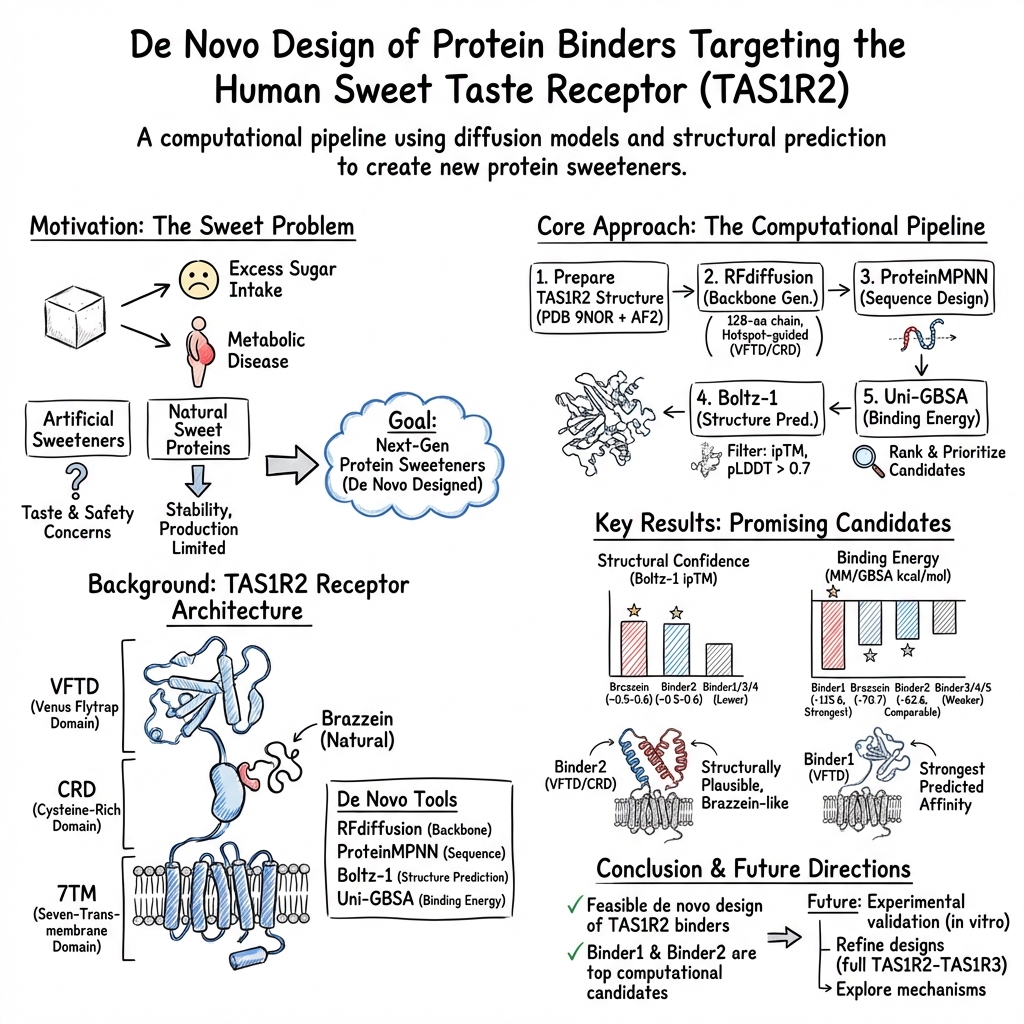
De novo design of protein binders targeting the human sweet taste receptor as potential sweet proteins
Abstract: Excessive consumption of dietary sugars is a major contributor to metabolic disorders, driving global interest in finding alternative sweeteners with reduced caloric impact. Natural sweet proteins, such as brazzein, offer exceptional sweetness intensity with little caloric contribution. However, their widespread use is limited by restricted natural diversity, low stability, and high production costs. Recent advances in structural biology and de novo protein design provide new opportunities to overcome these limitations through rational engineering. In this study, we report an integrated computational pipeline for the de novo design of protein binders targeting the human sweet taste receptor subunit TAS1R2, a key component of the heterodimeric class C G protein-coupled receptor mediating sweetness perception. The workflow combines diffusion-based backbone generation (RFdiffusion), neural network-guided sequence design (ProteinMPNN), structure-based filtering using Boltz-1, and binding energy evaluation via MM/GBSA calculations. Using the recently resolved cryo-EM structure of the TAS1R2 receptor, protein binders were designed to target both the Venus Flytrap Domain and the cysteine-rich domain of TAS1R2. A few designed binders exhibited favorable structural confidence and predicted binding energetics. In particular, Binder2 exhibited brazzein-like structural plausibility through specific short-range CRD contacts, while Binder1 displayed the strongest predicted binding affinity. Structural analyses of the binder-receptor complex revealed distinct binding modes and secondary structure profiles among the designs. This study demonstrates the feasibility of de novo designing protein binders that emulate key functional properties of natural sweet proteins, establishing a computational framework for the rational development of next-generation protein-based sweeteners.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
A simple explanation of the paper: Designing new “sweet” proteins from scratch
1) What is this paper about?
