UniX: Unifying Autoregression and Diffusion for Chest X-Ray Understanding and Generation
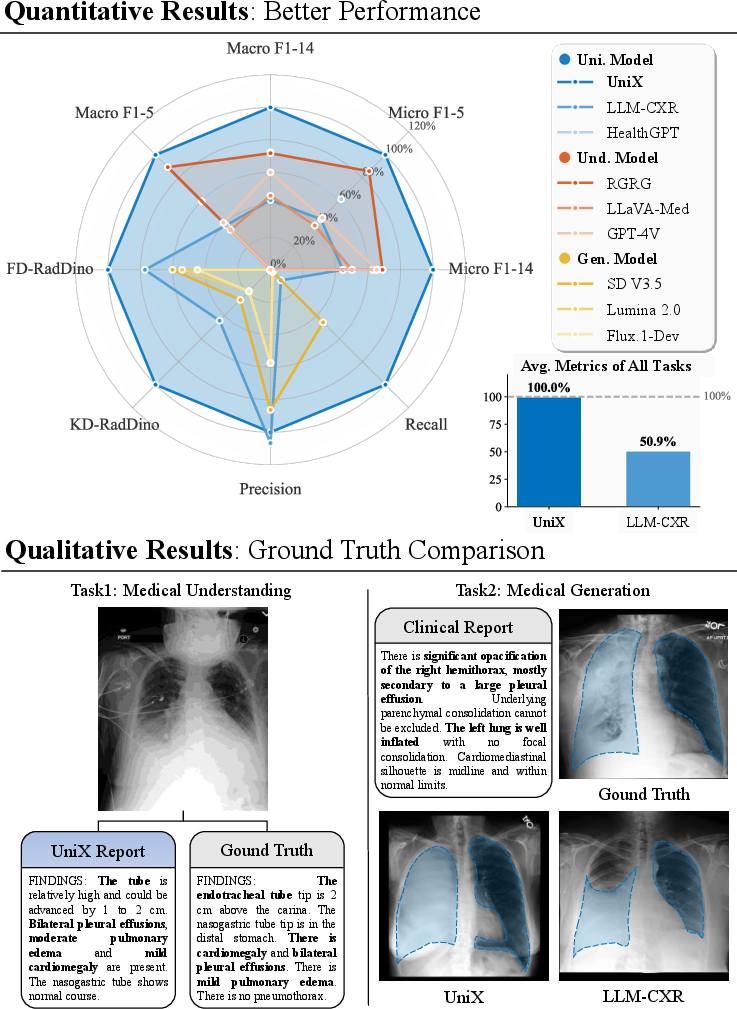
Abstract: Despite recent progress, medical foundation models still struggle to unify visual understanding and generation, as these tasks have inherently conflicting goals: semantic abstraction versus pixel-level reconstruction. Existing approaches, typically based on parameter-shared autoregressive architectures, frequently lead to compromised performance in one or both tasks. To address this, we present UniX, a next-generation unified medical foundation model for chest X-ray understanding and generation. UniX decouples the two tasks into an autoregressive branch for understanding and a diffusion branch for high-fidelity generation. Crucially, a cross-modal self-attention mechanism is introduced to dynamically guide the generation process with understanding features. Coupled with a rigorous data cleaning pipeline and a multi-stage training strategy, this architecture enables synergistic collaboration between tasks while leveraging the strengths of diffusion models for superior generation. On two representative benchmarks, UniX achieves a 46.1% improvement in understanding performance (Micro-F1) and a 24.2% gain in generation quality (FD-RadDino), using only a quarter of the parameters of LLM-CXR. By achieving performance on par with task-specific models, our work establishes a scalable paradigm for synergistic medical image understanding and generation. Codes and models are available at https://github.com/ZrH42/UniX.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
UniX: A simple explanation
What is this paper about?
This paper introduces UniX, a computer program that can both understand chest X-ray images (like writing a short report about what it sees) and create new, realistic chest X-ray images. The big idea is to do both jobs well in one system, without the two jobs getting in each other’s way.
