- The paper introduces CAPRMIL, which shifts context modeling to the patch embedding stage, reducing reliance on heavy MIL aggregators.
- It achieves linear complexity, cutting trainable parameters by up to 92.8% and inference FLOPs by up to 99% while maintaining competitive accuracy.
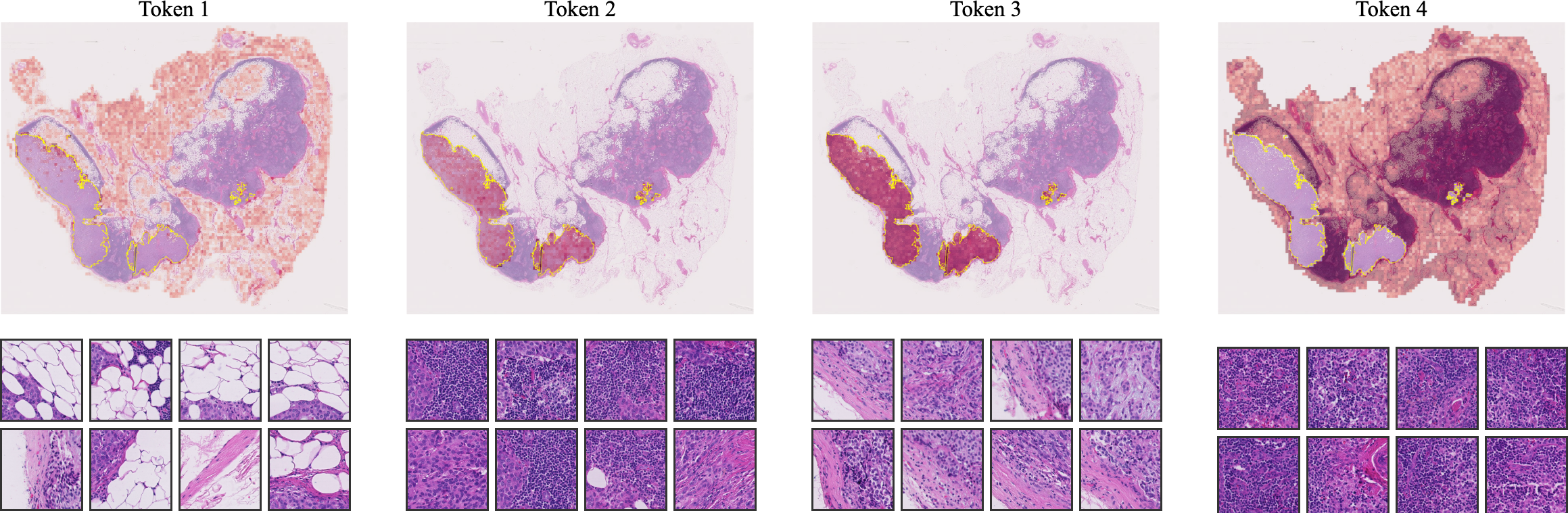
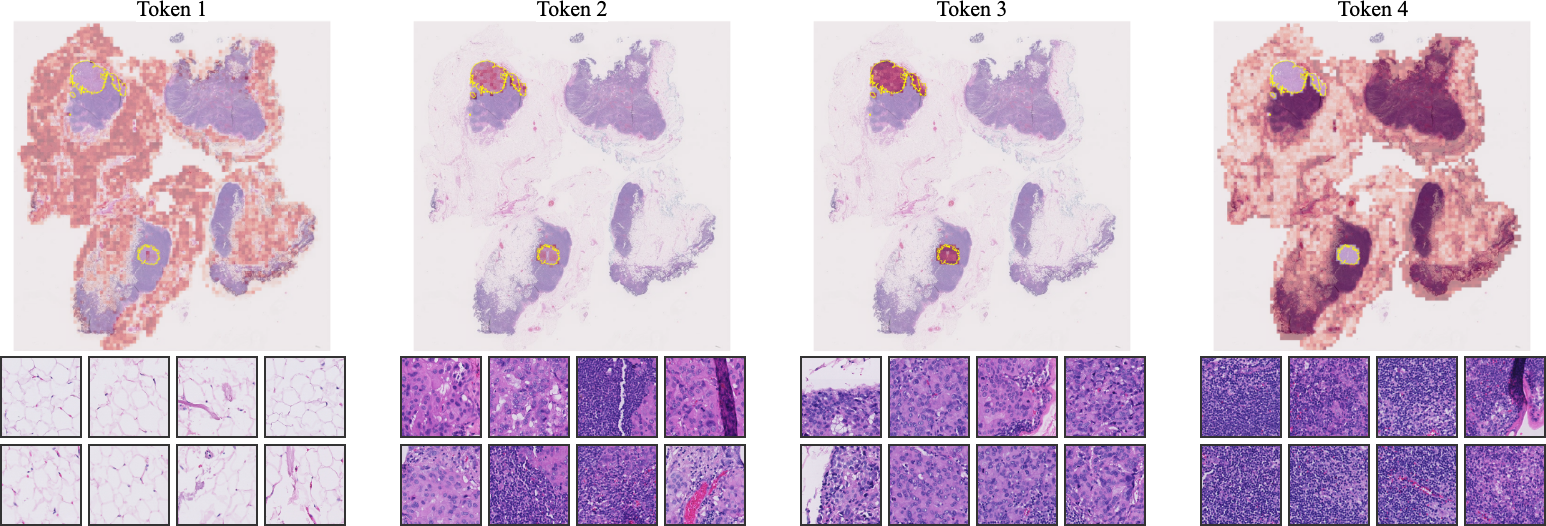
- Visualizations demonstrate that CAPRMIL clusters reflect morphologically coherent regions, enhancing interpretability for clinical applications.
CAPRMIL: Context-Aware Patch Representations for Efficient and Scalable Multiple Instance Learning
Introduction and Motivation
The digitization of pathology has necessitated robust, scalable frameworks for the analysis of gigapixel-scale Whole Slide Images (WSIs), where only slide-level supervisory signals are available. Standard embedding-based and attention-based MIL architectures, such as ABMIL, CLAM, TransMIL, and recent probabilistic and prototype-based MILs, have demonstrated success but are constrained by quadratic complexity of attention, susceptibility to overfitting, and heavy dependence on sophisticated aggregators. This work introduces CAPRMIL, a context-aware MIL paradigm that fundamentally shifts the locus of correlation modeling from the MIL aggregator to the patch embedding stage, drawing inspiration from operator learning advances in neural PDE solvers.
