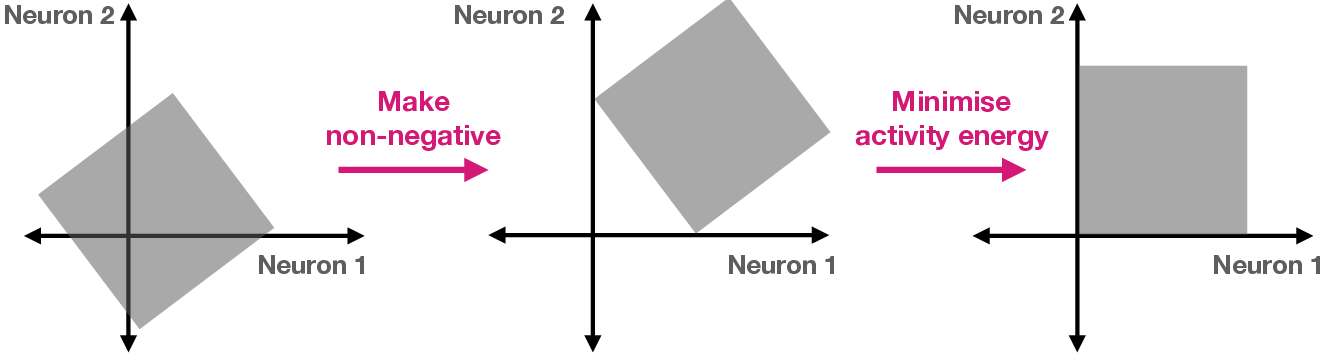
- The paper reveals that a few essential biological constraints (nonnegativity, energy efficiency, wiring costs) largely determine the algorithms neural networks implement.
- It demonstrates that top-down task demands and architectural constraints require these minimal details to explain the observed single-neuron tuning.
- Empirical evidence across domains supports linking these constraints to more interpretable, monosemantic representations in both biological and artificial systems.
How Much Neuroscience Does a Neuroscientist Need to Know? – An Authoritative Summary
Introduction and Central Claims
The paper "How much neuroscience does a neuroscientist need to know?" (2601.02063) interrogates the extent to which the brain’s learned algorithms are constrained by biological detail. The central thesis is that although biological realities undeniably impact neural computation, remarkably few neurobiological details substantively constrain which algorithms neural circuits can embody—provided the substrate is a neural network. The authors argue that a confluence of task demands, architectural (connectionist) constraints, and a select set of biological implementation details—most notably nonnegativity of activations, energetic constraints, and wiring costs—suffice to explain much of the observed regularity in neural algorithms down to the level of single-neuron tuning. This perspective motivates a unified approach between mechanistic interpretability in artificial neural networks (ANNs) and systems neuroscience.
