared decision-making UIs.
- Assumptions/dependencies: Requires prospective validation and regulatory clearance; avoid indication bias; equitable performance across populations.
- Foundation models for brain disease integrating multi-modal, longitudinal data
- Sector: Software/AI, academia
- What: Scale Proton to incorporate spatial transcriptomics, proteomics, imaging, wearables, and longitudinal health data for richer reasoning about brain diseases.
- Tools/workflows: Continual KG updating, multi-modal pretraining, federated learning with privacy safeguards.
- Assumptions/dependencies: Data sharing agreements, harmonization standards, compute resources.
- KG-informed safety pharmacology and off-target risk prediction
- Sector: Pharma/biotech safety
- What: Predict neurological adverse events and off-targets of candidate drugs by traversing multi-relational paths in NeuroKG.
- Tools/workflows: Integrated safety dashboards, alerting for high-risk mechanisms, in vitro neurotoxicity verification.
- Assumptions/dependencies: Requires curated adverse event ontologies and post-marketing data integration.
- Public health policy modeling for dementia prevention
- Sector: Policy, health economics
- What: Use Proton-prioritized drug classes and risk modifiers to simulate population-level impacts of prescription patterns on incident dementia.
- Tools/workflows: Microsimulation, cost-effectiveness analyses, scenario planning for formularies.
- Assumptions/dependencies: Strong causal evidence is needed; heterogeneous treatment effects must be characterized.
- Education and workforce development in graph AI for biomedicine
- Sector: Academia, professional training
- What: Build curricula, workshops, and hackathons around Proton/NeuroKG to train interdisciplinary teams.
- Tools/workflows: Open datasets and model weights; best-practice evaluation (disease-centric splits).
- Assumptions/dependencies: Sustained funding and community governance; periodic dataset/model refreshes.
- Strategic R&D portfolio analytics
- Sector: Finance, life-science investing
- What: Use Proton signals to prioritize investment into neurological programs aligned with genetically anchored or convergent network mechanisms.
- Tools/workflows: Overlay Proton scores with market, IP, and trial risk analytics.
- Assumptions/dependencies: Model outputs complement, not replace, diligence; transparency into uncertainty and bias is essential.
Notes on feasibility and responsible use
- Model/data dependencies: Performance depends on NeuroKG completeness, quality, and bias; well-studied nodes may be overrepresented. Periodic, transparent KG updates and bias audits are critical.
- Causality: EHR-based associations are not causal; RCTs or quasi-experimental designs are needed before clinical adoption.
- Model generalization: Disease-centric data splits reduce leakage but cannot eliminate all indirect signals; cross-system validation (molecular → organoid → clinical) is advised.
- Safety and ethics: Do not initiate or alter medical treatment based solely on model predictions; clinical decisions require licensed clinicians and evidence-based guidelines.
- Regulatory pathways: Adoption in regulatory or CDS contexts requires rigorous validation, documentation, and compliance with privacy and safety standards.
Glossary
- ADARs: Enzymes that edit RNA by converting adenosine (A) to inosine (I), influencing RNA stability and translation. "ADARs (Adenosine Deaminase Acting on RNA)"
- APEX2: An engineered ascorbate peroxidase used for proximity labeling to identify proteins near a bait protein in living cells. "an ascorbate peroxidase proximity labeling assay (APEX2)"
- AUROC: Area Under the Receiver Operating Characteristic curve; a performance metric for binary classifiers measuring tradeoff between true- and false-positive rates. "achieved high link prediction performance (AUROC ; accuracy )"
- Autophagy: Cellular degradation and recycling process that removes damaged organelles and proteins via lysosomes. "enriched in autophagy regulation"
- Bayesian hyperparameter optimization: A probabilistic strategy (often using surrogate models like Gaussian processes) to efficiently search hyperparameter space. "Through Bayesian hyperparameter optimization"
- Cox proportional hazards model: A semiparametric survival model that relates covariates to event hazard rates, assuming proportional hazards over time. "We used Cox proportional hazards models to estimate the association between drug exposure and incident dementia,"
- Cortical organoid: A 3D in vitro model derived from human stem cells that recapitulates aspects of cortical brain development and cell types. "cortical organoids derived from patients"
- CRISPR/Cas9 screen: A genome-wide perturbation approach using CRISPR/Cas9 to knock out or modify genes to discover those affecting a phenotype. "an unbiased whole-genome CRISPR screen"
- Dopaminergic (DA) neuron: A neuron that synthesizes and releases dopamine, critical in motor control and affected in Parkinson’s disease. "loss of dopaminergic (DA) neurons in the substantia nigra pars compacta"
- Doubly robust Cox regression model: A survival analysis approach combining outcome modeling and weighting so estimates remain consistent if either model is correctly specified. "doubly robust Cox regression model."