This paper is about creating brand‑new proteins that can make things taste sweet, without adding sugar or many calories. The scientists designed tiny proteins that are predicted to stick to the human sweet taste receptor (the part of your tongue that senses sweetness). If these designed proteins work, they could become next‑generation sweeteners that are very sweet, low‑calorie, and more stable than natural sweet proteins found in some tropical fruits.
2) What questions did the researchers want to answer?
The team wanted to know:
- Can we design, on a computer, brand‑new proteins that attach to the human sweet taste receptor (specifically the TAS1R2 part) in the right place and in the right way?
- Can these designed proteins look and behave like known sweet proteins (such as brazzein) that trigger sweetness?
- Which designs are most likely to stick strongly and correctly to the receptor, so they have the best chance to taste sweet in real life?
3) How did they try to solve it? (What methods did they use?)
Think of the sweet taste receptor like a lock on your taste buds. Sweet things are “keys” that fit the lock and make you taste sweetness. The scientists used a step‑by‑step computer “factory” to design new keys:
- Step 1: Shape maker (RFdiffusion)
- This tool “sketched” many possible shapes for a new protein (like modeling clay into different key shapes) that could fit onto the receptor. They focused on two parts of the TAS1R2 receptor:
- The Venus Flytrap Domain (VFTD), which acts like a clamshell that closes when something sweet binds.
- The Cysteine‑Rich Domain (CRD), which helps send the “sweet” signal.
- Step 2: Detail the shape with “material” (ProteinMPNN)
- Once the shapes were set, this tool chose the exact amino acids (the building blocks of proteins) to build each shape so the protein would be realistic and stable.
- Step 3: Quality check (Boltz‑1, an AlphaFold‑like predictor)
- This AI checked if the designed proteins would fold correctly and fit well on the receptor. It gave confidence scores—basically, how sure the computer is that the design makes sense.
- Step 4: “Stickiness” test (MM/GBSA energy)
- Finally, they estimated how strongly each designed protein would stick to the receptor. Lower (more negative) energy numbers mean stronger sticking.
They also compared their designs to brazzein, a real sweet protein from fruit, to see how close their designs came to something known to work.
4) What did they find, and why is it important?
- Several designs passed the computer checks. Two stood out:
- Binder1: Predicted to stick the strongest to the receptor (best “stickiness” score).
- Binder2: Looked the most realistic overall and had computer scores most similar to brazzein. It also made specific, short‑distance contacts in the CRD of the receptor (the kind of detailed fit you want to see).
- Different designs attached to different parts of the receptor:
- Binder1 and Binder2 mostly targeted the “clamshell” VFTD area (a known sweet‑binding spot).
- Others targeted the CRD or nearby areas.
- Many designs were made mostly of helices (a common, sturdy protein shape), which computers can design confidently.
Why this matters:
- Natural sweet proteins can be extremely sweet with almost no calories, but they can be hard to make and not always stable during cooking or storage.
- If we can design sweet proteins from scratch, we could improve stability, reduce costs, and tailor how they taste—possibly avoiding aftertastes that some artificial sweeteners have.
5) What does this mean for the future?
This work shows that it’s possible to design potential sweet proteins on a computer by aiming them at the human sweet taste receptor. The next steps are crucial:
- Make the top designs (like Binder1 and Binder2) in the lab.
- Test if they really bind to the receptor, actually taste sweet to people, are safe, stable in foods, and affordable to produce.