What questions were the researchers asking?
- Can one model be good at both “understanding” X-rays (reasoning and reporting) and “generating” X-rays (making high‑quality images)?
- How can we stop these two goals from fighting each other? Understanding needs summarizing meaning, while generation needs drawing tiny pixel details.
- Can the “understanding” part help the “generating” part make more accurate, medically consistent images?
How did they do it?
Two skills, two goals
- Understanding = turning an image into meaning and words. Think of it like smart autocomplete that looks at the picture and writes the report one word at a time.
- Generation = turning instructions into a detailed image. Think of it like starting from TV static and slowly clearing it up into a clear X-ray.
Because summarizing meaning and drawing pixels are very different, trying to do both with the exact same brain usually causes problems.
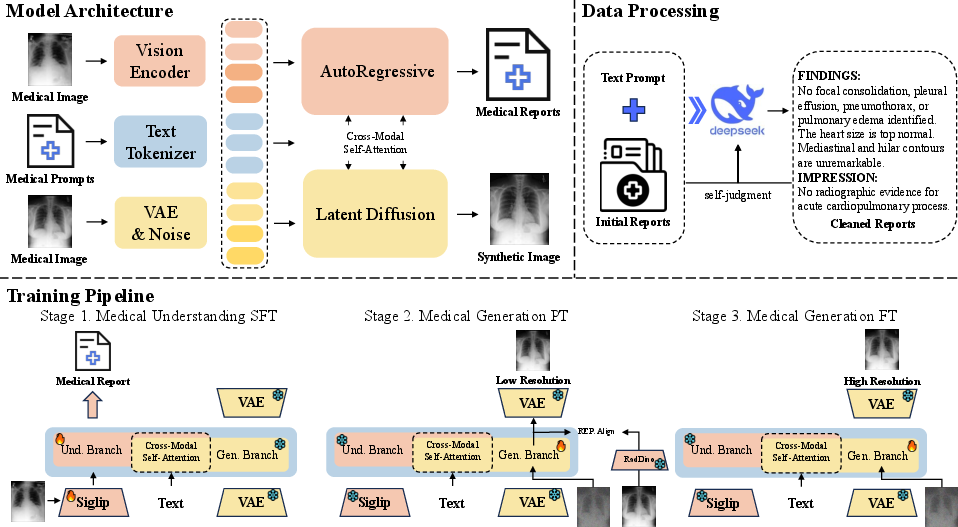
A two-branch design: one brain for each job
- Understanding branch (autoregression): Works like predictive text. It looks at the image and writes the report word by word, building up meaning.
- Generation branch (diffusion): Works like “unblurring” an image. It starts with noisy “static” and removes noise step by step until a realistic X-ray appears.
To make generation efficient, UniX draws in a compressed space (like editing a small “zip” version of the image) and then converts it back to a full image. This uses a tool similar to zipping and unzipping files.
Letting the two branches “talk”
UniX uses a mechanism called cross-modal self-attention. In everyday terms: while the image is being drawn, the understanding branch whispers helpful hints like “put a small shadow in the lower left lung” or “make the heart slightly enlarged.” This guidance helps the generated image match medical meaning, not just look pretty.
Training in three steps
To keep things stable and avoid the two jobs fighting, UniX learns in stages:
- Stage 1: Train understanding only. The model learns to read X-rays and write good reports.
- Stage 2: Freeze understanding, train generation at low resolution. The model learns to draw basic, low-res X-rays guided by the understanding features.
- Stage 3: Still keep understanding frozen, fine-tune generation at higher resolution for sharper, more detailed images.
Cleaning the training data
Real hospital reports can be messy (extra notes, formatting junk). The authors used a LLM to clean reports, keeping only the medically important parts. Cleaner text helps the model learn the right connections between images and words.
What did they find?
- Stronger understanding: UniX improved its report-quality score (Micro-F1) by about 46% compared to a previous unified model.
- Better image generation: UniX improved a key image-quality metric (FD-RadDino, where lower is better) by about 24%.