CAPRMIL Framework
CAPRMIL decouples the challenges of context modeling and bag-level aggregation, ensuring that each patch embedding is enriched with global, morphology-aware context before entering a lightweight MIL aggregation module. The pipeline is as follows:
- Patch Extraction and Projection: WSIs are tessellated and encoded into feature embeddings by a frozen backbone (e.g., a large-scale foundation model), then linearly projected into a compact latent space for computational efficiency.
- Context-Aware Patch Encoding: A stack of CAPRMIL Blocks augments each patch with global context using multi-head self-attention not over the native sequence of patches, but over a compact set of context/morphology tokens derived via soft clustering of patch representations. This mechanism is mathematically and algorithmically analogous to efficient transformer solvers for PDEs such as Transolver and Transolver++.
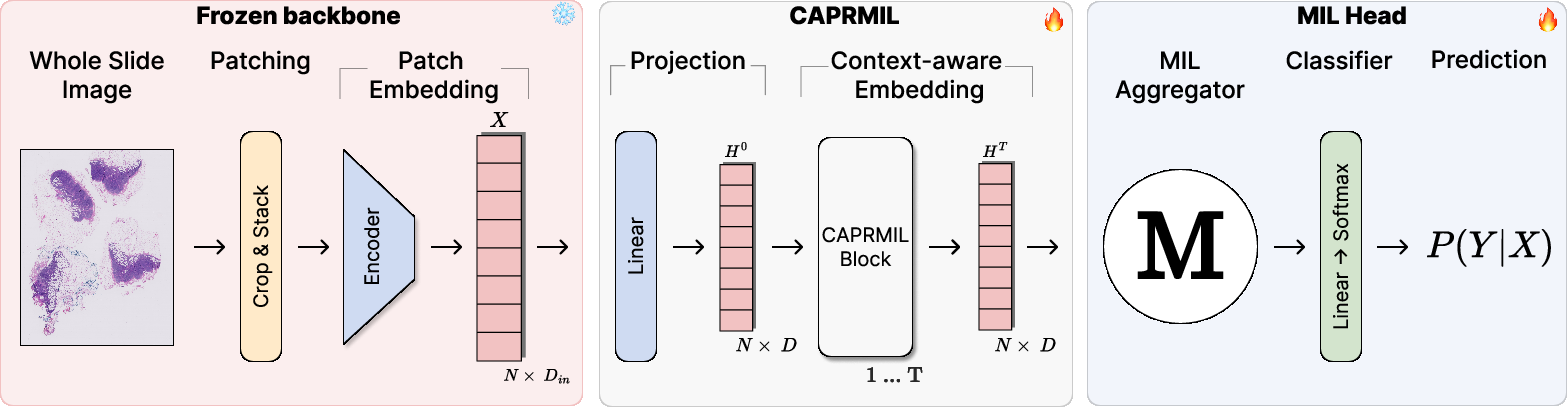
Figure 1: The CAPRMIL framework, from patch sampling to slide-level prediction via context-aware patch encoding and pooling.