If these steps succeed, we could have new, low‑calorie sweeteners that taste good, are more stable than current sweet proteins, and help reduce sugar intake—supporting better health without sacrificing sweetness.
Knowledge Gaps
Knowledge gaps, limitations, and open questions
Below is a concise, actionable list of what remains missing, uncertain, or unexplored in the paper.
- No experimental validation of designed binders: no synthesis, expression, folding, or biophysical binding assays (e.g., SPR/BLI/ITC) to confirm receptor engagement.
- Unknown functional modality: whether Binder1–Binder5 act as agonists, partial agonists, antagonists, or allosteric modulators is not tested (e.g., G protein/PLC signaling, Ca2+ flux assays in TAS1R2/TAS1R3-expressing cells).
- Receptor context incompleteness: designs and evaluations are performed primarily against TAS1R2; the functional heterodimer (TAS1R2–TAS1R3) and its inter-domain coupling (VFTD→CRD→TMD) in a membrane context are not modeled or tested.
- Static-state targeting: binders are designed to a single cryo-EM snapshot; multi-state/ensemble considerations (active vs inactive conformations) and state-selective design are not explored.
- Membrane and glycan environment omitted: extracellular glycosylation and the lipid membrane—both critical for class C GPCR conformation—are absent from modeling and energy calculations.
- Reliance on MM/GBSA end-state estimates: binding free energies lack entropic contributions, solvent/membrane realism, and kinetic parameters; no alchemical free-energy or long-timescale MD validation is performed.
- Conflicting predictors remain unresolved: Binder1 shows the best MM/GBSA energy while Binder2 shows the strongest structural confidence (ipTM/confidence_score); no calibration to determine which metric best predicts true activation/sweetness.
- Hotspot selection uncertainty: VFTD/CRD hotspots derive from prior NMR reports; alignment with the newest cryo-EM-defined orthosteric/allosteric sites is not validated (e.g., via computational alanine scanning or mutational sensitivity analysis).
- Fixed binder length (128 aa) and topology: no exploration of sequence-length optimization, alternative topologies (e.g., small disulfide-rich scaffolds like brazzein), cyclic peptides, or multivalent architectures to balance potency, stability, and cost.
- Helical-design bias: acknowledged preference for helical scaffolds (RFdiffusion + ProteinMPNN) is not mitigated; beta-sheet/disulfide-rich scaffolds—common in natural sweet proteins—are under-sampled.
- Undisclosed binder sequences: the paper does not provide sequences for Binder1–Binder5, impeding reproducibility, synthesis, and independent validation.
- Standalone stability unassessed: no in silico or experimental evaluation of thermal stability, pH tolerance, protease resistance, or long-term shelf stability of the designed binders.
- Manufacturability gap: expression systems (E. coli, yeast), yields, solubility, purification feasibility, and production costs are not evaluated.
- Safety and regulatory unknowns: immunogenicity/allergenicity (in silico epitope prediction), toxicity, and regulatory pathways for de novo sweet proteins are not addressed.
- Population variability: TAS1R2/TAS1R3 polymorphisms that influence sweet perception are not considered; binder robustness across common variants remains unknown.
- Cross-reactivity risk: potential binding to TAS1R1/TAS1R3 or other class C GPCRs (umami/bitter pathways) is not assessed, risking off-target taste effects.
- Competitive vs allosteric mechanism: it is unclear whether designed binders compete with natural sweeteners (e.g., glucose/sucralose) or act allosterically; no competition assays performed.
- Downstream signaling and conformation coupling: the capacity of binders to induce TAS1R2 VFTD closure and propagate conformational changes through CRD/TMD (i.e., true receptor activation) is not tested.
- Binding kinetics and sensory time-intensity: on/off rates, onset/lingering profiles, and correlation with human sensory dynamics are not evaluated.
- Saliva and food matrix effects: binder stability in saliva (proteases, ionic strength), pH ranges of foods/beverages, and processing conditions (heat, shear) are untested.
- Interface mapping depth: beyond two CRD contacts in Binder2, a comprehensive per-residue interaction analysis (hydrogen bonds, salt bridges, hydrophobic hotspots) and mutational robustness is lacking.
- Statistical robustness: significance testing on predicted metrics uses small sample sizes and model-derived scores; no experimental ground truth to substantiate statistical conclusions.
- MSA dependence for de novo sequences: Boltz-1 runs with MSA may not yield reliable confidence for novel sequences lacking homologs; effects of using single-sequence/no-MSA modes are not quantified.
