- The paper introduces the NccDCM framework to construct scalable, dynamic causal brain networks from EEG data using a hierarchical mixed-effects model.
- It leverages truncated Chen-Fliess series and evolutionary optimization (JADE) to achieve reliable estimation of excitation/inhibition ratios and causal couplings.
- The study provides robust group comparisons in epilepsy, demonstrating improved sensitivity over conventional connectivity methods for detecting network disruptions.
Differential Dynamic Causal Nets: Model Construction, Identification, and Group Comparisons
This essay provides a comprehensive technical summary and analysis of "Differential Dynamic Causal Nets: Model Construction, Identification and Group Comparisons" (2601.21478). The work introduces the NccDCM framework, a scalable, data-driven approach for constructing and comparing dynamic causal brain networks derived from EEG with explicit neural mass modeling and hierarchical mixed-effects parameter structure. The methodology addresses challenges of model identifiability, high-dimensional inference across heterogeneous subjects, and efficient estimation in nonlinear stochastic dynamical systems, with concrete applications to epilepsy-related neuropathology.
Methodological Framework: NccDCM Architecture and Inference
The proposed NccDCM constructs a conditionally coupled dynamic causal network at whole-brain scale, where each node corresponds to a neural mass model (Jansen-Rit), parameterized separately per subject and group, and edges encode directed causal transmission with biophysical interpretability.
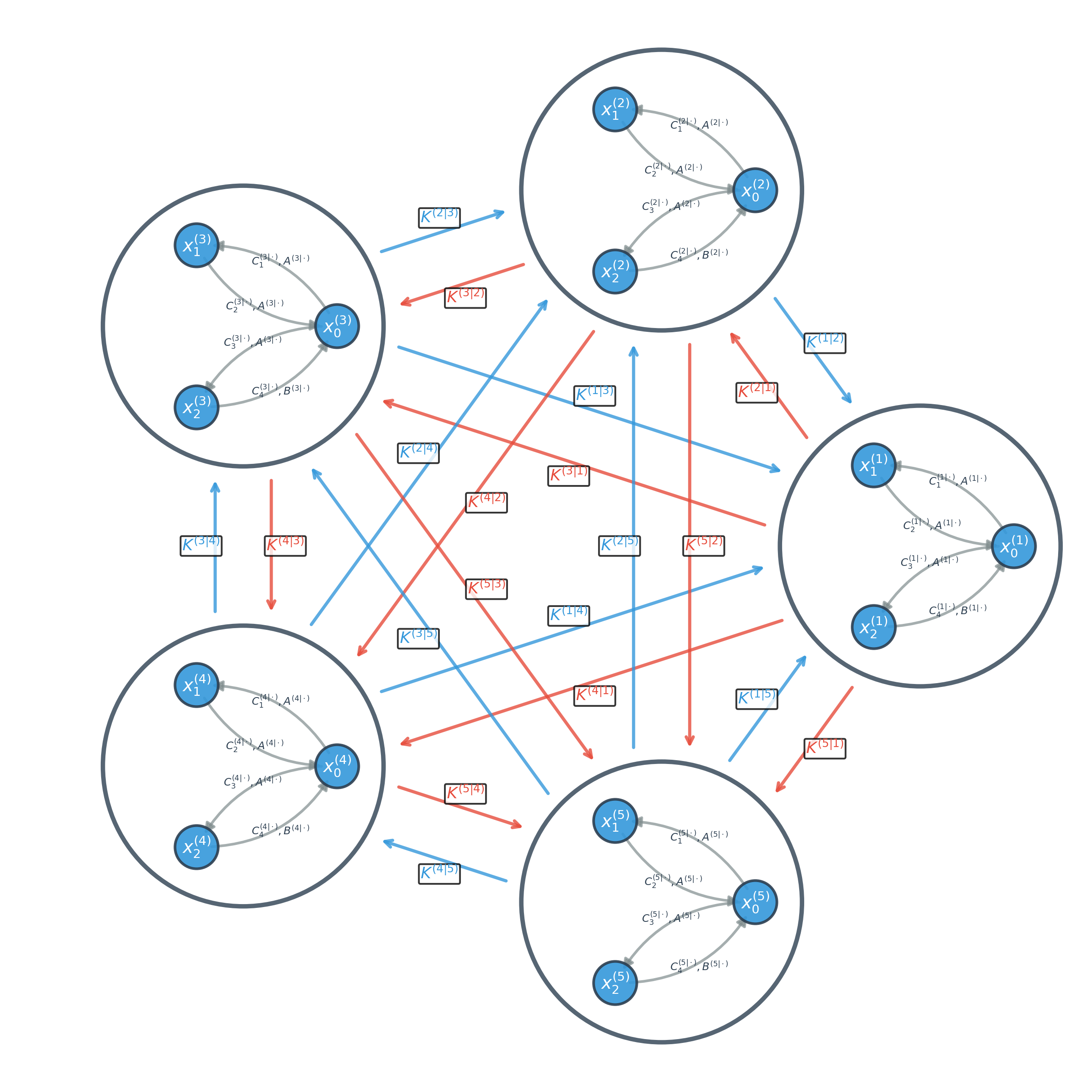
Figure 1: Schematic diagram of a five-channel NccDCM, each node parameterized by a local Jansen-Rit mass model and inter-node couplings denoted by transmission parameter K(i∣j).
The network is hierarchically structured:
- Node parameters (e.g., excitatory/inhibitory gains, synaptic parameters) vary by subject and follow a mixed-effects model.
- Edge parameters represent extrinsic causal couplings, allowing for heterogeneity and group-level effects.
- Within-node dynamics are governed by six-dimensional SDEs per Jansen-Rit formulation.
