- The paper introduces a framework that integrates a Copy-Paste Mechanism for input augmentation and a Cluster Maximum Mean Discrepancy block for domain alignment.
- It leverages pseudo-domain clustering via HDBSCAN with an MMD objective to align features, boosting segmentation accuracy on Fundus and M&Ms benchmarks.
- Experimental results demonstrate improved Dice and Jaccard indices, effectively bridging mixed-domain gaps without needing explicit domain labels.
Domain-Invariant Mixed-Domain Semi-Supervised Medical Image Segmentation via Clustered Maximum Mean Discrepancy Alignment
Introduction
Semi-supervised learning (SSL) and domain adaptation (DA) are cornerstones for medical image segmentation tasks, yet routine clinical datasets feature both annotation scarcity and complex domain heterogeneity—arising from multi-institutional data collection and various imaging devices. Conventional SSL/DA approaches frequently assume known, single, or clearly labeled domain partitions; this assumption fails under mixed-domain conditions, where domain labels for unlabeled samples are unknown and shifts are nontrivial. "Domain-invariant Mixed-domain Semi-supervised Medical Image Segmentation with Clustered Maximum Mean Discrepancy Alignment" (2601.16954) targets this complex regime by advancing both input-level and representational solutions for mixed-domain segmentation.
Methodology
The proposed framework, illustrated in Figure 1, adopts a teacher–student architecture—leveraging a small labeled set from a known domain and a large unlabeled set composed of mixed, unlabeled domains.
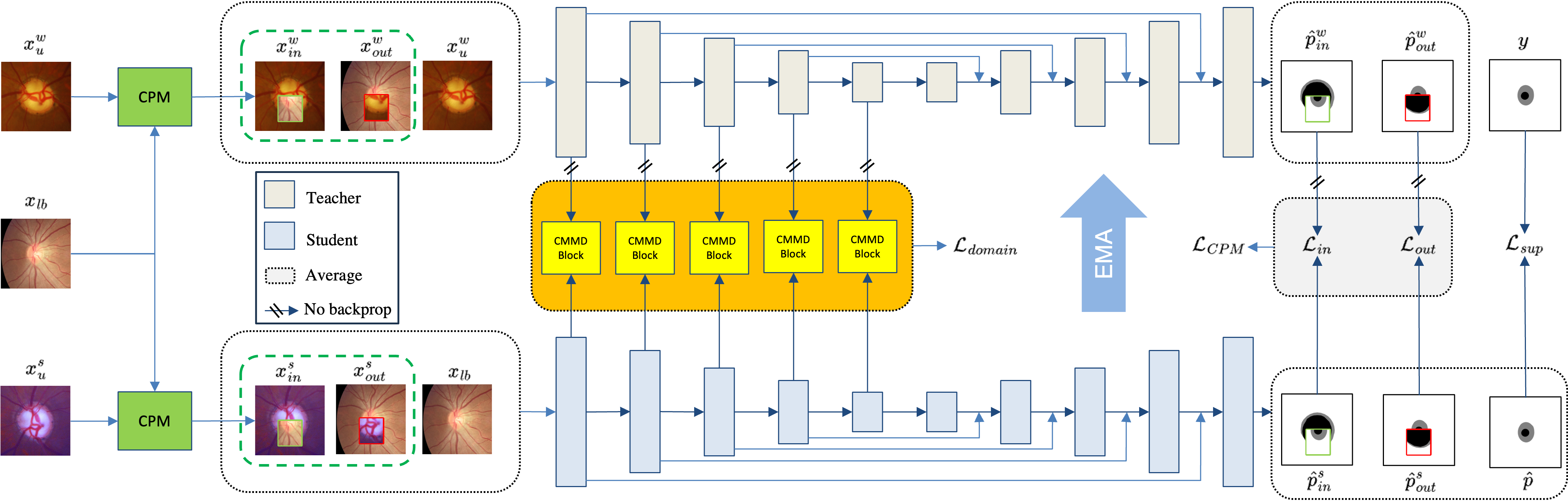
Figure 1: Framework overview integrating cross-domain data augmentation and feature alignment via CPM and CMMD modules within a teacher-student semi-supervised system.