Within each CAPRMIL Block, soft clustering maps patches to M≪N clusters, which are aggregated into tokens; self-attention is performed in this low-dimensional space, and global context is broadcast back to patches using the soft assignment matrix.
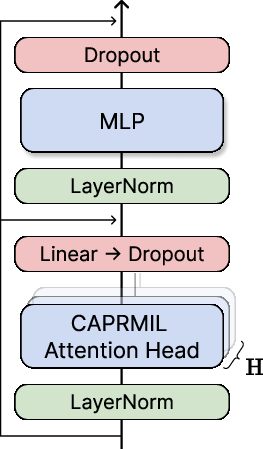
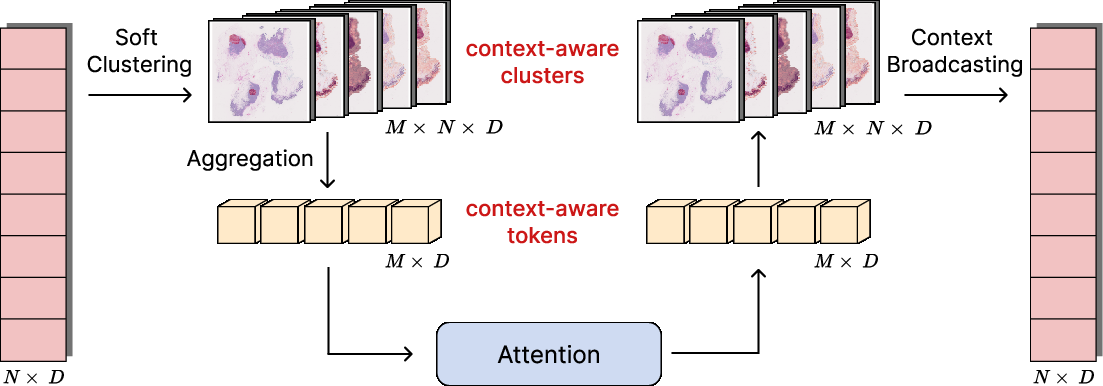
Figure 2: (a) Transformer-based CAPRMIL Block design. (b) Per-head clustering, token aggregation, attention, and context broadcast.
- MIL Aggregation and Classification: The context-enriched patch embeddings are pooled (mean, attention, or gated attention), and a simple classifier is trained on the resulting slide-level vector.
This approach results in overall linear complexity with respect to the number of input patches, a significant reduction from the O(N2) complexity of classic transformer-based aggregators.
Experimental Evaluation
Datasets and Protocols
CAPRMIL is extensively evaluated on four major computational pathology benchmarks:
- CAMELYON16 (tumor detection, binary classification, bags up to 20,000 patches)
- TCGA-NSCLC (lung cancer, binary classification)
- PANDA (prostate ISUP grading, 6-class classification)
- BRACS (breast lesion coarse classification, 3-class problem with atypical, benign, malignant categories)
For all datasets, patches are extracted at canonical resolutions and featurized using the frozen UNIv1 encoder.
Discriminative Power and Efficiency
CAPRMIL, even when coupled with vanilla mean-pooling aggregation, achieves competitive slide-level classification performance—matching or closely trailing SOTA methods—with dramatic gains in efficiency. Notable highlights include:
Resource Utilization
When analyzing practical deployment considerations (GPU memory, training time, accuracy), CAPRMIL stands out for minimal hardware demand yet maintains top-tier accuracy.