- Causal inference is achieved by fitting these models directly to observed multichannel EEG, rather than relying on a-priori parameterization or homogeneous assumptions.
Crucially, estimation leverages Chen-Fliess series expansions to construct tractable loss functions (forward predictive errors), enabling data-driven optimization via a genetic algorithm, JADE, tailored for highly non-convex, partially observed dynamical systems.
Statistical Modeling and Inference Procedures
Hierarchical Parameterization and Mixed-Effects
The key statistical advancement lies in the explicit mixed-effects parameterization of both node and edge dynamics, enabling:
- Within-group variability and between-group contrasts (case-control, preictal-ictal).
- Subject-level inference, critical for heterogeneous clinical populations.
- Efficient post-hoc statistical analysis, using MANOVA and permutation-based significance testing for both means and variances of estimated effective connectivity parameters.
Estimation and Identifiability
The forward modeling step, grounded in truncated Chen-Fliess expansions, is followed by evolutionary global optimization (JADE), robust to non-convexities and local minima. The work demonstrates:
- Individual excitatory/inhibitory parameters (A, B) are non-identifiable, but their ratio (A/B)—the excitation/inhibition (E/I) balance—is well identified and highly clinically relevant.
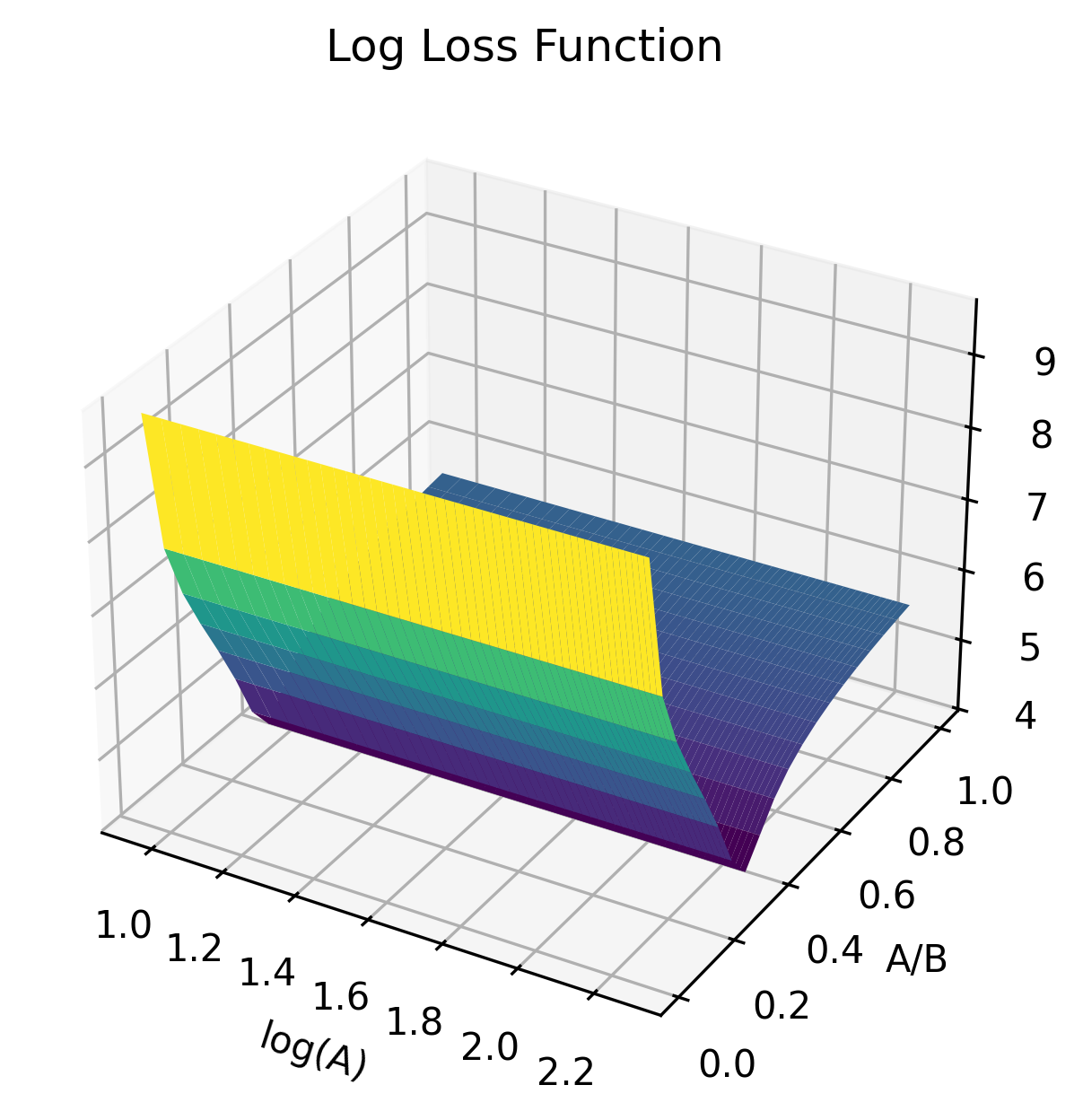
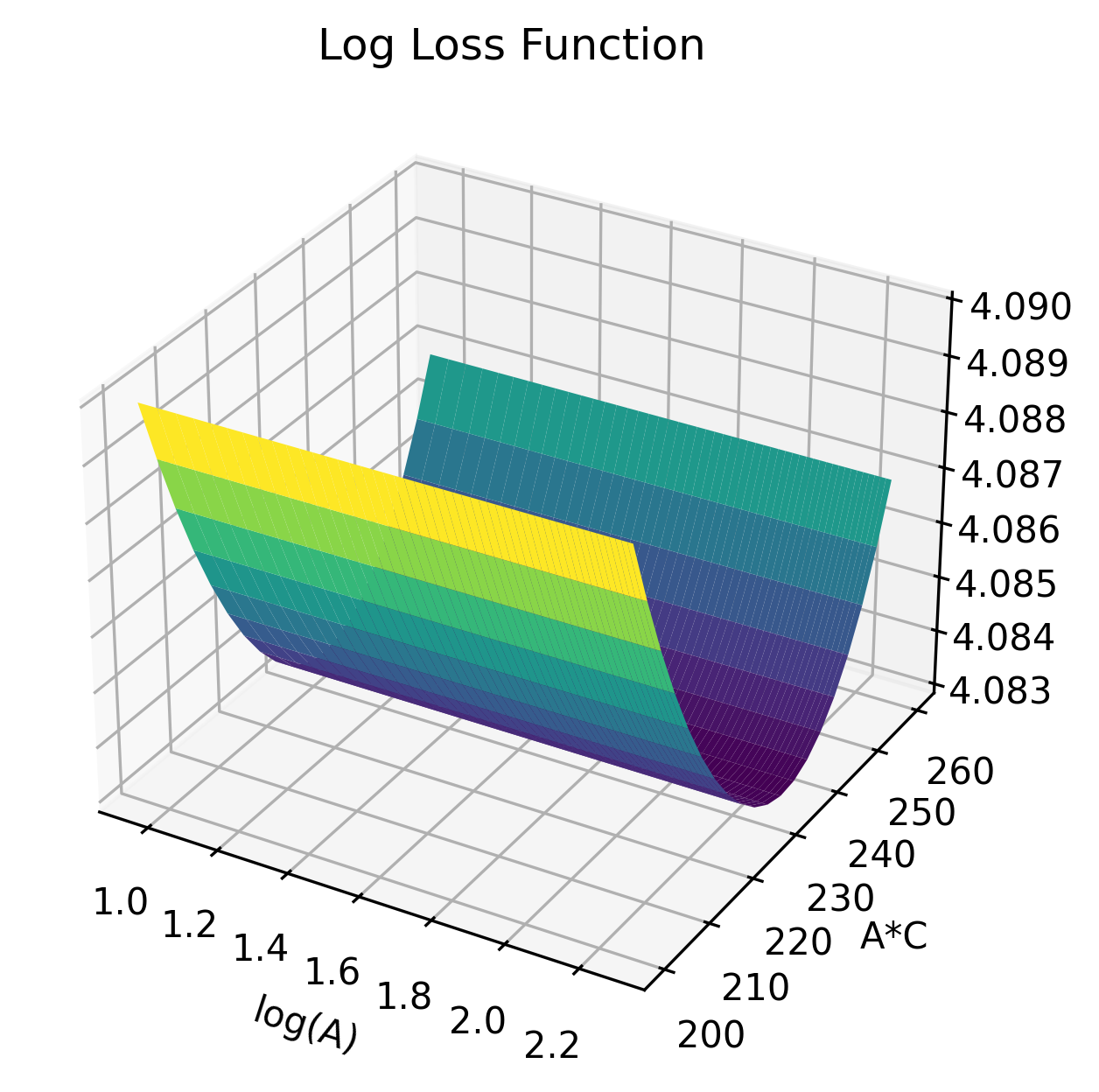
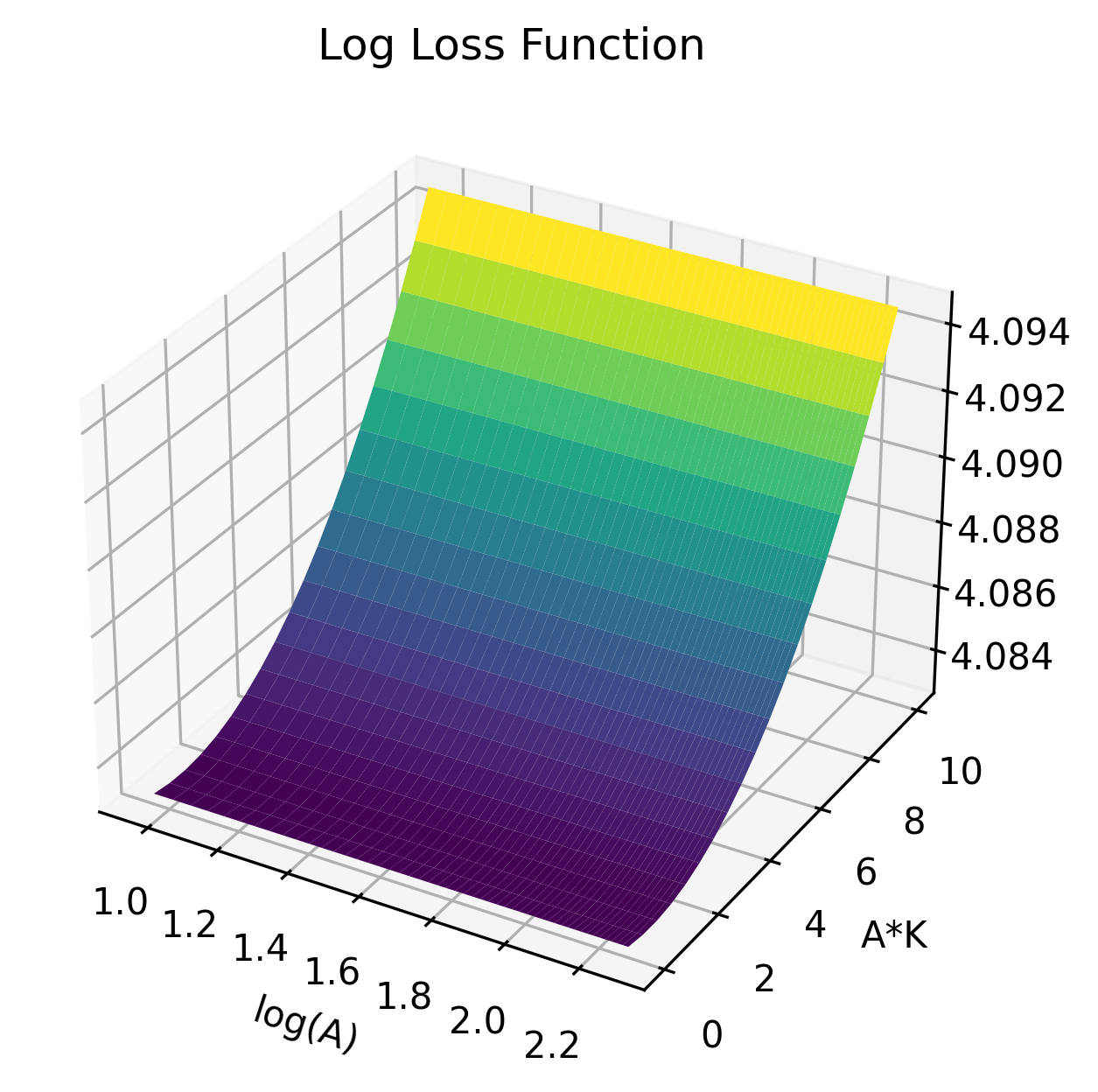
Figure 3: Surface plots of loss functions in (logA,A/B) and other parameter pairs, demonstrating flatness in logA and clear minima along the E/I axis, indicating identifiability structure.
Application to Epileptic Brain Network Analysis
The NccDCM framework is applied to a pediatric epilepsy EEG dataset, where both real and simulated data are parsed into seizure (ictal), pre-seizure (preictal), and control conditions. The analysis pipeline consists of:
- Subject-level heterogeneity assessment via hierarchical clustering on channel-wise features, justifying the mixed-effects model approach.
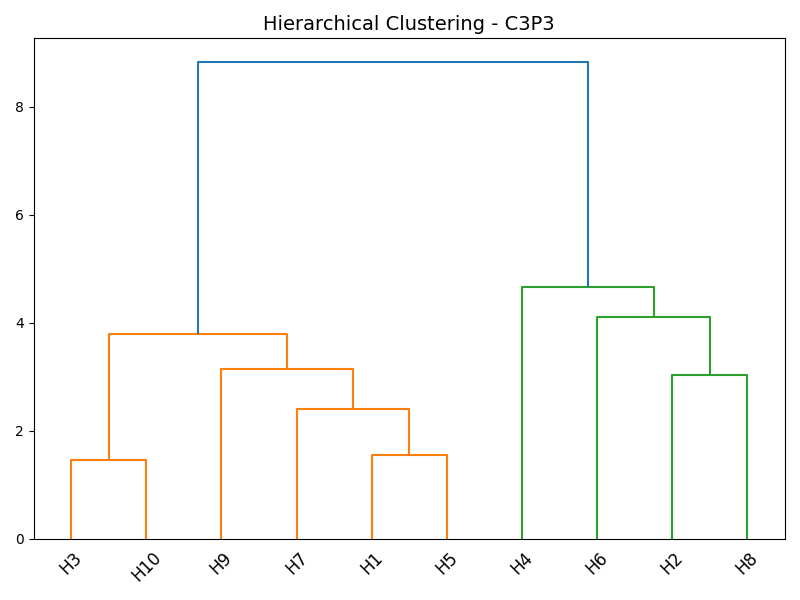
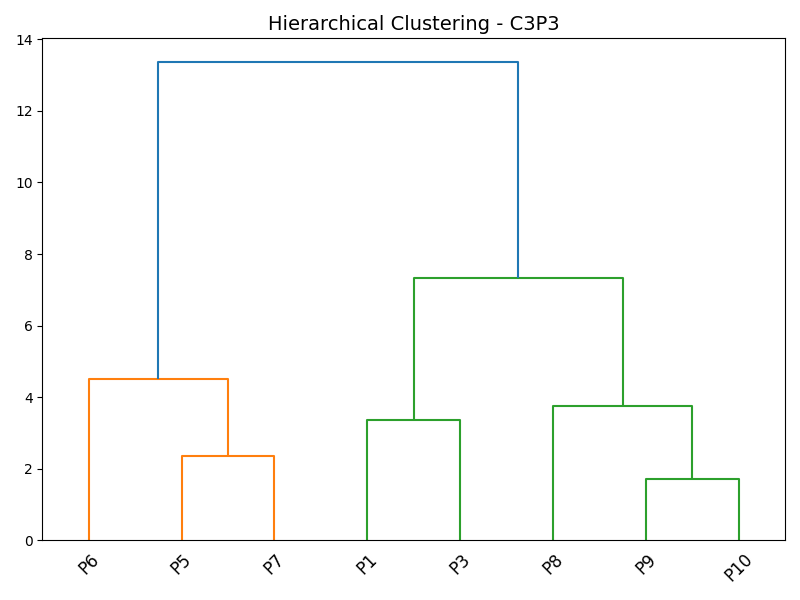
Figure 4: Dendrograms showing clear subject heterogeneity within both control and case groups at the level of the C3P3 channel.