- Efficient: It did this with roughly one-quarter of the parameters of a popular baseline (LLM-CXR), so it’s smaller but better.
- Competitive with expert systems: Even though UniX handles two jobs, it performed similarly to specialized single-task models in both understanding and generation.
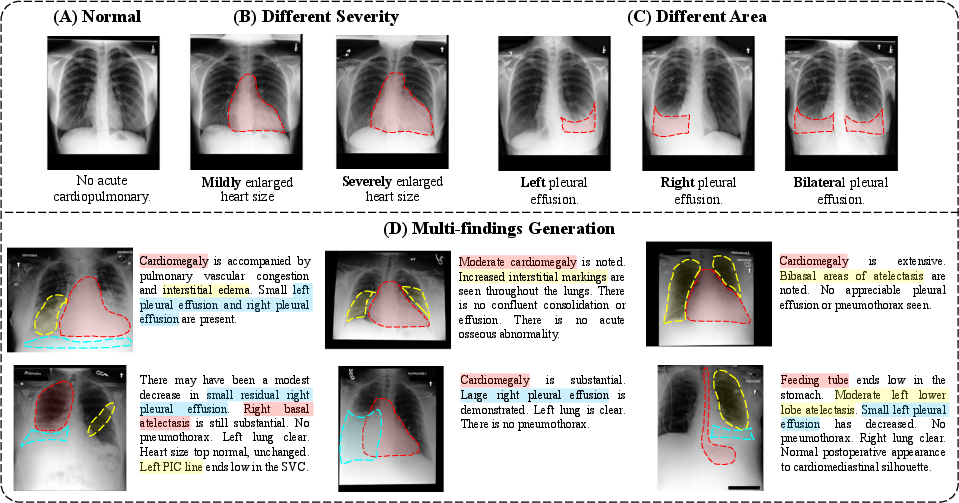
- More consistent across diseases: It generated clearer, more accurate images for many types of findings (like lung issues) and aligned well with the clinical text.
An important lesson from their tests: after training the understanding branch, it’s best to keep it frozen while training the generation branch. If you keep updating both at once, they interfere and both get worse.
Why does this matter?
- Better clinical tools: A system that both understands and generates X-rays could help radiologists check findings, create teaching examples, and explain cases more clearly.
- Higher-quality synthetic data: Realistic, medically accurate synthetic X-rays can help train and improve other medical AI systems, especially for rare conditions.
- Efficiency and reliability: Splitting the two tasks into their own branches reduces internal conflicts and makes the system more stable and accurate.
- A general blueprint: The “two-branch + guided conversation” idea can be used beyond chest X-rays, potentially improving other medical imaging areas (like CT or MRI) and even non-medical image tasks.
In short, UniX shows that you can get the best of both worlds—smart understanding and sharp generation—by giving each task its own space to think, and letting them collaborate at the right moments.
Knowledge Gaps
Knowledge gaps, limitations, and open questions
Below is a concise list of what remains missing, uncertain, or unexplored, framed to be concrete and actionable for future research.
- External generalization: Validate UniX on out-of-institution chest X-ray datasets (e.g., CheXpert, NIH CXR14, RSNA Pneumonia, Open-i) to quantify domain-shift robustness and performance drop.
- Modality transferability: Assess transfer to other medical modalities (CT, MRI, ultrasound) and views (lateral CXR), including multi-view consistency and cross-view conditioning.
- Clinical validation: Conduct blinded reader studies with radiologists to assess clinical correctness, utility, and safety of both generated reports and images versus automated metrics.
- Metric reliability: Evaluate correlation between FD-/KD-RadDino and clinician judgments; test sensitivity of RadDino-based metrics to image artifacts and pathology severity/location.
- Uncertainty and calibration: Measure calibration of report-generation outputs and uncertainty propagation (e.g., expected calibration error, selective prediction) rather than relying solely on CheXbert F1.
- Rare/long-tail pathologies: Benchmark generation and understanding on rare findings (e.g., pneumothorax tension, subtle interstitial patterns) with stratified analyses of fidelity and detection.