- Absolute energy magnitudes and uncertainty: MM/GBSA values (e.g., −115.6 kcal/mol) are not accompanied by uncertainty estimates, replicate runs, or cross-method validation (e.g., different GB/PB models).
- Multi-domain engagement designs: constructs that simultaneously engage VFTD and CRD to better transmit conformational changes are not attempted; potential for cooperative interfaces remains unexplored.
- Antagonism risk in real use: binders might occlude the orthosteric pocket and reduce sweetness from sugars; no assays to test whether they enhance or suppress sweetness in the presence of dietary sweeteners.
- Co-formulation synergy: interaction with existing sweeteners or flavor modulators (synergy/antagonism) is not explored.
- Sensory validation pathway: no plan for human sensory testing (thresholds, relative sweetness to sucrose, aftertaste profile) following in vitro efficacy.
- Data and model transparency: modeling parameters (e.g., Boltz-1 configuration, random seeds), structural models, and docking coordinates are not shared, limiting reproducibility and comparative benchmarking.
Practical Applications
Below are practical applications derived from the paper’s findings, methods, and innovations. They are grouped into immediate and long-term opportunities, with links to relevant sectors, prospective tools/products/workflows, and key assumptions or dependencies that affect feasibility.
Immediate Applications
- Industry (food & beverage): Computational pre-screening of protein sweeteners
- Application: Adopt the paper’s integrated pipeline (RFdiffusion → ProteinMPNN → Boltz-1 → Uni-GBSA) to rapidly generate and triage candidate sweet proteins before wet-lab investment.
- Tools/workflows: Containerized pipeline with preset TAS1R2 hotspot conditioning; plug-in to existing cheminformatics stacks; cloud/HPC execution.
- Assumptions/dependencies: In silico metrics (ipTM, complex_pLDDT, GBSA) correlate sufficiently with real binding and sensory sweetness; access to cryo-EM-derived TAS1R2/TAS1R3 structures.
- Academia (computational biology, structural biochemistry): Reproducible binder-design studies of taste GPCRs
- Application: Use the pipeline to probe structure–function relationships in TAS1R2 (e.g., VFTD vs CRD targeting) and to design mutational scans for mechanism dissection.
- Tools/workflows: Open codebases for RFdiffusion/ProteinMPNN/Boltz-1; Uni-GBSA scoring; Jupyter-based analysis; CLD statistical visualization.
- Assumptions/dependencies: Availability of MSA data; model generalization across GPCR conformational states.
- Academia/Industry (sensory science): Hypothesis-driven experimental validation of Binder1/Binder2
- Application: Express, purify, and test the two top in silico candidates for receptor binding and sweetness (cell-based assays, human panels).
- Tools/workflows: Yeast or bacterial expression (Komagataella phaffii, E. coli, or Aspergillus niger); receptor activation assays; standardized sensory protocols.
- Assumptions/dependencies: Predicted binding energy (Binder1: very strong; Binder2: brazzein-like plausibility) translates into perceptible sweetness without off-flavors.
- Software (bioinformatics/tooling): Packaging the sweet-protein design pipeline as a product
- Application: Deliver a SaaS or on-prem toolkit that automates target setup, hotspot conditioning, sequence design, structure filtering, and energy ranking for TAS1R2 binders.
- Tools/workflows: API endpoints for job submission; dashboard with ipTM/pLDDT/GBSA rankings; batch generation and result tracking.
- Assumptions/dependencies: Licensing compatibility of open-source components; compute cost and turnaround times; user-friendly target setup.
- Academia/Industry (method transfer): Extension to related taste receptors and flavor modulators
- Application: Apply the same pipeline to the umami receptor (TAS1R1/TAS1R3) and explore bitter blockers or taste modulators by retargeting hot-spot conditioning.
- Tools/workflows: Receptor structure preparation for other GPCRs; binder length/architecture tuning; multi-target screening.
- Assumptions/dependencies: Availability and accuracy of target structures; appropriate hotspot definitions; different activation mechanisms across receptors.
- Policy/Health (public health nutrition): Technical evidence to support sugar-reduction initiatives
- Application: Use the paper’s framework as justification for investment in protein-based sweeteners as low-calorie sugar alternatives in reformulation programs.