- Channel- and group-specific parameter fitting to estimate individually parameterized networks and perform group comparisons.
- Post-estimation statistical testing—via ANOVA and permutation—to identify significant differential causal subnets in both means and variances of E/I ratio, within-channel coupling, and transmission parameters.
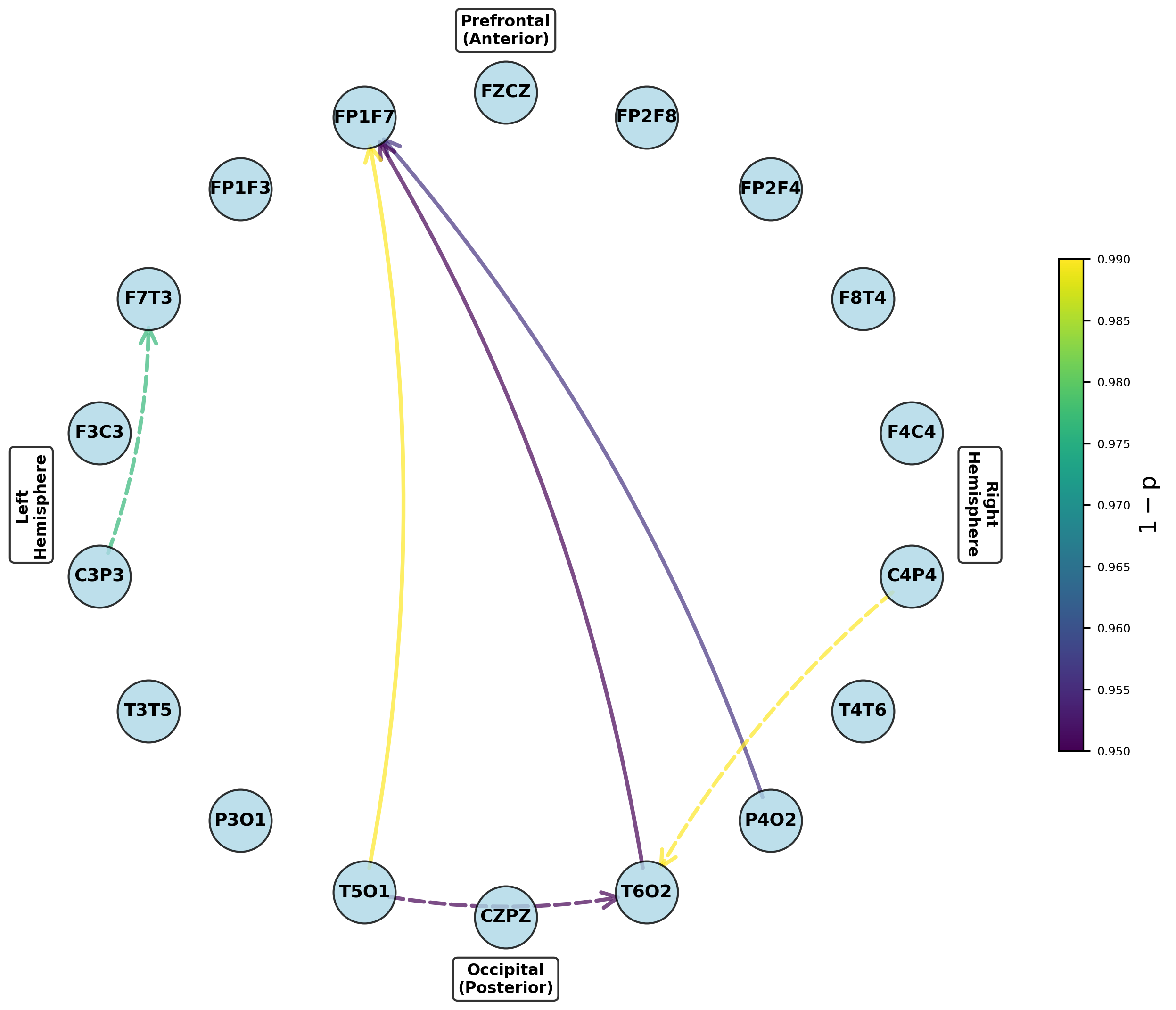
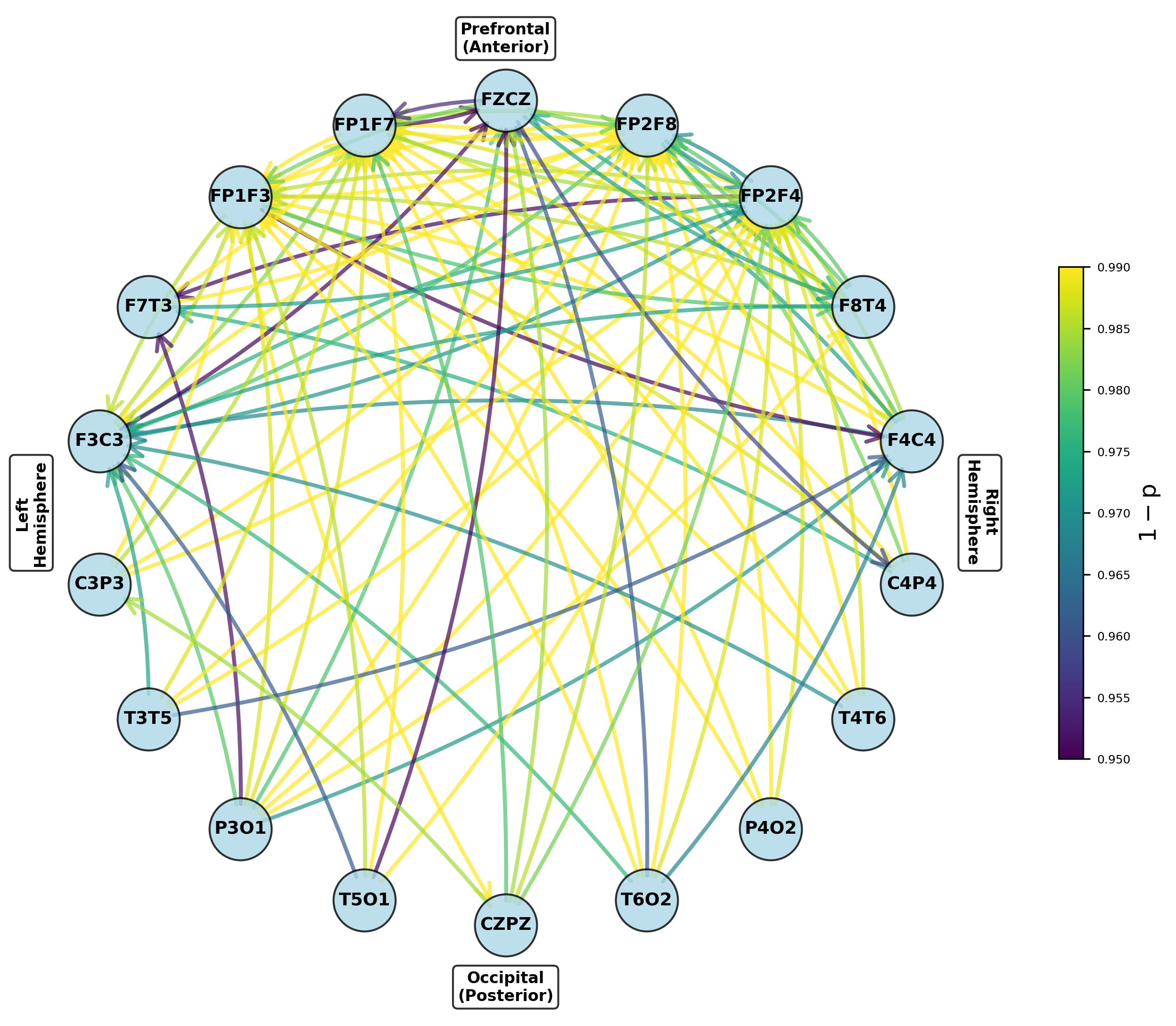
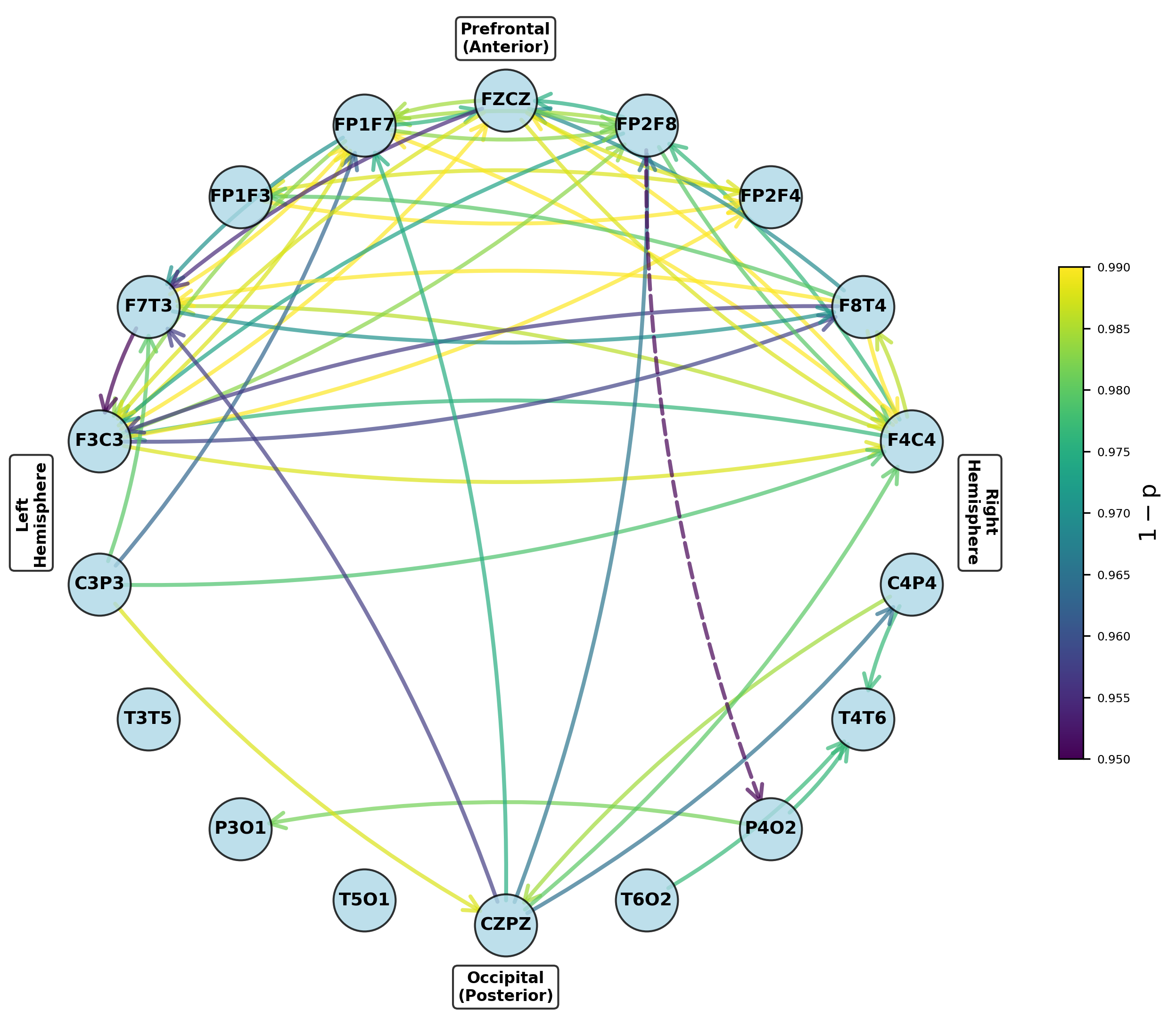
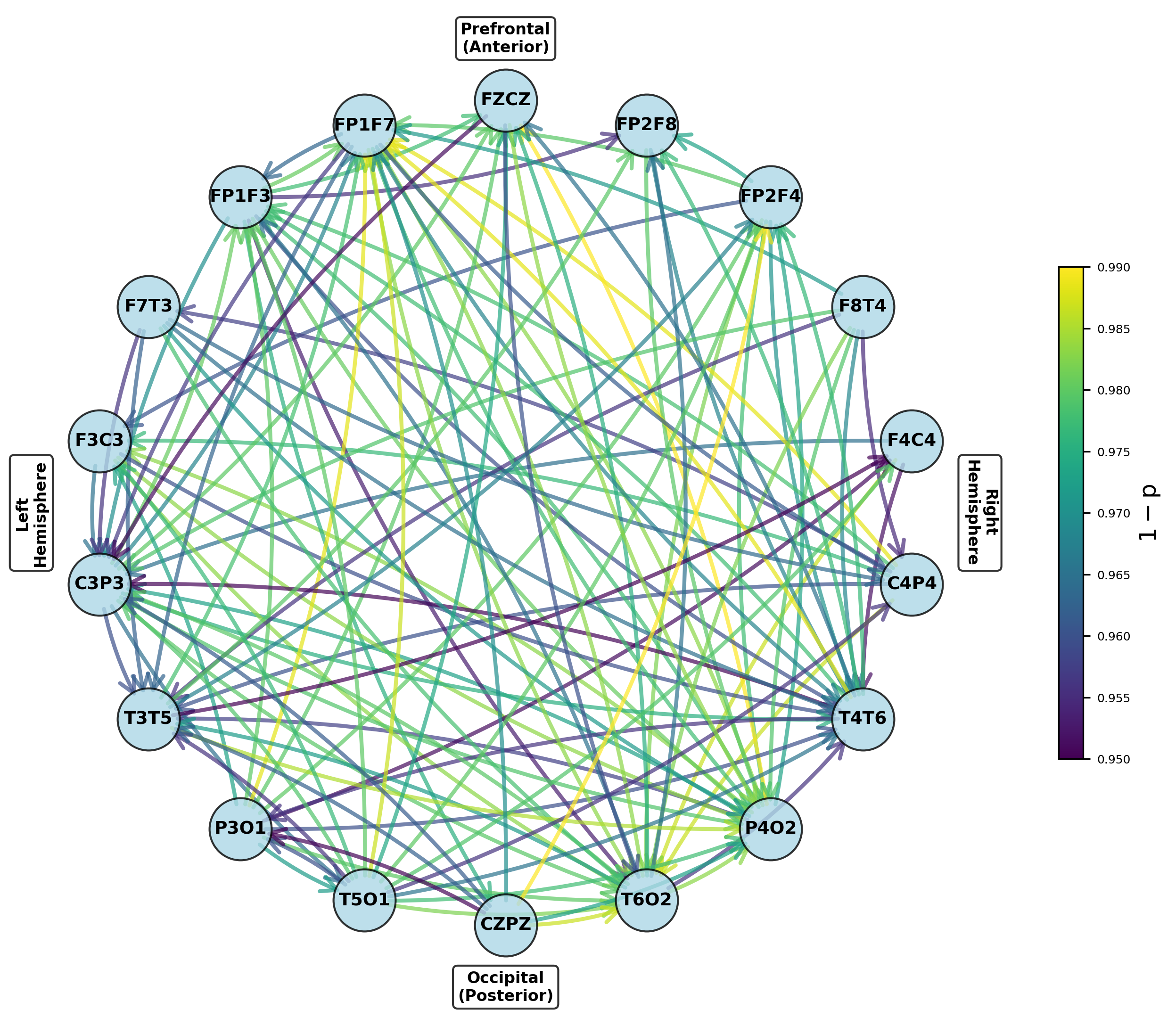
Figure 5: Networks of parameters showing significant group differences in dominant frontal and temporoparietal regions after multiple testing correction.
- Variance-based differential network analysis elucidates