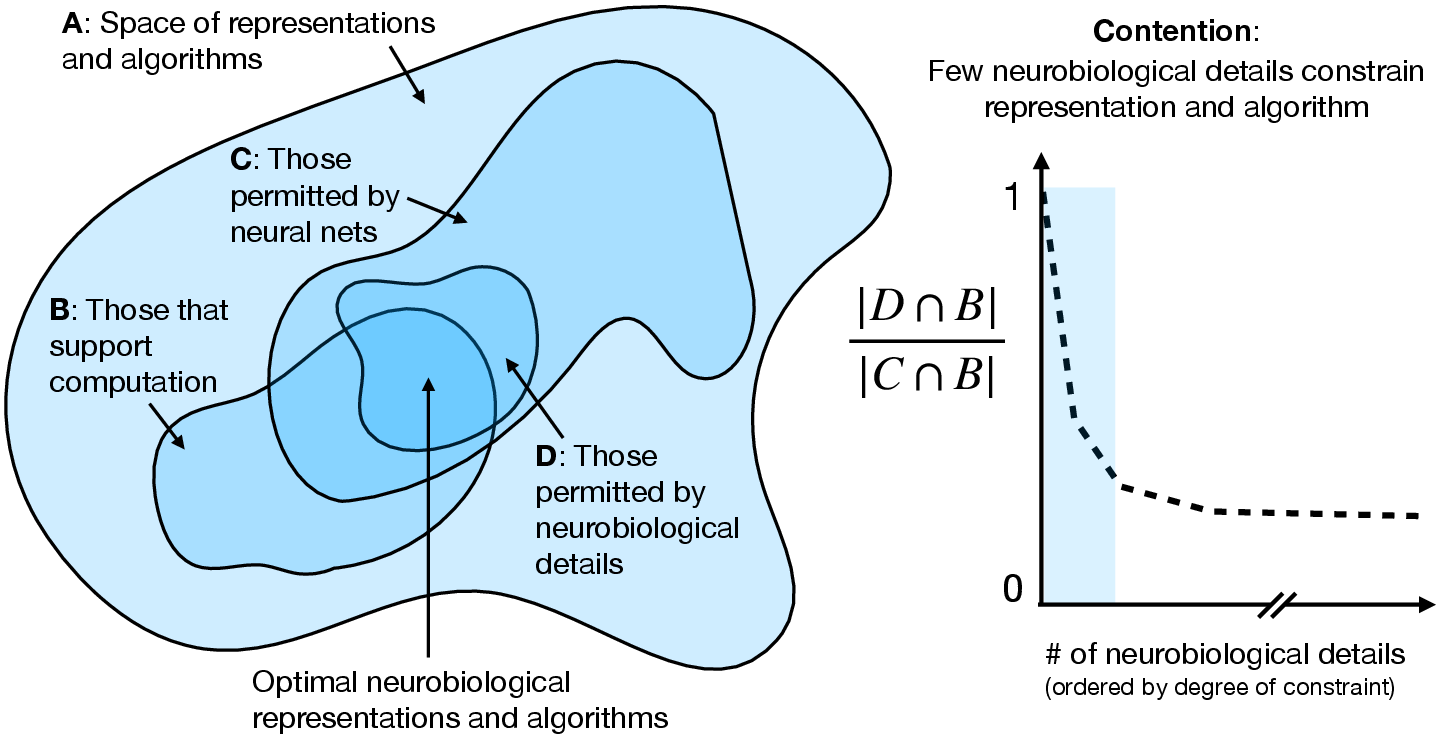
Figure 1: Conditioned on a neural network substrate, few biological details further constrain the space of effective algorithms implementing brain computations.
From Top-Down Task Constraints to Algorithmic Implementation
The paper situates its analysis within Marr’s levels—computational, algorithmic, implementation—with a focus on how implementation-level details back-constrain the space of plausible algorithms capable of task performance. Top-down task constraints (from ethology, cognition, psychology) are vital but insufficient: they delimit the computational goals but underdetermine their realization in neural circuits. Historically, many effective models of neural computation (e.g., cerebellar circuitry, reinforcement learning in dopamine systems, continuous attractor networks, efficient coding theory) have succeeded by positing only the most minimal biological details—point neurons, weighted connections, absence of spike timing, or rich diversity in inhibitory interneurons or synaptic rules [e.g., (2601.02063) references].
This top-down progression, however, does not explain why particular implementations (down to single-neuron tuning) are observed in neural data, nor does it address the challenge of mechanistically inferring the brain’s algorithm from neural activity. Architectural constraints—such as connectionist (ANN-like) representations—further reduce the hypothesis space, but neural populations can still solve the same task in myriad ways due to symmetries in the parameterization space.
Biological Constraints as Symmetry Breakers in Neural Representation
The authors' precise technical insight is that specific biological implementation details serve to break important symmetries (scaling, rotation, permutation) present in generic connectionist systems, thereby biasing learning toward “brain-like” solutions and making individual neuron tuning functional and interpretable.
