- The paper presents a universal harmonization module that mitigates domain shifts through a two-stage feature normalization and restoration cascade.
- It employs a domain-gated head for dataset-free inference, achieving up to +8.4% DSC improvement across diverse brain lesion datasets.
- Ablation studies confirm that integrating instance-wise standardization with polynomial affine transformations is critical for robust performance in joint training.
Universal Harmonization for Joint Medical Image Segmentation
Medical image segmentation in the clinical domain is fundamentally challenged by the pervasive heterogeneity in available datasets. Key sources of variation include imaging modalities, acquisition protocols, and annotation standards across different institutions. Existing deep learning segmentation methods—despite leveraging joint training (JT) paradigms—commonly fail to reconcile such heterogeneity, resulting in brittle models that are highly sensitive to domain and modality shifts, especially when leveraging architectures founded on layer normalization (e.g., transformers). These domain shifts degrade performance and impede generalization. The problem is exacerbated by divergent class definitions and unaligned label spaces across datasets, often requiring labor-intensive dataset-specific models or inference-time identification routines.
U-Harmony Framework and Methodological Innovations
The paper introduces U-Harmony, a universal harmonization module that integrates with generic 3D medical segmentation architectures, both convolutional and transformer-based, to enable robust joint learning on highly heterogeneous datasets. Its key technical contributions are twofold: (1) a two-stage feature normalization and restoration mechanism, and (2) a domain-gated head for dataset-free inference.
The harmonization pipeline replaces conventional normalization layers by first performing instance-wise feature standardization, followed by a polynomially-parameterized affine transformation, and finally, a restoration stage that re-injects critical domain-specific statistical information. This architecture ensures feature distributions are regularized to a common reference, thus mitigating domain shifts, but are later adapted to retain information unique to each dataset (Figure 1).
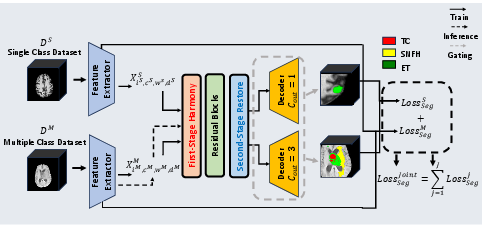
Figure 1: Overview of the U-Harmony framework illustrating the harmonization-restoration cascade and domain-gated head for instance-agnostic multi-domain inference.
The domain-gated head employs domain prototypes and dynamically infers the appropriate domain contribution at test time without side-channel metadata or explicit dataset identification, supporting seamless handling of data with previously unseen or ambiguous origins.
Experimental Setup and Comparisons
U-Harmony was evaluated across three highly heterogeneous, public brain lesion datasets: UCSF-BMSR, BrainMetShare, and BraTS-METS 2023. These datasets vary in MRI modalities, annotation schemes, and anatomical focus. Models were assessed in both single-dataset and joint-training regimes, including settings with unaligned modality and class compositions.
Baseline comparisons covered a spectrum of modern segmentation models (i.e., nn-UNet, SwinUNETR, 3D UNet, V-Net, nnFormer, TransUNet) and methods tailored to handle multi-domain or expert-ensemble scenarios (e.g., CVCL, MoME, MultiTalent).
Results: Robust Joint Training Across Domains
Superiority Under Domain Shift:
U-Harmony demonstrates consistent improvements in Dice Similarity Coefficient (DSC) across all tested domains, surpassing both CNN and transformer baselines by 1.6–3.4 percentage points in average DSC. Particularly, when applied within the SwinUNETR framework, the full U-Harmony model achieves significant performance gains on both single and joint segmentation settings.
Generalization in Joint Settings:
Under joint training with unaligned modalities and annotation classes, U-Harmony is able to maintain source domain performance while generalizing effectively to target domains. Its performance exceeds that of competing methods, with up to +8.4% improvement in DSC relative to multi-domain baselines. Visually, this manifests as superior boundary delineation and adaptive segmentation, even under severe domain and class shifts (Figure 2).
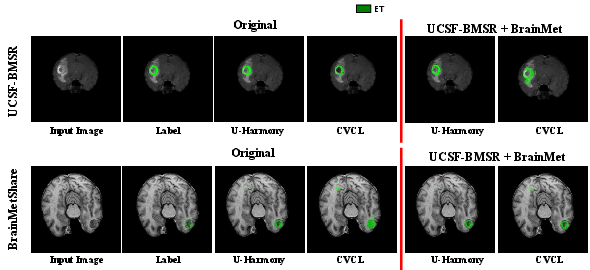
Figure 2: Comparative segmentation visualization on single and joint-task transitions, highlighting U-Harmony’s superior boundary accuracy and adaptivity versus CVCL.
Ablation and Layer Utilization Analysis:
Ablation studies confirm the contribution of the harmonization-restoration cascade: first-stage harmonization alone is insufficient; the restoration step is critical for maintaining domain-relevant features. Incorporating higher-order polynomial terms in the affine transformation further boosts robustness. Increased stacking of U-Harmony layers correlates with consistently superior segmentation performance, indicating the capacity for deeper harmonization to capture complex domain interrelations (Figure 3).
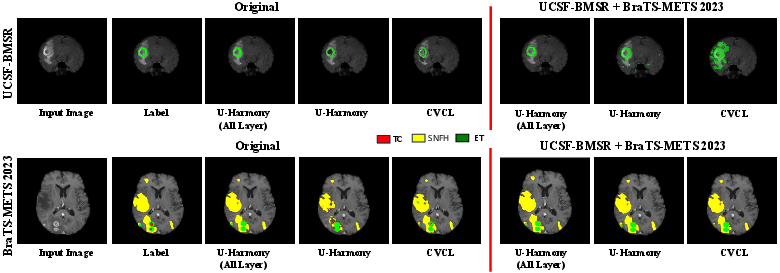
Figure 3: Segmentation results with ablations on harmonization, showing U-Harmony’s robustness under domain shift and expansion to extra (out-of-domain) classes.
Dataset-Free Inference:
The domain-gated head enables accurate inference without requiring prior knowledge of test data provenance, lagging behind an oracle (with explicit dataset labels) by only 1.3% DSC, but offering dramatically improved practicality for real-world deployment scenarios.
Implications and Future Directions
The U-Harmony module re-defines the practical limits of joint training for 3D medical image segmentation. The framework makes several strong claims: it systematically mitigates detrimental domain and modality shifts, requires no inference-time dataset identification, and enables a single segmentation model to function across diverse clinical data sources without sacrificing source-domain performance or incurring performance/robustness trade-offs typical to prior DG methods.
Practically, these advances support scalable deployment of medical AI in distributed and federated clinical networks, where pooling and harmonizing data from multiple sources is essential. Theoretically, the success of the harmonization-restoration cascade opens new directions for modular, learned normalization schemes addressing other multi-domain, multi-task scenarios, and provides a template for handling adaptation to out-of-training-distribution data—critical for trustworthy clinical deployment.
Potential future research includes extending U-Harmony to additional medical domains, leveraging it for foundation models in medical vision, exploring fine-grained class expansion in federated and privacy-preserving regimes, and integrating advanced meta-learning or continual learning strategies that further minimize catastrophic forgetting during sequential dataset expansion.
Conclusion
U-Harmony provides a unified, modular approach for universal harmonization in 3D medical segmentation, achieving robust joint training across heterogeneous, real-world datasets while eliminating the need for explicit domain identification at inference. The harmonization-restoration pipeline sets a new benchmark for adaptable segmentation, with strong evidence for both practical utility and theoretic extensibility in clinical AI.