- Spatial control and grounding: Quantify location/severity control (e.g., bounding-box/heatmap consistency, Grad-CAM alignment) rather than qualitative exemplars; add structured spatial constraints during generation.
- Cross-modal self-attention ablation: Compare the proposed cross-modal self-attention against standard cross-attention, gating, and adaptor-based fusion; visualize attention maps to verify semantic guidance and reduce spurious couplings.
- Training strategy alternatives: Systematically test joint/alternating updates, curriculum schedules, branch-specific adapters (LoRA), and knowledge distillation to reduce interference without freezing one branch.
- Initialization effects: Ablate Janus-Pro initialization versus medical-only pretraining or random init to quantify dependency on natural-image semantics and potential misalignment with clinical textures.
- VAE capacity and resolution: Analyze how VAE downsampling ratio and channel count affect structural fidelity; scale to clinical resolutions (1024–2048 px) and assess sharpness, micro-calcifications, and fine lung markings.
- Scaling laws: Explore performance–parameter tradeoffs by varying LLM and diffusion backbone sizes; characterize diminishing returns and optimal capacity allocation across branches.
- Data cleaning impacts: Provide quantitative analyses of DeepSeek report cleaning (e.g., fidelity to original diagnoses, removal of clinically relevant context, bias introduced); release reproducible prompts and rules.
- Robustness to noisy/incomplete prompts: Stress-test understanding and generation with real-world, free-form clinician inputs, incomplete histories, and contradictory directives; measure failure modes and graceful degradation.
- Privacy and memorization: Conduct membership inference and nearest-neighbor analyses to measure patient-level leakage from generated images; include CheXGenBench privacy metrics and formal audits.
- Safety evaluation: Identify and quantify harmful failure cases (e.g., anatomically implausible images, hallucinated devices/pathologies); develop and test safety filters/validators for deployment.
- Closed-loop synergy: Test whether synthetic images can improve the understanding branch (e.g., semi-supervised or augmentation pipelines); measure gains and risks of synthetic-to-real transfer.
- Pathology-specific alignment: Extend per-pathology evaluation to severity grading, laterality, and multi-finding interactions; verify co-occurrence realism and causal consistency with reports.
- Adversarial robustness: Evaluate susceptibility to adversarial prompts and perturbations (text/image), including prompt injection or conflicting instructions during joint attention.
- Interpretability: Provide token-/pixel-level explanations linking generated image regions to specific report statements; assess clinician trust via interpretable rationales.
- Efficiency and latency: Measure training/inference speed and memory for both branches (including sampling steps), and benchmark feasibility on clinical hardware; explore fast samplers or distillation for diffusion.
- Multi-view and temporal coherence: Extend to longitudinal CXR series and multi-view synthesis; quantify temporal consistency and progression realism (e.g., disease evolution).
- Structured conditioning: Investigate conditioning on structured inputs (labels, masks, contours, device metadata) and compare against text-only conditioning for controllability and fidelity.
- Generalization to multilingual settings: Test understanding/generation with multilingual reports; evaluate cross-lingual alignment and bias introduced by English-centric cleaning and metrics.
- Reproducibility: Release full data-cleaning scripts, model checkpoints per stage, and detailed seeds/settings; run cross-lab reproducibility studies to quantify variability.
- Fairness and bias: Audit performance across demographics (age, sex, race), clinical devices (portable vs. fixed), and acquisition protocols; implement bias mitigation strategies where gaps are found.
Practical Applications
Immediate Applications
Below are actionable use cases that can be deployed now (primarily in research, education, and non‑diagnostic clinical support settings), derived from UniX’s dual‑branch architecture, cross‑modal self‑attention, data cleaning pipeline, and training strategy.
- Radiology report drafting assistant for chest X-rays (Healthcare, Software)
- Use the autoregressive understanding branch to auto‑draft structured, pathology‑aware reports that radiologists can edit and sign.
- Potential workflow: PACS/RIS plug‑in that ingests DICOM, generates a draft, and surfaces key findings and uncertainties.