- Epitranscriptomic mechanisms: Chemical modifications of RNA (e.g., methylation, editing) that regulate RNA function without changing the underlying sequence. "association between BD and RNA alterations by epitranscriptomic mechanisms"
- False discovery rate (FDR): The expected proportion of false positives among declared significant findings when performing multiple hypothesis tests. "FDR "
- Gene set enrichment analysis (GSEA): A statistical method to test whether predefined sets of genes show statistically significant, concordant differences between conditions. "via gene set enrichment analysis (GSEA)"
- Genome-wide association study (GWAS): A study scanning the genome for common variants associated with a trait or disease across many individuals. "genome-wide association studies (GWAS) implicate hundreds of loci"
- Hazard ratio (HR): A measure in survival analysis comparing the event rate between two groups over time. "(hazard ratio [HR] , 95\% CI , -value ;"
- Heterogeneous graph transformer: A transformer architecture adapted to graphs with multiple node and edge types, enabling relation-aware message passing. "a heterogeneous graph transformer that generates testable hypotheses"
- ICD-10: The 10th revision of the International Classification of Diseases, a standardized diagnostic coding system. "specific ICD-10 diagnosis codes"
- In silico: Performed via computer simulation or computational analysis rather than in wet-lab experiments or living organisms. "in silico screens performed by Proton"
- Induced pluripotent stem cell (iPSC): A somatic cell reprogrammed to a pluripotent state, capable of differentiating into various cell types. "iPSC-derived midbrain DA neurons"
- Inverse probability treatment weighting (IPTW): A weighting technique using propensity scores to create a pseudo-population that balances covariates between treated and control groups. "we used inverse probability treatment weighting (IPTW) to balance treatment and control groups"
- Knowledge graph: A structured representation of entities and their relationships in a graph form, supporting reasoning and inference. "a knowledge graph contextualized to the adult human brain."
- Kolmogorov–Smirnov statistic: A nonparametric statistic comparing cumulative distributions; used in GSEA to assess enrichment. "a Kolmogorov-Smirnov statistic adjusted for multiple hypothesis testing and gene set size."
- Link prediction: The task of inferring missing or future edges between nodes in a graph based on observed structure and attributes. "using a self-supervised link prediction objective"
- Mendelian randomization: A causal inference method using genetic variants as instrumental variables to estimate the effect of an exposure on an outcome. "Mendelian randomization experiments"
- MYTH: Membrane Yeast Two-Hybrid, a split-ubiquitin system for detecting protein-protein interactions in membranes. "a split-ubiquitin yeast two-hybrid system (MYTH)"
- NeuroKG: A multimodal, brain-centered knowledge graph integrating genes, proteins, cell types, regions, drugs, and phenotypes. "NeuroKG, a knowledge graph contextualized to the adult human brain."
- Normalized enrichment score (NES): A GSEA-derived metric scaling enrichment scores to account for gene set size and multiple testing. "normalized enrichment score (NES)"
- Off-label use: The prescription of an approved drug for an indication, dose, or population not specified in its official labeling. "prescribed off-label"
- Orbitrap Astral Mass Spectrometer: A high-resolution mass spectrometry instrument platform enabling deep, accurate proteomic profiling. "using an Orbitrap Astral Mass Spectrometer"
- PaperQA2: A generative AI literature-search agent that retrieves and synthesizes evidence from scientific papers. "PaperQA2, a generative literature search agent"
- Protein-protein interaction (PPI): Physical or functional associations between proteins that influence cellular pathways and complexes. "protein-protein (PPI) with -synuclein"
- Random walk with restart (RWR): A network algorithm that repeatedly walks from a source node with a chance to return, ranking nodes by proximity. "a random walk with restart (RWR) algorithm"
- Rare-variant association study: A genetic study focusing on low-frequency variants to detect associations with diseases or traits. "rare-variant association studies"
- Relational transformer: A transformer architecture designed to model typed relations among entities, enabling reasoning over structured graphs. "a 578-million-parameter relational transformer"
- Self-attention: A mechanism that computes weighted interactions among elements of an input to capture contextual dependencies. "use self-attention and next token prediction objectives"
- Self-supervised learning: Learning from unlabeled data via proxy objectives (e.g., predicting masked or missing information). "self-supervised link prediction objective"
- Silhouette score: A clustering quality metric comparing cohesion (within-cluster) and separation (between-cluster) for each point. "neurodegenerative disease silhouette score "
- Single-nucleus RNA-sequencing (snRNA-seq): Sequencing RNA from isolated nuclei to profile gene expression at single-cell resolution, useful for frozen tissues. "single-nucleus RNA-sequencing (snRNA-seq)"
- SGLT-2 inhibitor: A class of drugs that lower blood glucose by inhibiting renal sodium–glucose cotransporter 2. "The SGLT-2 inhibitor dapagliflozin"
- Survival analyses: Statistical methods for time-to-event data assessing how covariates affect the time until an event occurs. "reduced seven-year dementia risk in survival analyses"
- Vitamin D receptor (VDR): A nuclear receptor that mediates gene regulatory effects of the active form of vitamin D. "via the nuclear vitamin D receptor (VDR)"
Collections
Sign up for free to add this paper to one or more collections.