Copy-Paste Mechanism (CPM)
The Copy-Paste Mechanism (CPM) directly augments the training set by spatially exchanging informative regions between labeled and unlabeled samples, promoting synthetic diversity at the pixel level. Within each CPM operation, binary masks delineate the transferred region, creating compositional examples that not only increase the effective size and heterogeneity of training data but also facilitate the bridging of distributional gaps without explicit domain information. Consistency regularization with pseudo-labels (via Dice and cross-entropy losses) ensures supervision across both weak (teacher) and strong (student) augmentations.
Cluster Maximum Mean Discrepancy (CMMD) Block
To explicitly address domain discrepancies at the feature level, the Cluster Maximum Mean Discrepancy (CMMD) block executes two main procedures (see Figure 2):
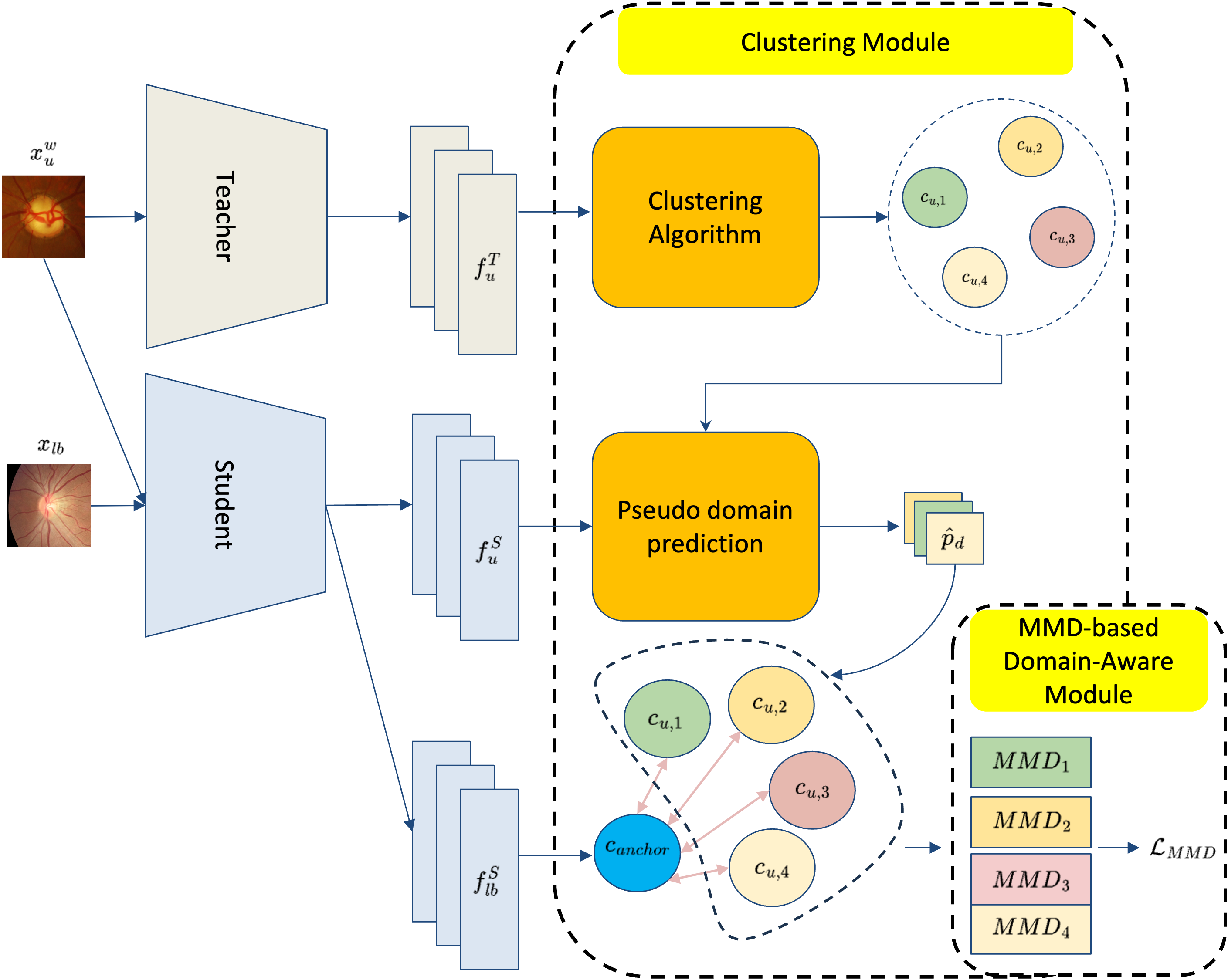
Figure 2: The CMMD block discovers pseudo-domains by density-based clustering and aligns their feature distributions to labeled anchors via the MMD objective.