- Assumptions/dependencies: Assistive use only (non‑autonomous); local domain adaptation to hospital data; governance for human‑in‑the‑loop review; integration with PACS/RIS and EHR.
- Image–report consistency checker using alignment metrics (Healthcare, Quality/Compliance, Software)
- Leverage UniX’s alignment score and cross‑modal features to flag report–image mismatches (e.g., missing or contradictory findings).
- Potential tool: a QA dashboard that audits recent reports and highlights cases for secondary review.
- Assumptions/dependencies: Access to imaging and reports; validated thresholds per site; guardrails to avoid over‑flagging; radiologist oversight.
- Pathology‑controlled case generator for training and competency assessment (Education, Academia)
- Use diffusion generation guided by understanding features to synthesize cases with controllable lesion type, location, and severity for resident training, OSCEs, and CME.
- Potential product: an “X‑ray curriculum generator” that builds practice sets tied to specific learning objectives.
- Assumptions/dependencies: Clear labeling of synthetic data; adherence to institutional training policies; evaluation with PRDC/FD‑RadDino to ensure fidelity/diversity.
- Targeted data augmentation for imbalanced pathologies (Academia, Healthcare AI R&D)
- Generate synthetic images for rare findings to balance training sets for classifiers/VLMs and improve downstream performance.
- Potential workflow: “Generate‑to‑Learn” pipeline that selects under‑represented classes, synthesizes cases, retrains models, and re‑evaluates.
- Assumptions/dependencies: Utility validated on task‑specific metrics; monitoring for synthetic artifacts or distribution shift; governance for synthetic/real data ratios.
- LLM‑based report cleaning and normalization (Software, Academia)
- Deploy the paper’s data cleaning pipeline (e.g., DeepSeek‑assisted prompts) to strip non‑diagnostic content, standardize phrasing, and improve training targets and downstream NLP/analytics.
- Potential tool: “RadClean” batch processing for legacy archives and new reports.
- Assumptions/dependencies: Clinician review of cleaning templates; safeguards to avoid removing clinically relevant nuance; site‑specific customization.
- Benchmarking and model selection with UniX’s metrics (Academia, Industry)
- Adopt FD‑RadDino, KD‑RadDino, and PRDC as standardized measures to compare generative fidelity, alignment, and diversity for medical imaging models.
- Potential product: an evaluation suite that plugs into training pipelines and CI/CD for medical AI.
- Assumptions/dependencies: Availability of RadDino feature extractors; reproducible protocol (e.g., CheXGenBench); careful interpretation of metric deltas across datasets.
- Parameter‑efficient unified modeling for prototyping (Software, Academia)
- Use UniX’s smaller footprint (≈1.5B vs. 12B) to build research prototypes that jointly perform report understanding and image generation without compromising quality.
- Potential tool: “UniX‑Lite” SDK/API for rapid lab deployment and ablation studies.
- Assumptions/dependencies: GPU resources for diffusion; adherence to open‑source licensing; local fine‑tuning for domain shift.
- Semi‑automated labeling and pre‑annotation (Healthcare AI Ops, Industry)
- Apply the understanding branch to pre‑label chest X‑rays for pathology categories and populate structured fields, accelerating dataset curation.
- Potential workflow: annotation platform integration with human validation and disagreement resolution.
- Assumptions/dependencies: Clear operating thresholds; radiologist adjudication; bias/fairness audits across patient subgroups.
Long-Term Applications
These use cases require further clinical validation, scaling, regulatory approvals, or extension of UniX’s approach to other modalities and settings.
- Clinically validated end‑to‑end decision support for chest X-rays (Healthcare, Policy)
- Combine report drafting, consistency checking, and risk triage into a single assistive platform with audit trails and continuous monitoring.
- Potential product: a certified CDS tool that prioritizes cases (e.g., suspected pneumothorax) and ensures report–image coherence.
- Assumptions/dependencies: Prospective trials, multi‑site generalization, regulatory clearance, robust fail‑safes, and medico‑legal frameworks.