- Tools/workflows: Policy briefs summarizing technical feasibility; R&D grant calls; public–private partnerships.
- Assumptions/dependencies: Demonstrated safety and consumer acceptance; cost-effective manufacturing; alignment with dietary guidelines.
- Education (graduate training, workforce development): Curriculum modules on de novo protein design for food applications
- Application: Integrate this pipeline and case study into courses/workshops on computational protein design, GPCR biology, and food ingredient innovation.
- Tools/workflows: Hands-on labs using RFdiffusion/ProteinMPNN/Boltz-1; capstone projects targeting TAS1R2.
- Assumptions/dependencies: Access to compute resources; educational licenses; faculty expertise.
Long-Term Applications
- Industry (food & beverage): Commercialization of de novo sweet proteins
- Application: Launch ingredient products based on optimized successors of Binder1/Binder2 that match or exceed brazzein’s sweetness with improved stability, flavor profile, and cost-of-goods.
- Tools/workflows: Directed evolution and stability engineering; microencapsulation/formulation; multi-sweetener blending (with stevia/mogrosides) to reduce off-notes.
- Assumptions/dependencies: Robust expression and purification at scale; consistent sensory performance across matrices; shelf-life and processing compatibility.
- Manufacturing/Biotech (fermentation): Strain and process development for high-yield production
- Application: Engineer Komagataella phaffii, E. coli, or Aspergillus niger strains to produce designed sweet proteins efficiently and economically.
- Tools/workflows: Codon optimization; secretion tags; bioreactor optimization; downstream purification and drying; quality control (LC-MS, stability assays).
- Assumptions/dependencies: Protein folding and post-translational modifications compatible with host; yield and purity meet cost targets; IP freedom-to-operate.
- Healthcare/Public health: Population-level sugar reduction and metabolic benefits
- Application: Incorporation of protein sweeteners in widely consumed products to reduce added sugar intake and support obesity/diabetes prevention strategies.
- Tools/workflows: Reformulation roadmaps with sensory equivalence targets; monitoring of health outcomes; equitable access strategies.
- Assumptions/dependencies: Long-term safety and gut microbiome neutrality; sustained consumer acceptance; competitive pricing vs sugar.
- Pharmaceuticals/Consumer health: Taste masking for oral medicines and functional foods
- Application: Use designed sweet proteins as taste modulators to improve palatability of bitter drugs, nutraceuticals, and pediatric formulations.
- Tools/workflows: Co-formulation with actives; receptor-targeted modulators for bitterness suppression; sensory validation in target populations.
- Assumptions/dependencies: Specificity to sweetness without unintended taste alterations; stability in pharmaceutical excipients; regulatory approval pathways.
- Software/Automation (R&D acceleration): Closed-loop design–build–test–learn platforms
- Application: Integrate the computational pipeline with lab robotics and high-throughput microfermentation to iterate binder design against sensory and binding assays.
- Tools/workflows: Automated cloning, expression, and screening; ML models retrained on experimental feedback; multi-objective optimization (sweetness, stability, cost).
- Assumptions/dependencies: Reliable high-throughput assays that correlate with consumer perception; data infrastructure; budget for automation.
- Personalized nutrition/Consumer tech: Preference-tuned sweeteners
- Application: Tailor binder variants to individual taste sensitivities or cultural preferences, potentially via regional product lines or personalized recommendations.
- Tools/workflows: Taste profiling; A/B testing at scale; digital twin models of receptor variants; adaptive formulation in smart kitchens or vending systems.
- Assumptions/dependencies: Genetic and phenotypic variability in taste receptors; economic feasibility of customization; privacy and data governance.
- Regulatory/Policy: New GRAS determinations and labeling frameworks for de novo proteins
- Application: Develop pathways for safety evaluation, allergenicity assessment, and labeling of computationally designed food proteins.
- Tools/workflows: Toxicology and allergenicity testing (in vitro/in vivo); exposure modeling; harmonized international guidelines; stakeholder consultations.
- Assumptions/dependencies: Transparent risk assessment; harmonization across jurisdictions; consumer trust.
- Finance/Market strategy: IP portfolios and licensing ecosystems for designed sweet proteins
- Application: Create IP around designs, workflows, and production methods; license to ingredient companies; build startup ventures focusing on protein sweeteners.