- Clustering Module: High-level features of the unlabeled set, produced by the teacher encoder, are clustered using HDBSCAN, a density-based method adept at discovering clusters of unknown multiplicity without requiring prior assumptions about the number of domains. The resulting pseudo-centroids label the student features, establishing soft correspondences between pseudo-domains.
- MMD-based Domain-Aware Module: The Maximum Mean Discrepancy (MMD) objective measures and minimizes the distance in RKHS between each pseudo-domain cluster and the "anchor" cluster derived from labeled samples. This block operates across all encoder layers, ensuring that skip connections feeding into the decoder transmit features purged of domain signals, and only class-relevant representations persist. Domain loss aggregates alignment costs across encoder stages, mitigating domain-specific clustering and promoting invariance.
Loss Composition
The total loss is the weighted sum of supervised loss, CPM loss, and domain loss, with the latter two gradually integrated via a dynamic annealing schedule. This staged integration stabilizes early learning while progressively enforcing domain invariance and data diversity.
Experimental Results
Robustness and domain invariance were tested on the Fundus and M&Ms (Multi-centre, Multi-vendor, Multi-disease) segmentation benchmarks. Metrics included Dice coefficient (DC), Jaccard index (JC), Hausdorff distance (HD), and Average Surface Distance (ASD).
Fundus: With only 20 labeled samples, the method achieves DC/JC of 86.99%/78.22%, outperforming state-of-the-art SSL (UA-MT, FixMatch, CPS) and DA approaches (FDA, UDA-VAE++) as well as recent dedicated mixed-domain methods such as MiDSS (reproduced DC/JC: 86.91%/78.12%). This is achieved without reliance on explicit domain indices. Notably, representation visualizations (see Figure 3) reveal reduced domain separation post-CMMD, indicating successful eradication of domain-specific clustering.
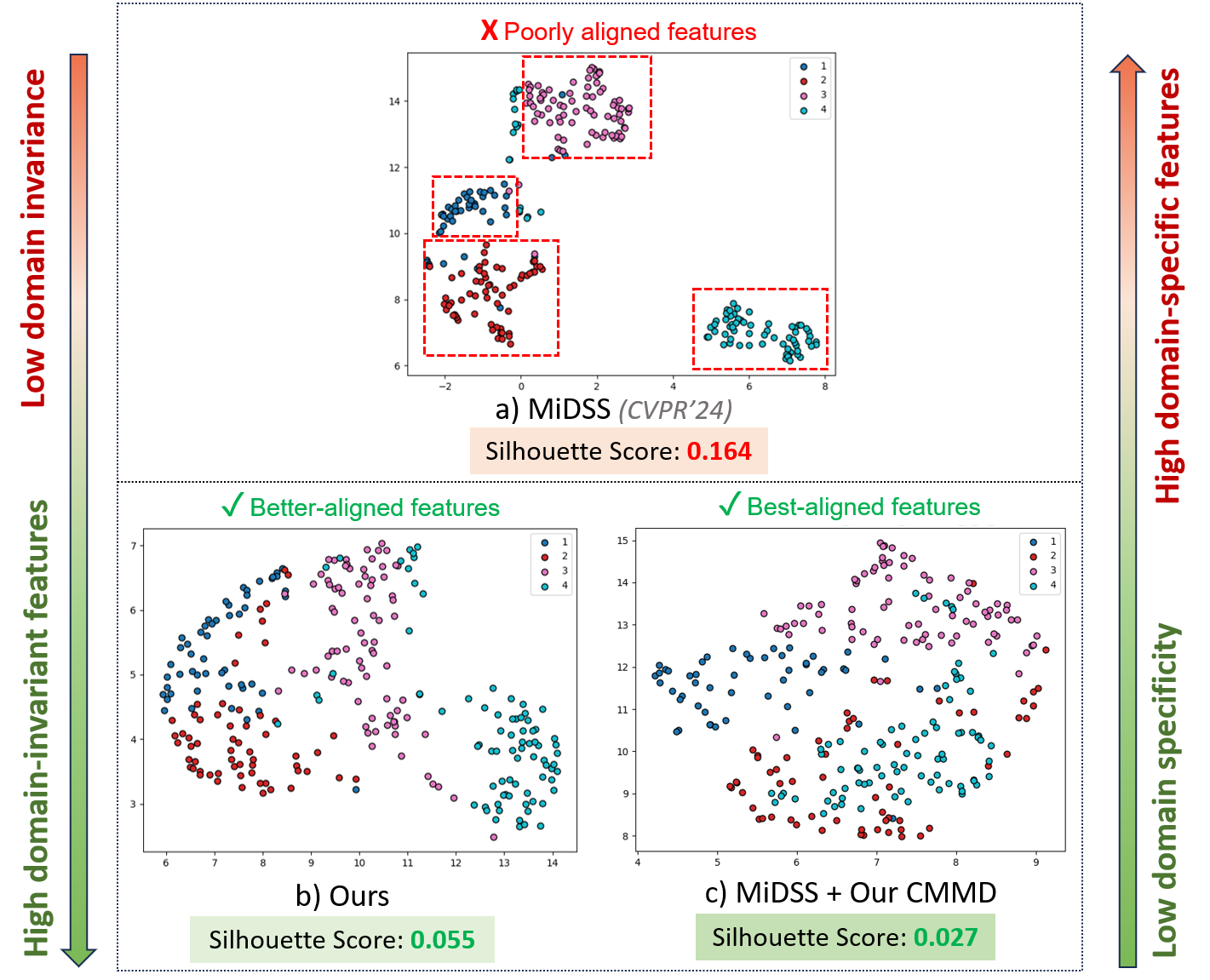
Figure 3: UMAP visualization reveals decreased inter-domain feature variance after applying the proposed CMMD, as evidenced by lower silhouette scores and more mixed clusters.
M&Ms: On this multi-vendor cardiac dataset, the approach sets a new performance upper bound with an average DC of 84.35 and JC of 75.33, with gap-bridging observable especially across scanner boundaries. Incorporating CMMD into the MiDSS pipeline further raises accuracy to DC/JC 87.18%/78.45%, validating CMMD’s plug-and-play property for domain-mixed pipelines.