- Cross‑modality unified foundation models (CT/MRI/Ultrasound) (Healthcare, Software, Robotics)
- Extend the dual‑branch (autoregression + diffusion) architecture and cross‑modal self‑attention to continuous imaging modalities for high‑fidelity synthesis and understanding.
- Potential tools: modality‑specific VAEs, multi‑scale diffusion backbones, unified report generation across modalities.
- Assumptions/dependencies: Large, diverse, modality‑specific datasets; scalable training; domain‑aware evaluation metrics.
- Privacy‑preserving synthetic cohort creation and data sharing (Policy, Healthcare, Academia)
- Use controllable generation to produce site‑calibrated synthetic datasets that protect PHI while preserving utility for algorithm development and benchmarking.
- Potential workflow: “Synthetic sandbox” for external collaborations and pre‑training.
- Assumptions/dependencies: Privacy leakage assessments; policy guidance on synthetic data governance; standards for disclosure and labeling.
- Human‑in‑the‑loop active learning pipelines (Industry, Healthcare AI Ops)
- Integrate UniX as a selector/generator to identify uncertain cases, synthesize counterexamples, and iteratively improve models with radiologist feedback.
- Potential product: an AL orchestration platform that couples uncertainty estimation with targeted synthetic augmentation.
- Assumptions/dependencies: Interfaces for expert review; reliable uncertainty quantification; monitoring for synthetic bias.
- Automated radiology QA and governance at scale (Policy, Healthcare Quality)
- Institutional adoption of alignment checks and pathology‑specific fidelity measures to continuously audit reporting quality and compliance.
- Potential tool: a governance cockpit that tracks drift, error rates, and corrective actions.
- Assumptions/dependencies: Policy buy‑in; clear escalation protocols; integration with quality management systems; fairness audits.
- Rare disease registries and clinical trial simulation (Healthcare, Academia, Policy)
- Generate realistic, pathology‑controlled cohorts to stress‑test algorithms and explore trial scenarios where real data is scarce.
- Potential workflow: synthetic arms for feasibility studies and power analyses.
- Assumptions/dependencies: Ethical frameworks; validation of clinical relevance; clear limitations communicated to stakeholders.
- Edge and resource‑constrained deployment (Software, Devices)
- Leverage parameter efficiency to move parts of the pipeline toward on‑prem or edge devices (e.g., local pre‑labeling, draft generation).
- Potential product: hospital‑grade appliances with secure inference and local governance.
- Assumptions/dependencies: Further model compression (quantization, distillation), hardware support for diffusion, robust MLOps.
- Generative simulation for device calibration and algorithm robustness (Industry, Regulatory Science)
- Use controlled synthesis to emulate acquisition differences, artifacts, and distribution shifts, aiding calibration and stress testing of CAD systems.
- Potential workflow: synthetic test benches for robustness and post‑market surveillance.
- Assumptions/dependencies: Domain models of noise/artifacts; collaboration with manufacturers; acceptance of synthetic testing in regulatory pathways.
Glossary
- Alignment Score: A metric assessing how well generated images align with text prompts. "we measure generation quality with FD-RadDino and KD-RadDino, assess imageâtext consistency with the Alignment Score, and evaluate accuracy and diversity using the four PRDC metrics."
- Autoregressive sequence modeling: Modeling where each token is predicted conditioned on previous tokens in sequence. "The understanding branch formulates multimodal comprehension as an autoregressive sequence modeling problem."
- Bottleneck-free multimodal interaction: Cross-modal interaction without compressive bottlenecks that could lose information. "echoing insights from BAGEL~\cite{deng2025emerging} on the value of bottleneck-free multimodal interaction."
- CheXbert F1 score: A radiology-specific F1-based metric for evaluating report correctness. "we primarily evaluate the medical reliability of generated reports using the CheXbert F1 score~\cite{smit2020chexbert} between generated and ground-truth reports."