- Tools/workflows: Patent filings (designs, methods, formulations); freedom-to-operate analyses; strategic partnerships.
- Assumptions/dependencies: Distinctiveness from natural sweet proteins and existing patents; capital availability; market adoption timelines.
- Cross-domain science: GPCR-targeted protein binders beyond taste
- Application: Leverage the pipeline to design protein binders that modulate other class C GPCRs (e.g., neurological targets), expanding therapeutic or sensory applications.
- Tools/workflows: Target-specific hotspot conditioning; functional assays; safety profiling.
- Assumptions/dependencies: High-quality structures of target GPCRs; translational relevance; regulatory complexity.
Notes on feasibility across applications:
- Binding does not guarantee perceptual sweetness; receptor activation, signaling cascades, and matrix effects matter.
- The pipeline shows a helical bias; diversified folds may be needed for broader functional landscapes.
- In-mouth proteolysis, heating, and pH variations require stability engineering to ensure real-world performance.
- Consumer acceptance depends on taste quality, safety perception, and labeling; early sensory co-design is critical.
- Compute resources and reproducibility of structural predictions are prerequisites for reliable deployment.
Glossary
- AlphaFold2: A deep learning system that predicts protein structures from amino acid sequences. "AlphaFold2- predicted model generated using the UniProt sequence Q8TE23."
- AlphaFold3: The latest AlphaFold model capable of predicting biomolecular interactions and complexes. "Boltz-1 (Wohlwend et al., 2025), an open-source implementation of AlphaFold3 (Abramson et al., 2024)."
- binding free energies: Quantitative measures of the thermodynamic favorability of binding between molecules, often reported in kcal·mol−1. "binding free energies were estimated using the Uni-GBSA workflow (Fig. 11)."
- BindCraft: A generative design framework for creating functional protein binders. "Recent generative frameworks, such as RFdiffusion, ProteinMPNN, and BindCraft, combined with structure-prediction advances exemplified by AlphaFold3 and Boltz, now permit the systematic exploration of de novo protein design with atomic-level precision."
- Boltz-1: An open-source implementation of AlphaFold3 used for structure prediction and complex evaluation. "Designed binder sequences from ProteinMPNN were evaluated using Boltz-1 (Wohlwend et al., 2025), an open-source implementation of AlphaFold3 (Abramson et al., 2024)."
- ChimeraX: A molecular visualization tool for rendering and analyzing 3D structures. "For visualization, both the designed binders and the final binder-receptor complexes were rendered in ChimeraX 1.7 (Meng et al., 2023) and Pymol 3.0.0 (DeLano, 2002)."
- compact letter displays (CLDs): A method to summarize statistical group differences by assigning letters to groups that are not significantly different. "Post hoc comparisons were performed using the Games-Howell test, and results were summarized with compact letter displays (CLDs) to indicate groups not significantly different at a = 0.05."
- complex_pde: Complex predicted distance error, a metric indicating uncertainty in predicted inter-residue distances within a complex. "For the complex predicted distance error (complex_pde) (Fig. 8b), brazzein and Binder2 were significantly lower than other binders (p<0.05)."
- confidence_score: A model-derived overall confidence metric for predicted structures or complexes. "The resulting complexes were ranked and filtered based on multiple evaluation metrics, including an overall confidence_score > 0.6, and the average predicted local distance difference test score of the complex (complex_pLDDT >0.7)."
- contact-bias residues: Specific target residues provided to design algorithms to bias binder placement toward desired interfaces. "provided to RFdiffusion as contact-bias residues to guide binder placement."
- Cysteine-Rich Domain (CRD): An extracellular modular domain rich in cysteines, often mediating interactions in class C GPCRs. "The CRD region has also been reported in mediating interactions with sweet proteins such as brazzein (Assadi-Porter et al., 2010)."
- cryo-EM: Cryogenic electron microscopy, a technique for determining high-resolution structures of biomolecules at low temperatures. "Using the recently resolved cryo-EM structure of the TAS1R2 receptor, protein binders were designed to target both the Venus Flytrap Domain and the cysteine-rich domain of TAS1R2."
- de novo protein design: The computational creation of novel protein sequences and structures from scratch. "Recent advances in structural biology and de novo protein design provide new opportunities to overcome these limitations through rational engineering."