Crucially, these constraints are not merely engineering artifacts but appear in both biological and high-performing artificial systems, rendering ANNs more interpretable and their units more monosemantic when such constraints are imposed [bricken2023monosemanticity].
Predictive and Explanatory Successes
Several lines of evidence substantiate these claims:
- Cross-domain Algorithmic Convergence: Task-optimized ANNs, with appropriate constraints, converge to solutions structurally and functionally matching recorded neural data in vision [yamins_performance-optimized_2014], audition [singer_sensory_2018], prefrontal cortex [mante_context-dependent_2013], and hippocampal circuits (e.g., grid cells, CANNs) [sorscher_unified_2023].
- Single-Neuron Tuning and Mechanism Identification: Single-neuron responses, shaped by these constraints, become diagnostic of the underlying algorithm (e.g., temporal difference-like dopamine signals, D1/D2 pathway selectivity, grid-cell modularity).
- Algorithm Selection by Constraint Competition: Synthetic tasks (such as the “twisted XOR”) show that the choice of algorithm implemented by the network can be determined by the balance of energetic costs associated with different representational bases, which ultimately manifest at the single-neuron level.
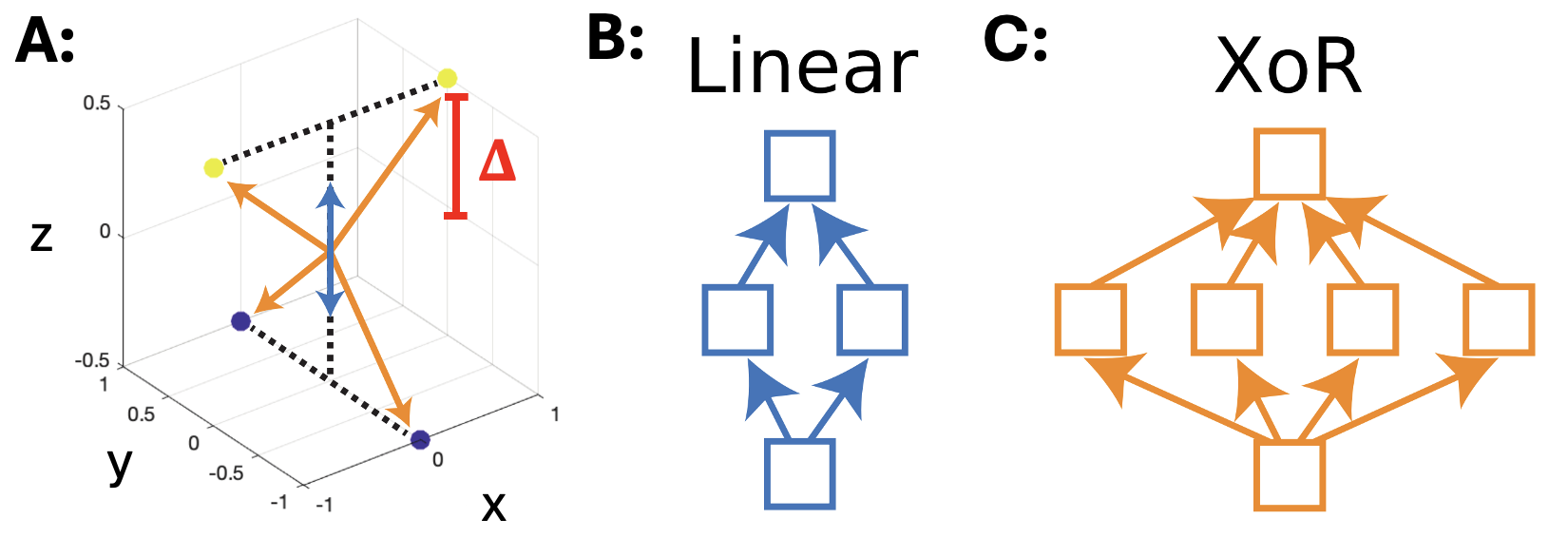
Figure 3: In the twisted XOR task, changing the separability parameter Δ alters the energetic cost landscape, inducing a shift in the number and specialization of neurons encoding the solution.