- Cross-entropy loss: Standard loss for next-token prediction measuring divergence between predicted and true distributions. "The cross-entropy loss is computed over the autoregressive predictions for all tokens in :"
- Cross-modal self-attention: An attention mechanism jointly operating over tokens from different modalities to enable dynamic interactions. "we introduce a cross-modal self-attention mechanism to dynamically guide the generation process with understanding features."
- Decoupled dual-branch design: An architecture that separates understanding and generation into distinct branches to avoid interference. "This section, we describe how UniX exploits its decoupled autoregressiveâdiffusion dual-branch design through data processing, model configuration, and a three-stage training pipeline."
- Discretized generation paradigms: Methods that generate images via discrete token vocabularies, potentially losing fine details. "most current unified medical foundation models still rely on discretized generation paradigms, whose outputs are constrained by vocabulary granularity and fail to recover fine structural details in medical images."
- FD-RadDino: A Fréchet-distance-like distribution metric computed in the RadDino feature space to assess image quality. "a 24.2\% gain in generation quality (FD-RadDino)"
- Feature interference: Harmful cross-task effects when shared representations carry conflicting objectives. "This approach, unfortunately, can introduce task competition and feature interference, which degrades performance in both understanding and generation."
- Generative trajectory: The evolution of the generated sample across diffusion steps or model iterations. "This design allows semantic representations from the understanding branch to directly modulate the generative trajectory"
- H-LoRA: Task-specific low-rank adaptation modules used to partially separate parameters for different tasks. "HealthGPT~\cite{lin2025healthgpt} mitigates this issue through task-specific H-LoRA modules, offering a structured compromise but not a fundamental solution."
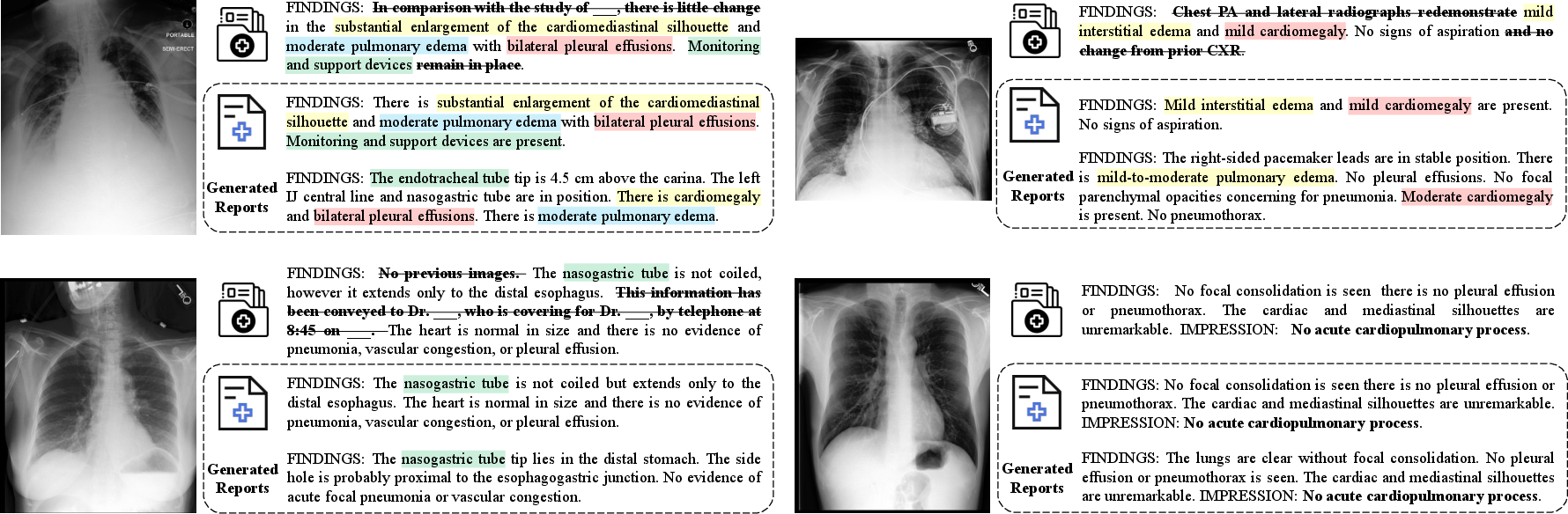
- Hallucinations (in medical reporting): Fabricated or unsupported content produced by a model in generated reports. "Figure~\ref{fig:und_visual} illustrates the critical role of data cleaning with DeepSeek in mitigating hallucinations."