- diffusion-based backbone generation: A generative modeling approach that samples protein backbones using diffusion processes. "The workflow combines diffusion-based backbone generation (RFdiffusion), neural network-guided sequence design (ProteinMPNN), structure-based filtering using Boltz-1, and binding energy evaluation via MM/GBSA calculations."
- dl_binder_design: A protocol/toolchain for deep-learning-guided binder sequence design. "Following backbone generation, candidate binders were subjected to sequence design using the ProteinMPNN-FastRelax workflow implemented in the dl_binder_design protocol (https://github.com/nrbennet/dl binder_design) (Bennett et al., 2023; Dauparas et al., 2022)."
- docking orientations: Relative positions and rotations of a ligand or binder when bound to a target, as predicted by modeling. "This suggests that Binder2 has stable docking orientations."
- FastRelax: A PyRosetta protocol that performs energy minimization and relaxation of protein structures. "followed by structural relaxation with PyRosetta FastRelax (Chaudhury, Lyskov, & Gray, 2010)."
- Games-Howell test: A statistical post hoc test suitable for unequal variances and sample sizes. "Post hoc comparisons were performed using the Games-Howell test, and results were summarized with compact letter displays (CLDs) to indicate groups not significantly different at a = 0.05."
- G protein-coupled receptor (GPCR): A large family of membrane receptors that transduce signals via G proteins. "Sweetness perception is mediated primarily by the human sweet taste receptor, a heterodimeric G protein-coupled receptor (GPCR) composed of the TAS1R2 and TAS1R3 subunits, which belong to the class C family of GPCRs."
- heterodimeric: Composed of two different subunits forming a functional dimer. "Sweetness perception is mediated primarily by the human sweet taste receptor, a heterodimeric G protein-coupled receptor (GPCR) composed of the TAS1R2 and TAS1R3 subunits, which belong to the class C family of GPCRs."
- Hot-spot conditioning: A design strategy that biases binder placement toward predefined critical residues or regions on a target. "Hot-spot conditioning was applied for binder design under two schemes."
- Hydrogen-bond assignments: Determinations of hydrogen bond donors/acceptors in a structure, often optimized during preparation. "Hydrogen-bond assignments were automatically optimized, and restrained minimizations were performed, optionally deleting water molecules to improve the quality of the structural models prior to energy evaluation."
- interfacial interactions: Contacts and forces at the interface between two binding partners (e.g., protein–protein). "A zoomed-in view highlights the interfacial interactions: residues ARG37 and THR55 in Binder2 (yellow sticks) form close contacts with residues GLY509 and ALA228 (cyan sticks) within the CRD of TAS1R2."
- ipTM: Inter-protein TM-score-like metric used to assess interface quality and multimer confidence in predicted complexes. "For the ipTM (Fig. 8a), brazzein and Binder2 exhibited the highest values between 0.5 and 0.6, significantly outperforming other designed binders (Binder1, Binder3, Binder4; p<0.05)."
- ligand: A molecule that binds to a specific site on a target protein, often triggering conformational changes. "A recent study showed that upon ligand binding, conformational rearrangements were predominantly observed in the VFTD of TAS1R2, leading to a closure of the orthosteric pocket, whereas the VFTD of TAS1R3 remained largely unchanged (Shi et al., 2025)."
- MM/GB(PB)SA: End-state free energy calculation methods combining molecular mechanics with Generalized Born or Poisson–Boltzmann solvent models. "Uni-GBSA: an open-source and web-based automatic workflow to perform MM/GB (PB) SA calculations for virtual screening."
- multiple sequence alignment (MSA): An alignment of three or more biological sequences used to infer evolutionary and structural information. "For each binder-receptor complex, structural predictions were generated with the -- use_msa_server option enabled to leverage multiple sequence alignment information."
- orthosteric pocket: The primary ligand-binding site on a receptor responsible for endogenous ligand recognition. "leading to a closure of the orthosteric pocket, whereas the VFTD of TAS1R3 remained largely unchanged (Shi et al., 2025)."
- PDB ID: The unique identifier for a structure in the Protein Data Bank. "PDB ID: 9NOR, chain B"
- pLDDT: Predicted Local Distance Difference Test score, indicating per-residue confidence in structure predictions. "The complex pLDDT scores (Fig. 8c) were consistently high (>0.78) across all complexes, confirming good overall model confidence."