group differences in heterogeneity across specific subnetworks.
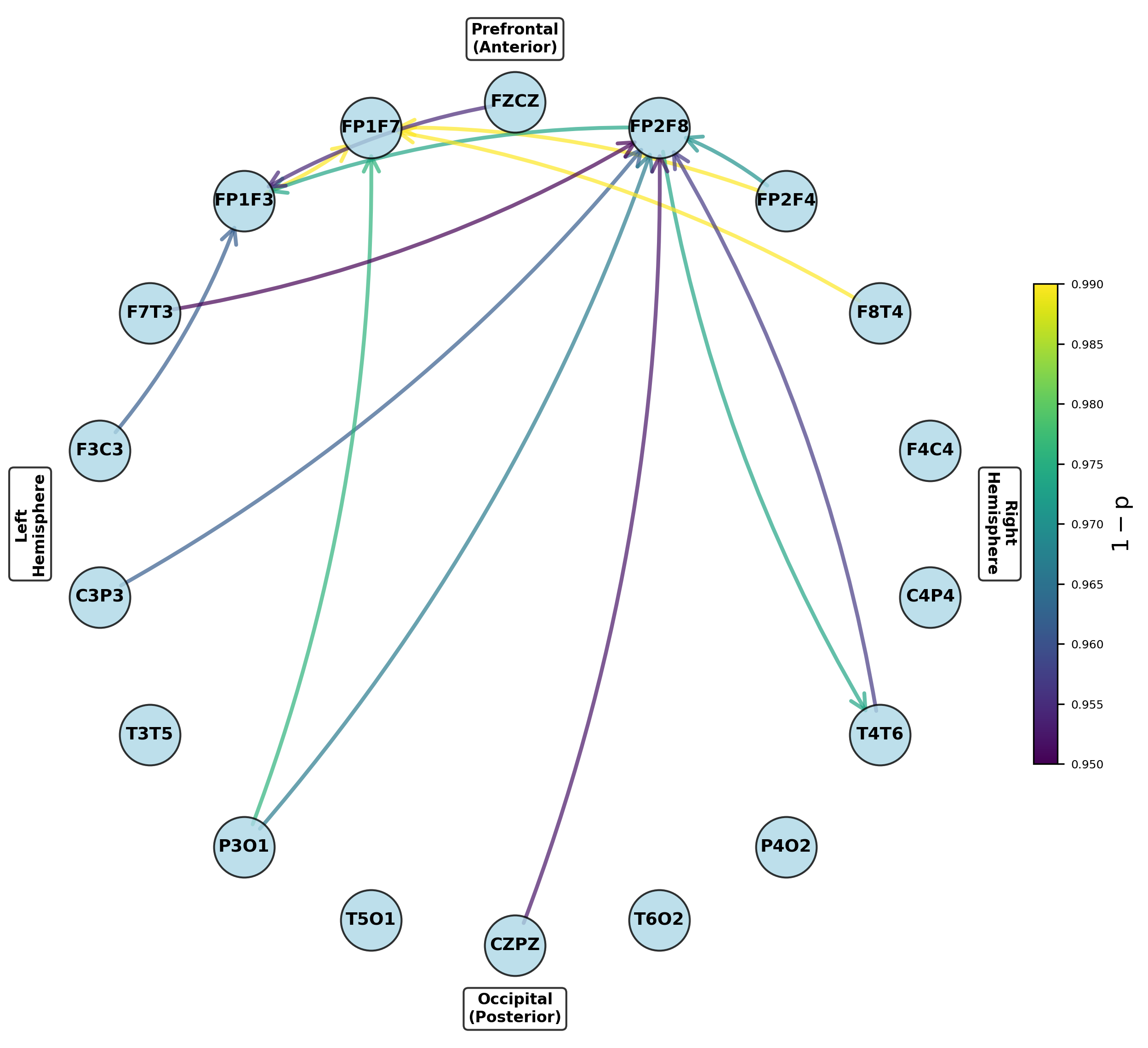
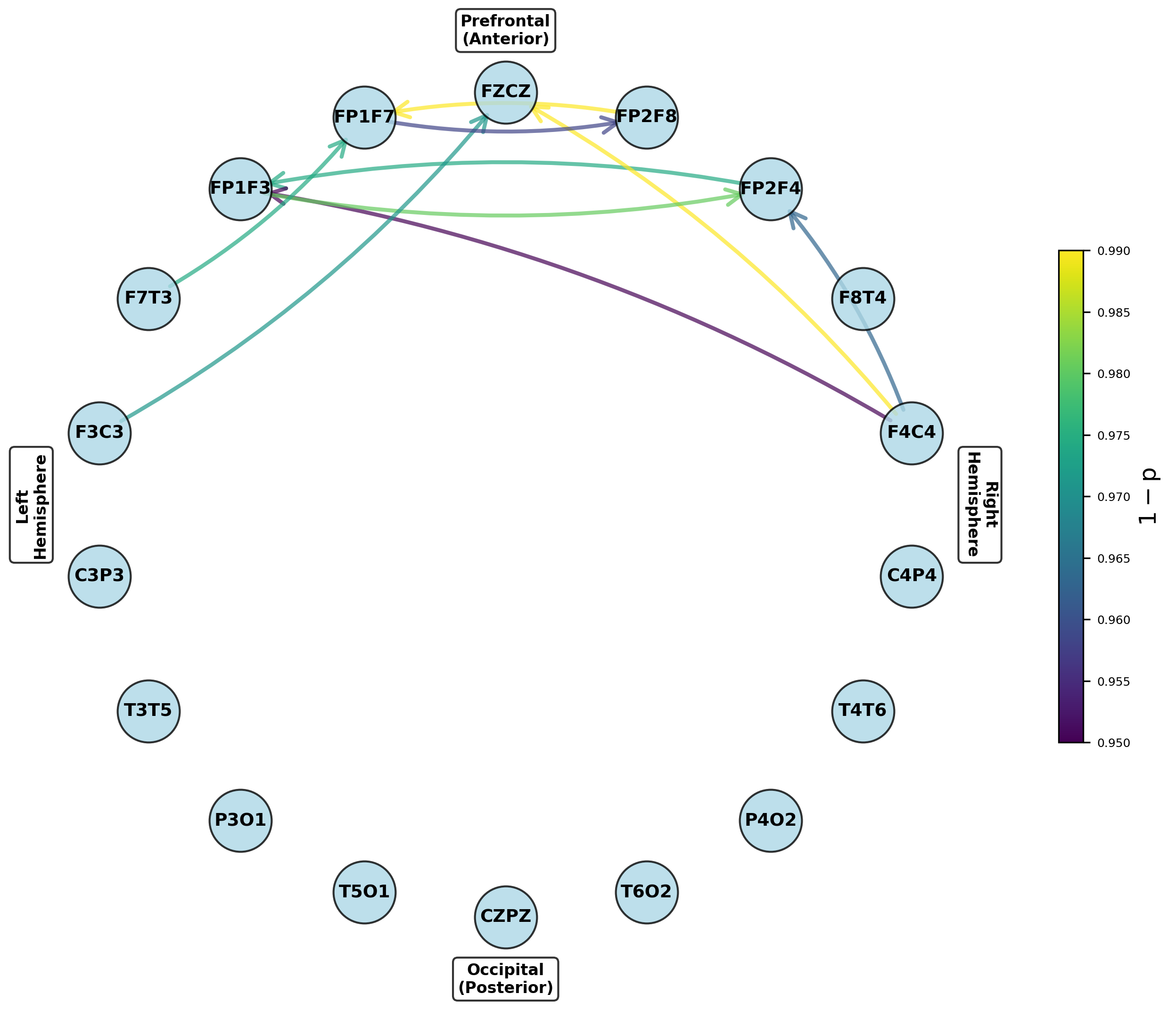
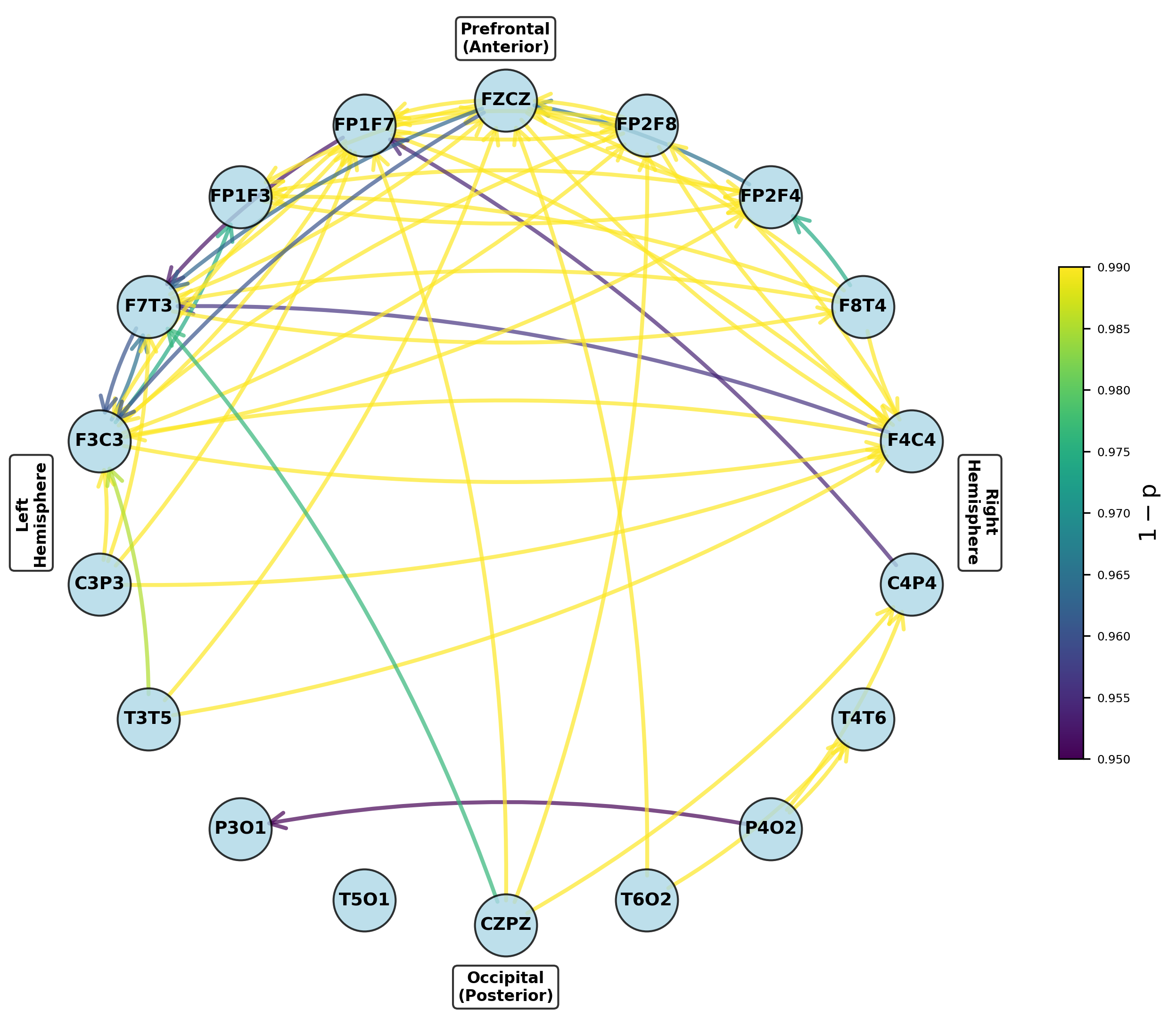
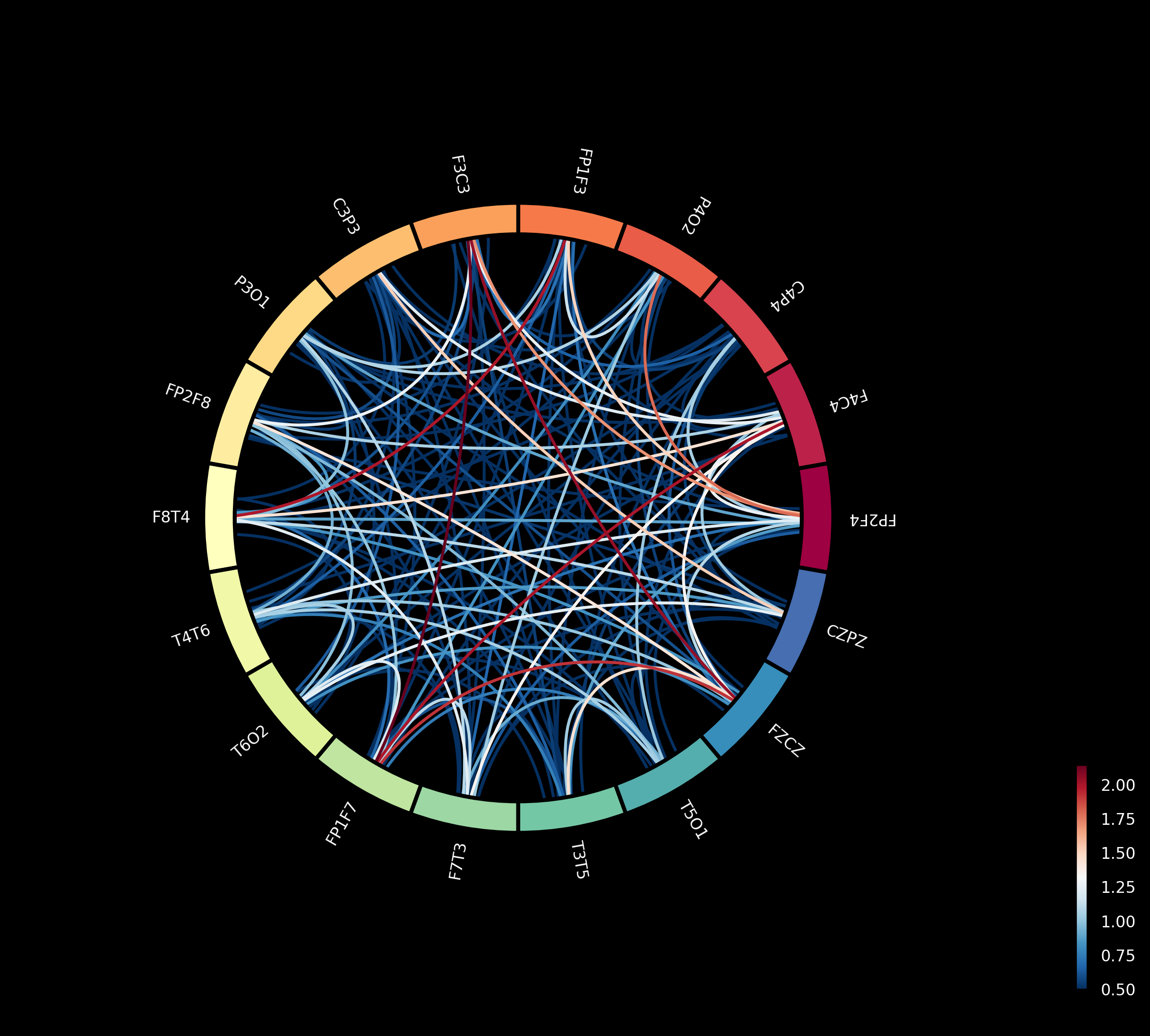
Figure 6: Networks highlighting channel pairs with significant variance differences between groups, notably in prefrontal outputs.