Implications for Theoretical Neuroscience and AI Interpretability
The authors' framework has broad consequences:
- Unification of Neurointerpretability and Mechanistic Interpretability in AI: Since the neural tuning and algorithmic structure result from a small set of constraints, both biological and artificial systems can be reverse-engineered via the same mechanistic methodologies.
- Limits of Biological Detail: Many biological features (spiking, dendrites, microcircuit details) may be largely orthogonal to algorithm selection in most brain regions, serving their implemented algorithm rather than fundamentally constraining it. However, some exceptions exist (timing-based computation, noncanonical plasticity).
- Minimal Sufficient Detail for Modeling and Interpretation: To capture the brain’s chosen algorithm, it is essential to include constraints like nonnegativity and energetic efficiency in models, as omitting them leaves critical interpretive ambiguity at the level of neurons.
- Pragmatic Modeling Guidance: The work substantiates the legitimacy—and, in some cases, necessity—of building single-neuron level explanations, as biological constraints “align” neuron tuning with the computational structure, making them robust markers of algorithm.
Future Directions
The thesis, while compelling, is not claimed to be proven for all brain systems, particularly in high-dimensional, complex domains such as vision or language where the task and optimal algorithm remain less well-characterized. Systematic computational and experimental work is needed to rigorously establish which biological details are essential for constraining algorithms in these domains versus which can be safely abstracted.
The framework provides a foundation for future research in:
- Predicting When and Why Modularity Emerges: Detailed theoretical and empirical studies, such as those based on the “range” rather than “independence” of input factors, will elucidate the exact conditions that lead to modular neuron tuning [dorrell_range_2025].
- Extending Mechanistic Interpretability: Further advances in interpretable and monosemantic neural architectures in AI can be directly mirrored in systems neuroscience, closing the loop between artificial and biological intelligence research.
- Decomposing Complexity in Brain Circuits: By identifying the minimal set of constraints required to explain neural algorithms at the level of interest, neuroscientists can focus on key explanatory variables, facilitating tractable modeling and experimental design.
Conclusion
This paper provides a rigorous argument that a limited set of biological implementation details—specifically, nonnegativity, energy efficiency, and wiring constraints—interact with task and architectural (connectionist) constraints to shape not only the algorithm that is learned in brains but also the interpretable structure observed at the level of single neurons. This perspective underpins the systematic use of single-neuron (rather than population-manifold) analyses to infer neural algorithms and strongly motivates the continued synthesis of computational neuroscience and mechanistic AI interpretability. Going forward, experimentalists and theorists alike are encouraged to maintain a broad technical familiarity with neuroscience, but to recognize that only a handful of biological features may be critical in delimiting the space of algorithms accessible to the brain.
References
- Paper: "How much neuroscience does a neuroscientist need to know?" (2601.02063)
- Bricken et al., On monosemanticity in sparse autoencoders [bricken2023monosemanticity]
- Dorrell et al., "Range, not Independence, Drives Modularity in Biological Inspired Representation" [dorrell_range_2025]
- Sorscher et al., "A Unified Theory for the Origin of Grid Cell Patterns" [sorscher_unified_2023]
- Yamins et al., "Performance-optimized hierarchical models predict neural responses in higher visual cortex" [yamins_performance-optimized_2014]
- Mante et al., "Context-dependent computation by recurrent dynamics in prefrontal cortex" [mante_context-dependent_2013]
- Koulakov et al., "Orientation Preference Patterns in Mammalian Visual Cortex: A Wire Length Minimization Approach" [koulakov_orientation_2001]
- Harris et al., "Synaptic energy use and supply" [harris_synaptic_2012]