- Pymol: A molecular graphics system used for visualizing and manipulating 3D biomolecular structures. "For visualization, both the designed binders and the final binder-receptor complexes were rendered in ChimeraX 1.7 (Meng et al., 2023) and Pymol 3.0.0 (DeLano, 2002)."
- ProteinMPNN: A neural network model that designs protein sequences compatible with given backbones and interfaces. "Sequence design was carried out with ProteinMPNN, which maps the generated backbones to amino acid sequences that are physically realistic and compatible with the targeted receptor interface (Dauparas et al., 2022)."
- PyRosetta: A Python interface to the Rosetta molecular modeling suite used for structure refinement and energy calculations. "followed by structural relaxation with PyRosetta FastRelax (Chaudhury, Lyskov, & Gray, 2010)."
- restrained minimizations: Energy minimization procedures applying positional or geometric restraints to maintain structural features. "Hydrogen-bond assignments were automatically optimized, and restrained minimizations were performed, optionally deleting water molecules to improve the quality of the structural models prior to energy evaluation."
- RFdiffusion: A diffusion-based generative model for designing protein backbones and interfaces. "Binder backbones were generated with the RFdiffusion binder design protocol (https://github.com/RosettaCommons/RFdiffusion) (Watson et al., 2023)."
- Schrödinger: A commercial computational chemistry platform used for structure preparation and modeling. "The filtered binder-receptor complex structures were further processed using Schrödinger 2023.4 (Madhavi Sastry, Adzhigirey, Day, Annabhimoju, & Sherman, 2013)."
- saturation transfer difference NMR spectroscopy: A nuclear magnetic resonance technique to detect ligand binding by transferring saturation from receptor to bound ligand. "Interactions between the human sweet-sensing T1R2-T1R3 receptor and sweeteners detected by saturation transfer difference NMR spectroscopy."
- seven-transmembrane (7TM) region: The heptahelical membrane-spanning domain characteristic of many GPCRs. "The sweet taste receptor subunit TAS1R2 belongs to the class C family of GPCRs and has three modular domains: the extracellular Venus Flytrap Domain (VFTD), a Cysteine-Rich Domain (CRD), and the seven-transmembrane (7TM) region (Fig. 7a)."
- structural filtering: The process of evaluating and discarding predicted complexes based on confidence and quality metrics. "In the third stage, structural filtering was performed using Boltz-1 (Wohlwend et al., 2025), a structure prediction model that evaluates binder-receptor interactions and eliminates unstable or misfolded candidates."
- structural scaffolds: Backbone frameworks that support a protein’s overall shape and facilitate sequence design. "we generated de novo binder backbone using RFdiffusion, which is a diffusion-based framework that produces diverse structural scaffolds and thereby enables broad sampling of conformational space for candidate binders (Watson et al., 2023)."
- threaded: The process of mapping a designed sequence onto a specific backbone conformation. "The newly designed sequence was then threaded back onto the backbone within the complex, followed by structural relaxation with PyRosetta FastRelax (Chaudhury, Lyskov, & Gray, 2010)."
- transmembrane domain (TMD): The membrane-embedded portion of a receptor comprising its transmembrane helices. "Each subunit contains an extracellular Venus Flytrap Domain (VFTD), a cysteine-rich domain (CRD), and a transmembrane domain (TMD)."
- Uni-GBSA: An automated workflow for performing MM/GB(PB)SA binding energy calculations. "binding free energies were estimated using the Uni-GBSA workflow (Yang, Bo, Xu, Xu, Wang, & Zheng, 2023), an automated pipeline for MM/GB(PB)SA calculations."
- UniProtKB: A comprehensive protein sequence and annotation database. "The amino acid sequence of the human taste receptor type 1 member 2 (TAS1R2) was obtained from UniProtKB (accession number Q8TE23) (Consortium, 2019), comprising 839 residues."
- Venus Flytrap Domain (VFTD): The extracellular ligand-binding domain of class C GPCRs that can adopt open/closed conformations. "Each subunit contains an extracellular Venus Flytrap Domain (VFTD), a cysteine-rich domain (CRD), and a transmembrane domain (TMD)."
- Welch's ANOVA: A robust analysis of variance method for comparing group means when variances and sample sizes are unequal. "Group differences were first assessed with Welch's ANOVA, which accommodates unequal variances and sample sizes."
Collections
Sign up for free to add this paper to one or more collections.