- Pair-matched preictal vs. ictal comparison reveals seizure-associated reconfiguration of dynamic causal nets.
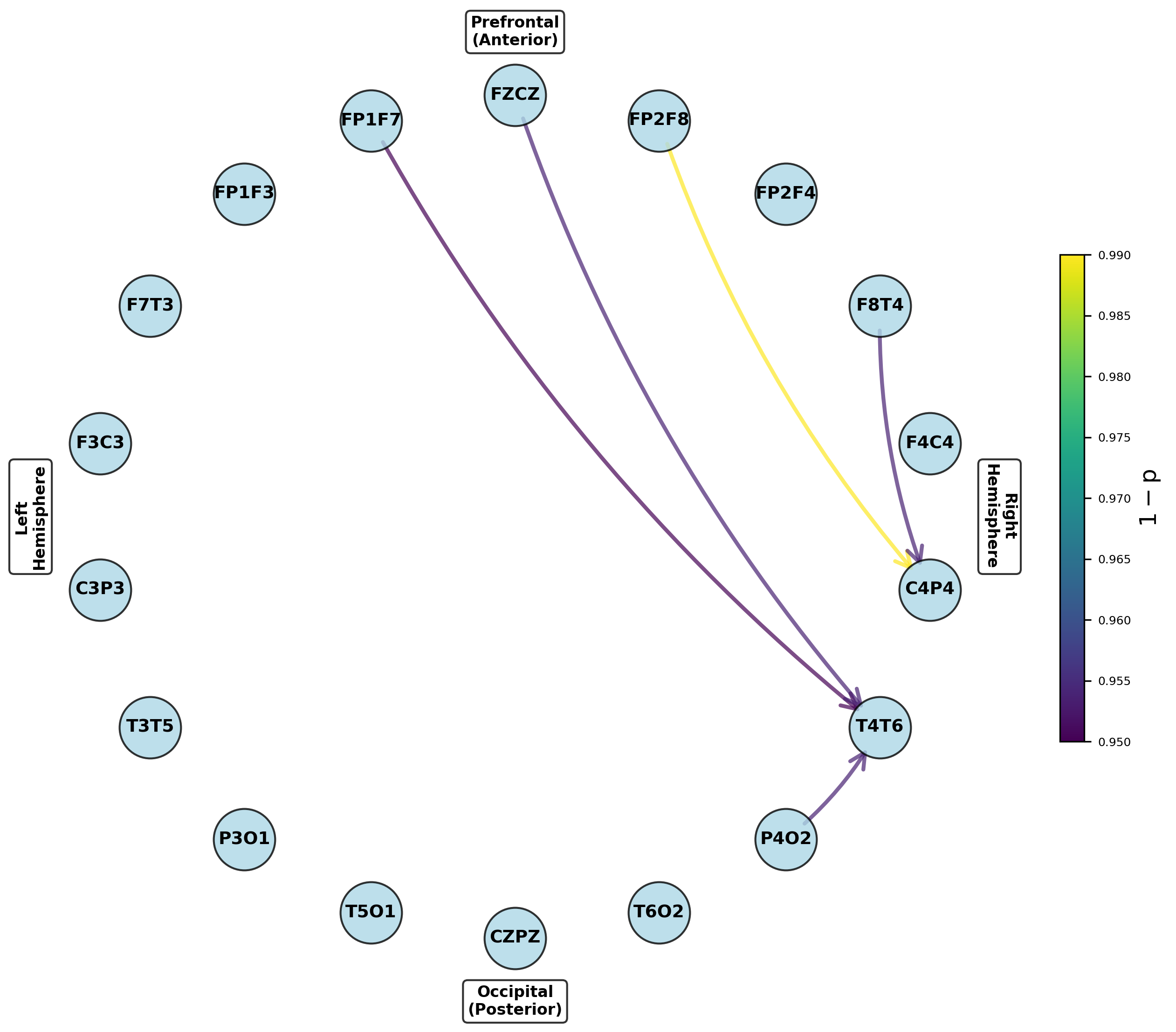
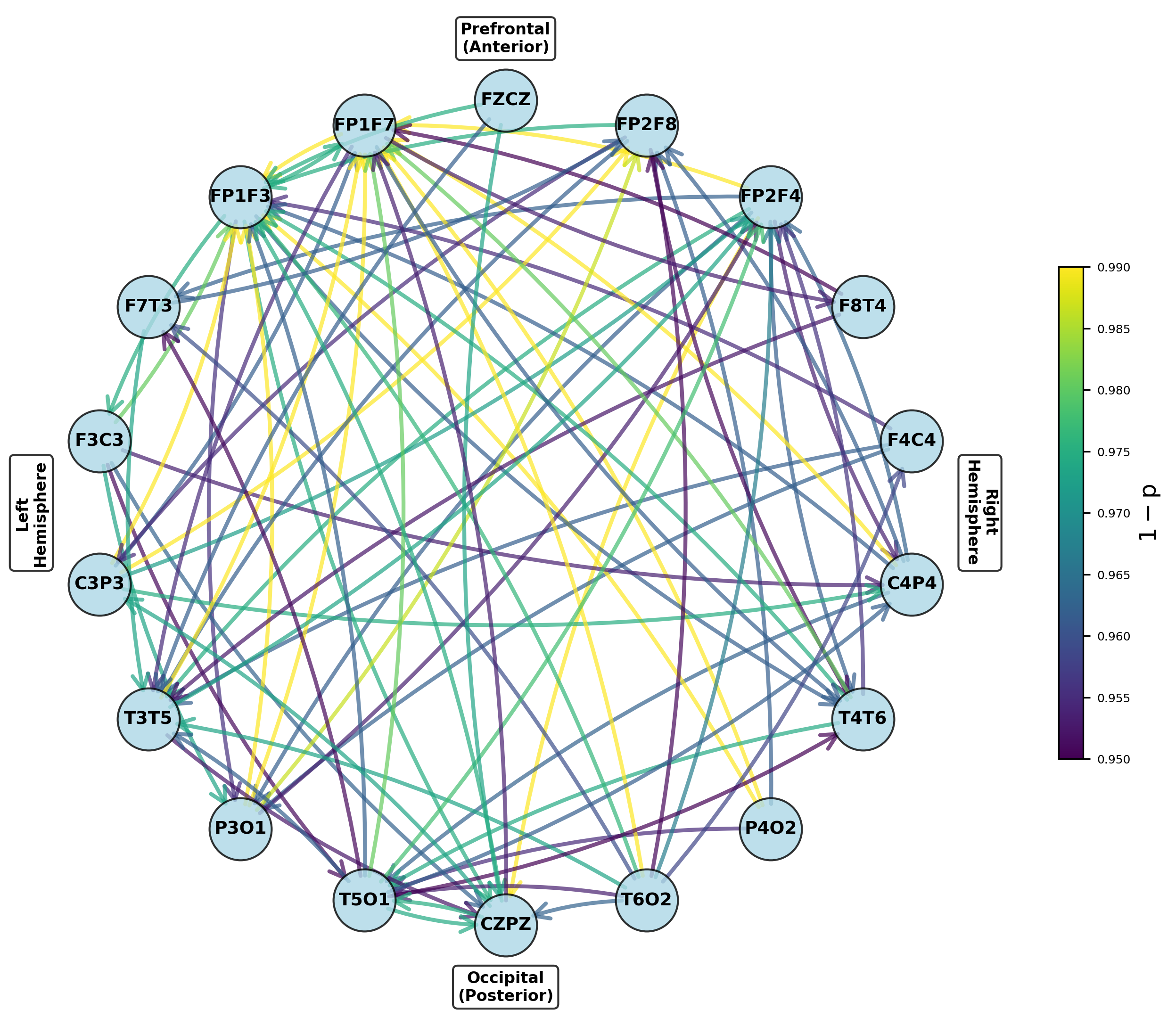
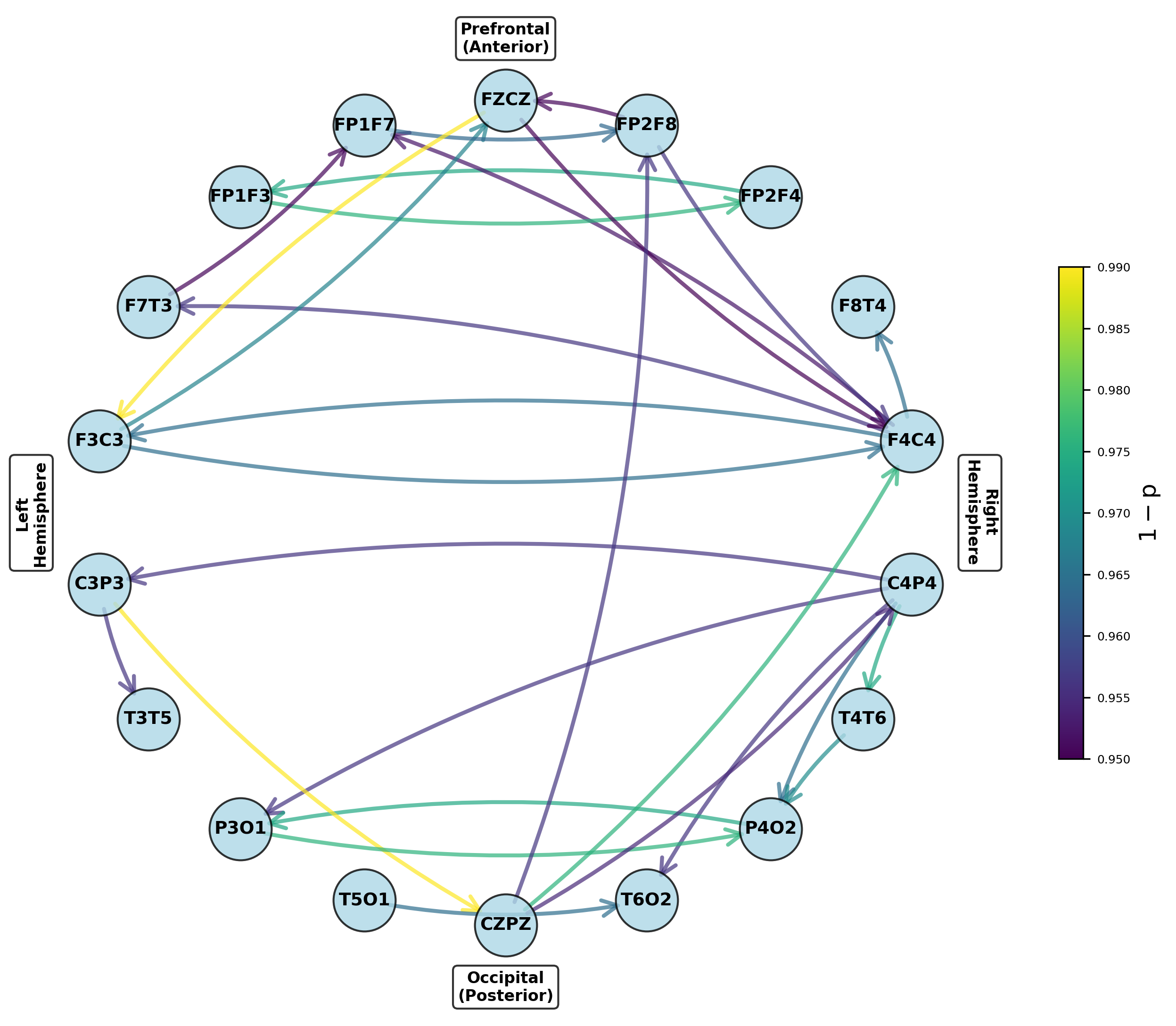
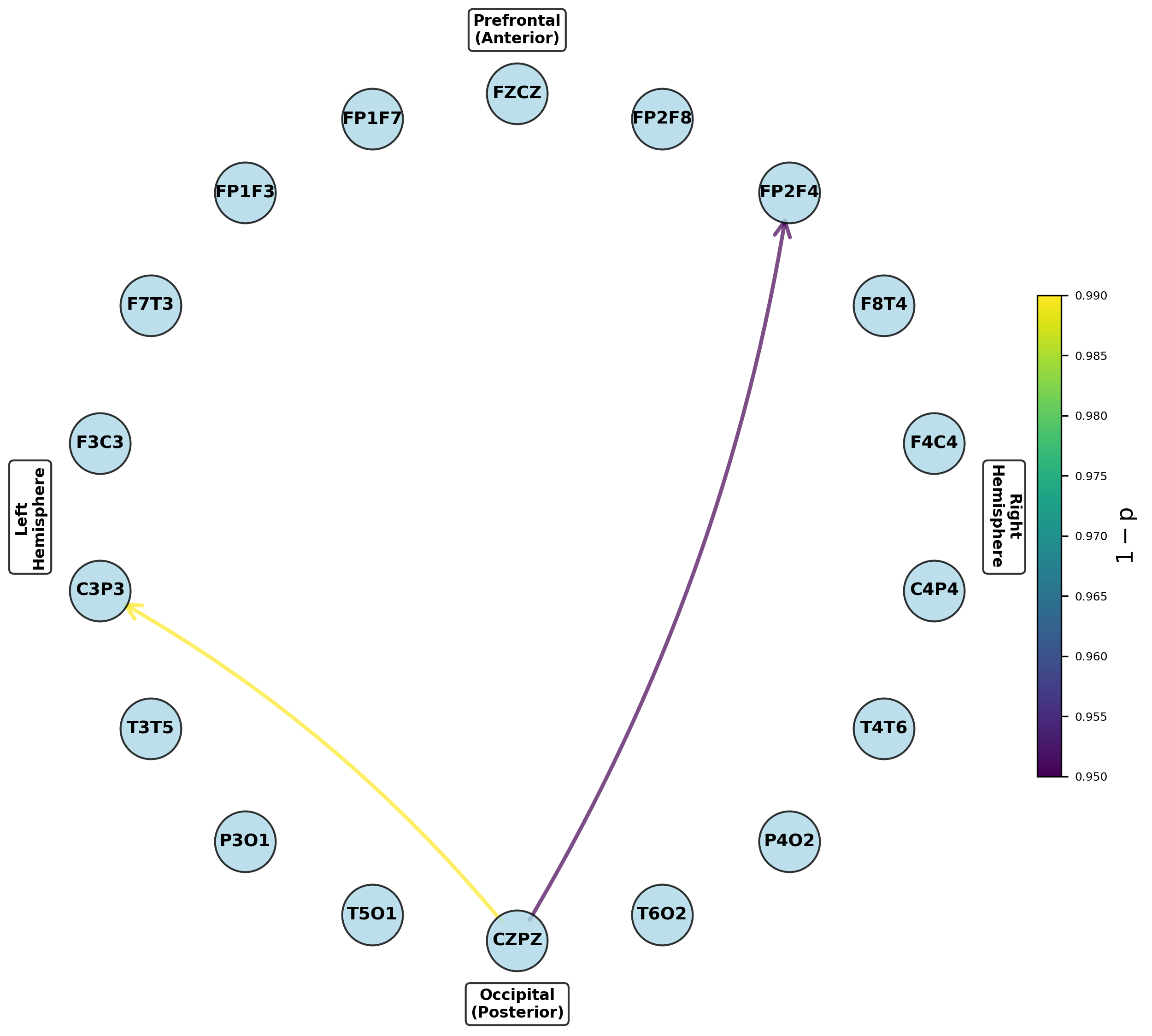
Figure 8: Networks for parameters exhibiting significant increases from preictal to ictal periods, indicating seizure onset network transitions.
- Comparison to conventional functional connectivity (lagged correlation) demonstrates greater sensitivity of NccDCM for detecting case-control differences post multiple testing corrections.
Simulation studies are systematically conducted to assess estimator accuracy for both local node models and multi-node networks. Median absolute relative errors (AREs) on key model-derived features—E/I ratio, within-channel and inter-channel coupling, firing threshold—are uniformly low (<10–15% under reasonable SNR), confirming practical estimability.