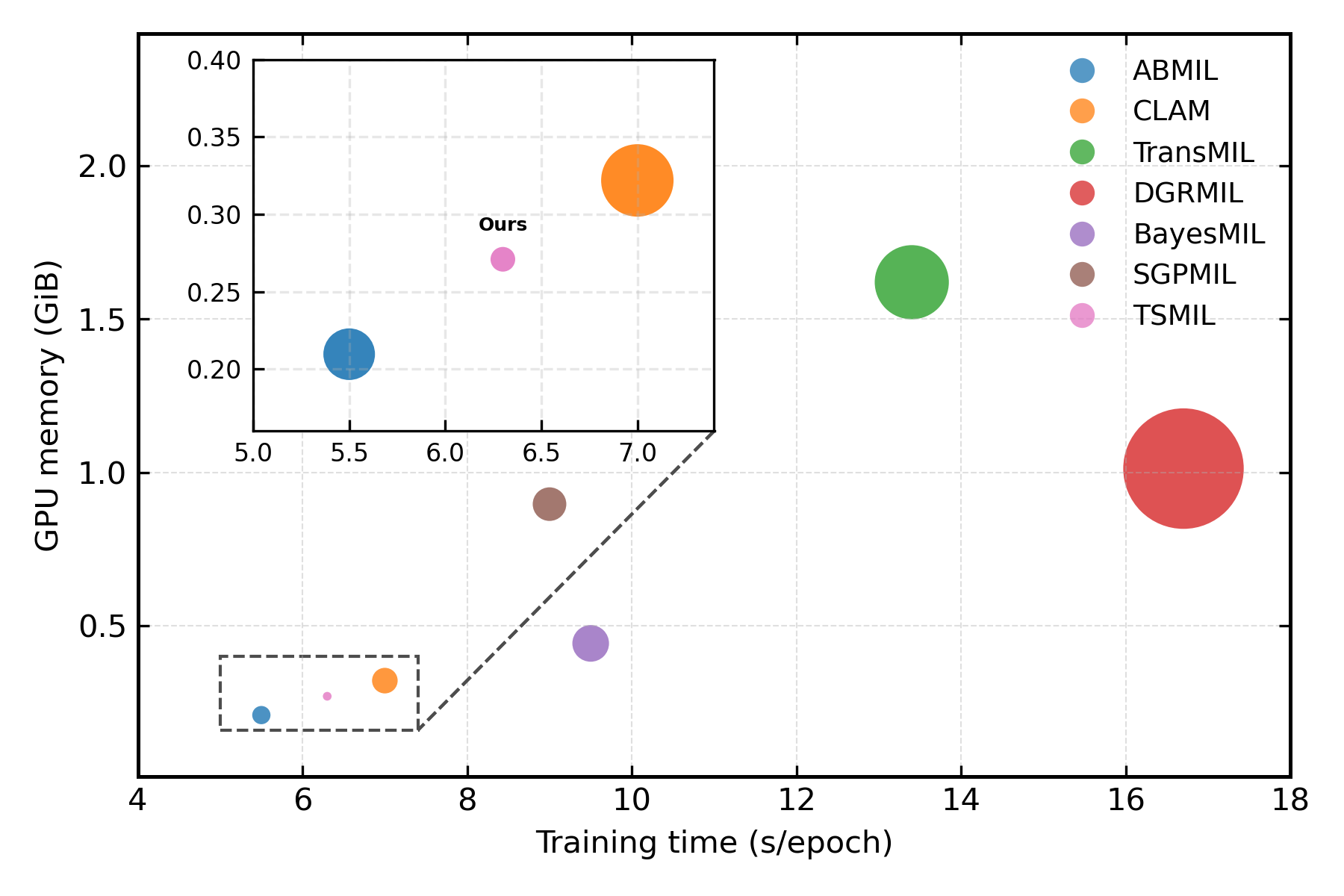
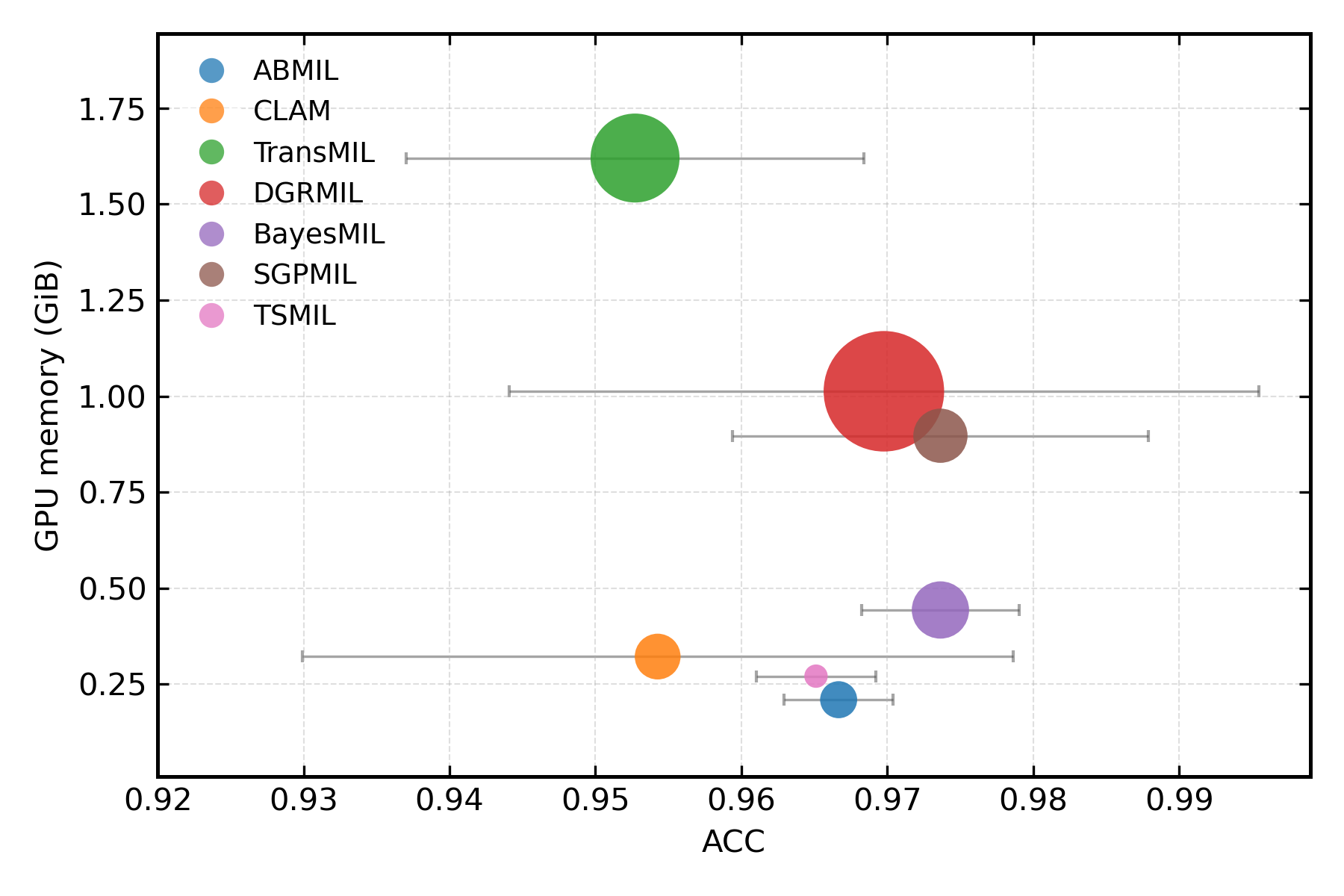
Figure 4: (a) GPU memory footprint and training time. (b) Memory efficiency versus classification accuracy across models.
Robustness and Modularity
Ablation studies on core architectural hyperparameters (clusters M, heads H, MLP expansion) reveal broad insensitivity, with optimality typically reached with low numbers (e.g., M=4, H=8). Further, swapping the final aggregation mechanism (mean, attention, gated) demonstrates that the gains are predominantly secured during the context-aware encoding phase, not aggregation—showing CAPRMIL's broad modularity.
Implications and Future Directions
CAPRMIL’s design leads to several direct theoretical and practical implications for MIL in computational pathology:
Conclusion
CAPRMIL demonstrates that efficient, context-aware patch tokenization fundamentally advances MIL for WSI analysis: context modeling at the embedding level removes dependence on high-parameter, computation-heavy aggregators, yielding robust, modular, and efficient pipelines. Future developments could leverage this framework for multimodal integration, uncertainty quantification, and scaling to even higher dimensional biomedical datasets.