Ablation Studies delineate the respective impact: CPM alone raises DC by ~3.8 points, CMMD provides a further boost, with presence of both necessary for closing the domain gap.
Findings and Implications
Compared to previous approaches, the following critical claims and outcomes are substantiated:
- CPM is essential for enriching cross-domain training data, mitigating overfitting to limited labels in scenarios with severe domain discrepancy.
- The CMMD block successfully learns domain-invariant features across entirely unknown, mixed domains. Unlike classic DA, which requires explicit label or two-domain scenarios, the clustering-plus-alignment mechanism dispenses with such assumptions.
- Visualization and numerical scores collectively demonstrate the contraction of the domain gap both qualitatively and quantitatively, with the framework surpassing dedicated mixed-domain methods on all core benchmarks.
- CMMD functions as a modular, parameter-free feature-space loss, enabling its seamless integration into other segmentation pipelines.
- Some empirical margin to MiDSS exists, but the consistent improvement is robust across multiple datasets and evaluation metrics, even when annotation is minimal.
Future Directions
The method’s cluster-based domain modeling opens new avenues for instance- and batch-level domain discovery in more complex segmentation pipelines and for modalities with higher data heterogeneity (e.g., histopathology, cross-protocol imaging). Deployment to transformers or non-convolutional architectures remains unexplored. Additionally, adaptive, learnable weighting schedules for loss mixing, and efficient kernel selection for MMD computation, will further enhance its applicability.
Conclusion
This work defines a principled, easily extensible framework for mixed-domain semi-supervised medical image segmentation, combining input-level augmentation with domain-invariant feature alignment. The CPM and CMMD modules, integrated into a teacher–student network, achieve strong performance under severe domain-label scarcity, and the approach generalizes to unknown and multi-modal distribution shifts. This framework forms a solid basis for advancing robust, practical medical imaging solutions in the era of multi-source, annotation-deficient data.