- KD-RadDino: A kernel/divergence-based distribution metric in RadDino feature space for generation evaluation. "we measure generation quality with FD-RadDino and KD-RadDino"
- Latent diffusion framework: A diffusion model operating in a compressed latent space instead of pixel space for efficiency and fidelity. "The generation branch adopts a latent diffusion framework that reconstructs medical images from high-level semantics extracted by the understanding branch."
- Latent space: A compressed representation space (e.g., from a VAE) where diffusion or learning operates. "diffusion is performed in a VAE-encoded latent space, which greatly improves efficiency and stability."
- Micro-F1: F1 score computed by aggregating contributions of all classes (micro-averaging), used for multi-label evaluation. "UniX achieves a 46.1\% improvement in understanding performance (Micro-F1)"
- Modality selectors: Indicators that switch projection parameters depending on whether a token comes from understanding or generation streams. "where the modality selectors and are defined as:"
- Multimodal token sequence: A unified sequence combining visual and textual tokens for joint processing. "we define a multimodal token sequence "
- Parameter sharing: Reusing the same model parameters across multiple tasks or heads. "Existing efforts, such as LLM-CXR~\cite{lee2023llm}, often employ parameter sharing and joint multi-task heads for integrated learning."
- PRDC metrics: A set of four measures—Precision, Recall, Density, Coverage—for evaluating fidelity and diversity of generated data. "and evaluate accuracy and diversity using the four PRDC metrics."
- QK normalization: Normalizing query-key projections in attention to stabilize training and improve performance. "The language backbone contains 24 transformer layers and incorporates QK normalization and QKV bias."
- QKV bias: Learnable bias terms added to Q, K, and V projections in attention layers. "The language backbone contains 24 transformer layers and incorporates QK normalization and QKV bias."
- RadDino (image features): A radiology-specific visual representation used to compute feature-based metrics and alignments. "aligning the eighth-layer hidden states of the generation branchâs LLM with RadDino image features using a similarity objective."
- RadGraph: A structure-based evaluation metric for radiology report quality focusing on clinical entities and relations. "Additional evaluation metrics, including BLEU~\cite{papineni2002bleu}, Radgraph~\cite{jain2021radgraph} and ROUGE-L~\cite{lin2004rouge}, are provided in the supplementary material."
- Representation Alignment (REPA): A training technique aligning hidden states with target feature spaces to speed convergence and improve conditioning. "we apply Representation Alignment~\cite{yu2024representation}, aligning the eighth-layer hidden states of the generation branchâs LLM with RadDino image features using a similarity objective."
- Semantic abstraction: Compressing visual information into high-level concepts rather than reconstructing pixels. "these tasks have inherently conflicting goals: semantic abstraction versus pixel-level reconstruction."
- Three-stage training strategy: A staged procedure to separately train and then coordinate understanding and generation branches. "we adopt a three-stage training strategy to progressively align the understanding and generation branches."
- VAE (Variational Autoencoder): A generative model that encodes images into a continuous latent space and decodes them back. "diffusion is performed in a VAE-encoded latent space"
- Velocity field (in diffusion): The vector field that indicates the direction to denoise samples during diffusion training. "the model learns to estimate the target velocity field by minimizing the mean-square error"
- Visionâlanguage pretraining: Pretraining techniques that jointly learn from images and text for downstream multimodal tasks. "visionâlanguage pretrainingâbased medical foundation models have shown remarkable success"
- Visual bag-of-words: A discrete codebook approach that tokenizes images into visual words for generation or understanding. "discrete generation methods based on visual bag-of-words~\cite{van2017neural, esser2021taming}."
Collections
Sign up for free to add this paper to one or more collections.