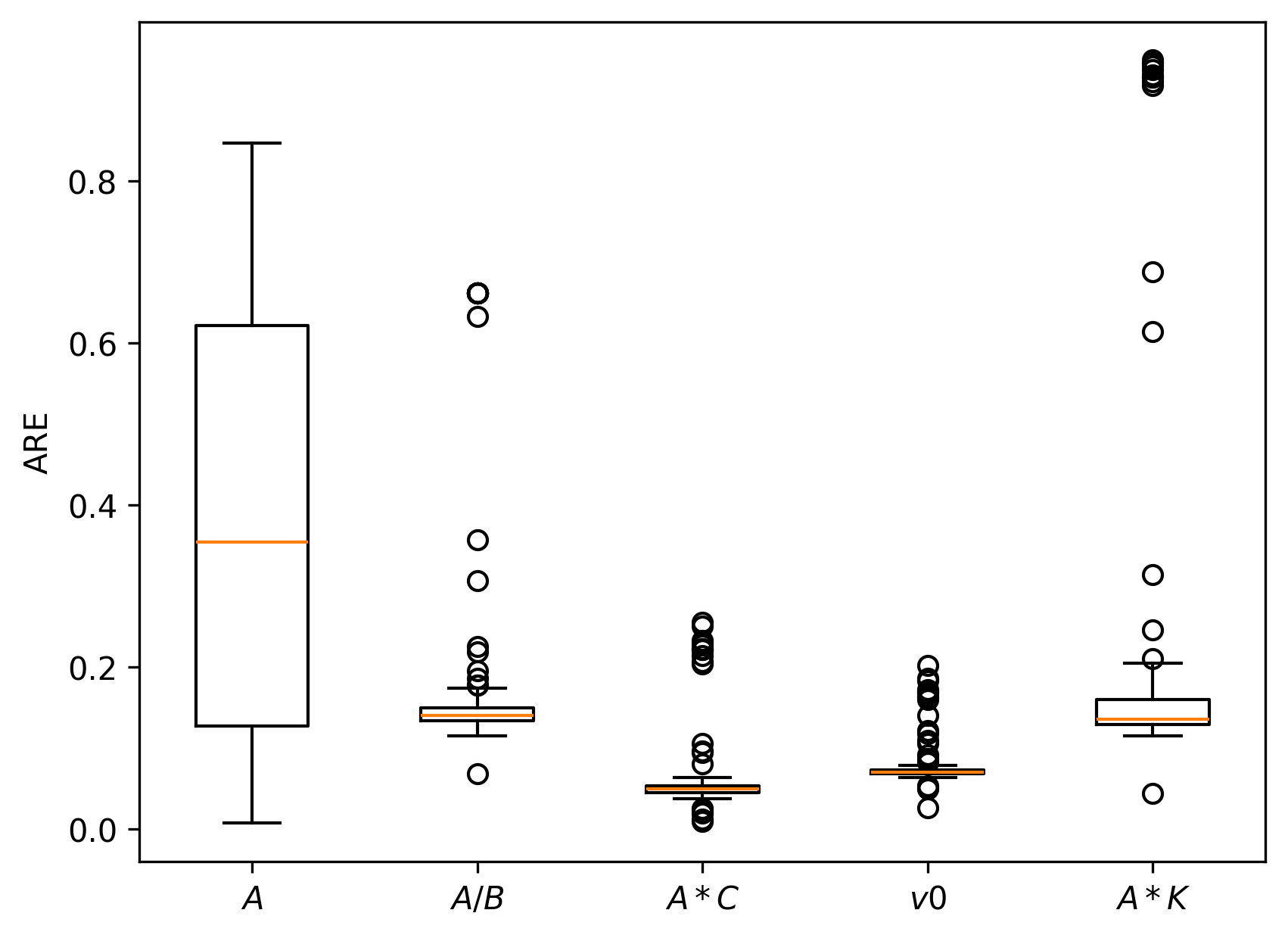
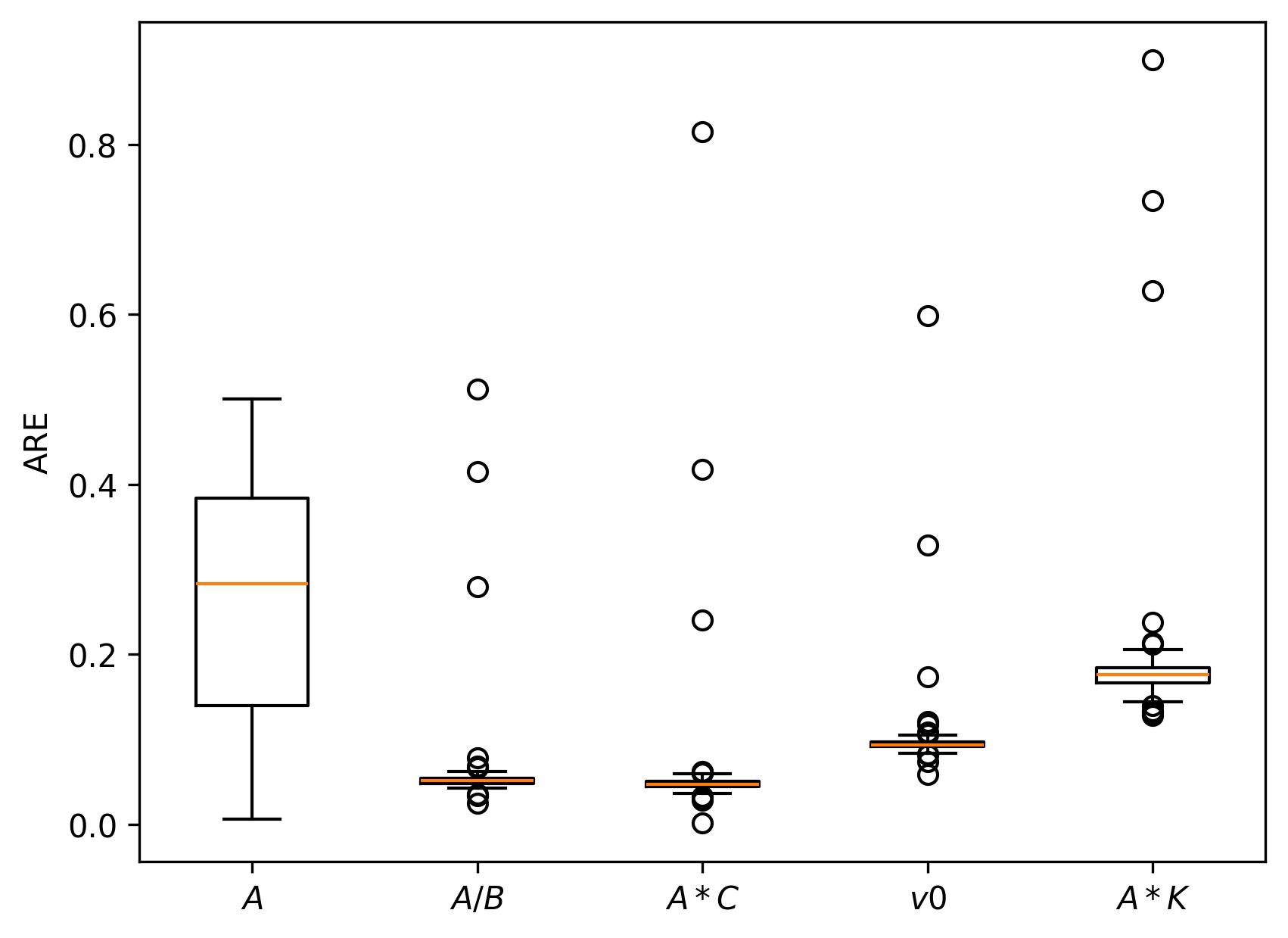
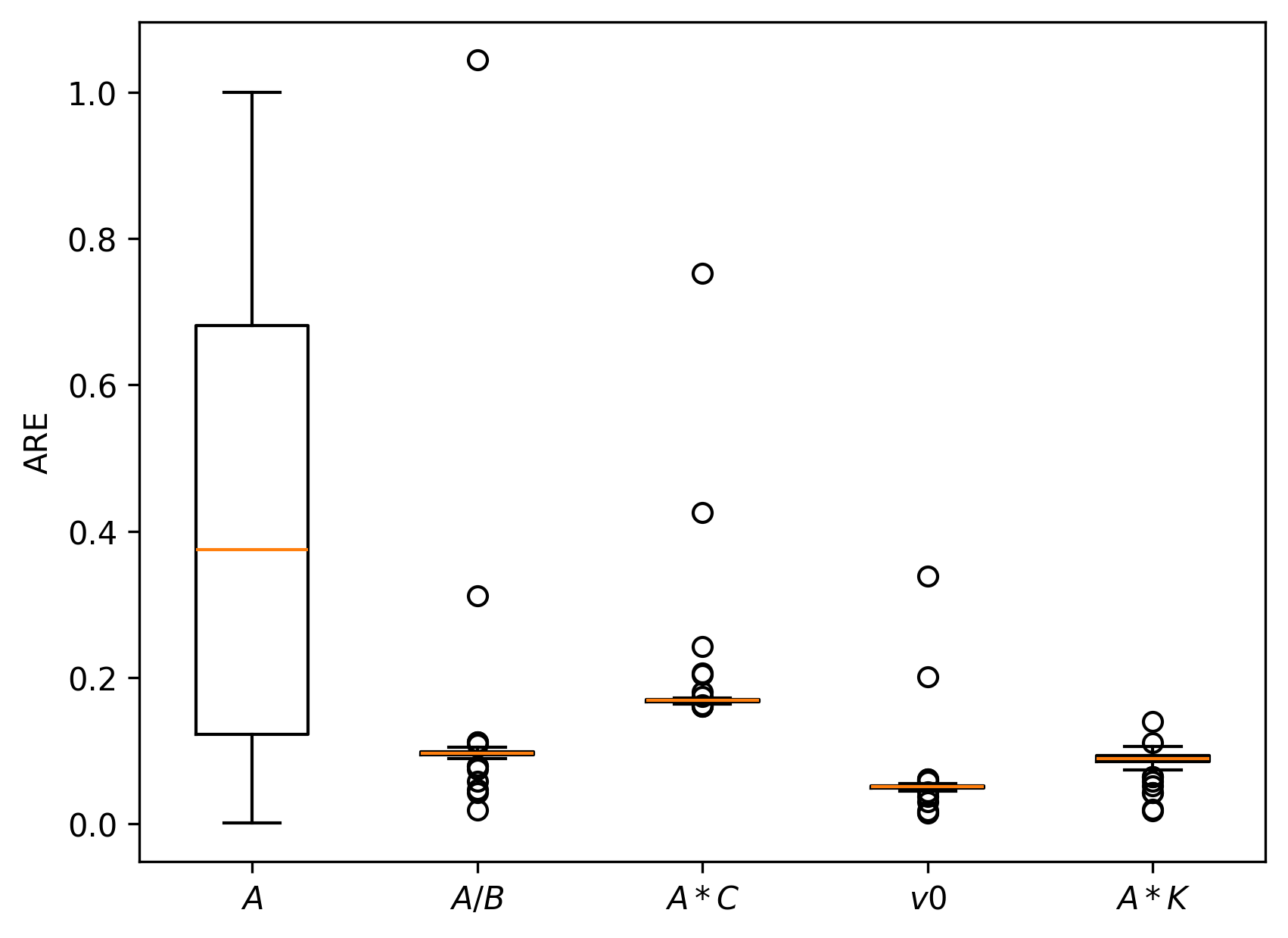
Figure 7: Boxplots of AREs for multi-node simulation under different parameter regimes, confirming robustness and accuracy of the loss-based estimation approach.
Identifiability analysis further clarifies that while excitation and inhibition cannot be uniquely separated, the ratio (E/I) is robustly estimable, aligning with prior theoretical results in conductance-based DCMs.
Implications: Theoretical and Practical Impact
This work advances the modeling of effective connectivity by introducing:
- A principled statistical approach to large-scale, heterogeneous, dynamic brain networks, breaking from conventional homogeneous, low-node DCMs.
- Scalable inference for clinical group comparisons, bridging microcircuit biophysics and macroscale network analysis, validated against real neuroclinical data.
- Robust numerical optimization for partially observed nonlinear SDE systems, opening new doors for interpretable, data-driven neural network modeling.
Key practical and theoretical implications include:
- Improved characterization of epilepsy as a network disorder, providing concrete evidence for network-level frontal and temporoparietal disruptions during seizures.
- Robustness against unmodeled individual heterogeneity (age, gender), as mixed-effects are explicit and parameter inference is channel- and individual-specific.
- Interpretability—estimated parameters directly relate to excitation/inhibition, connectivity, and threshold, enabling causal mechanistic hypotheses at the systems level.
Limitations and Future Directions
Acknowledged limitations include:
- Marginal screening in channel coupling estimation trades efficiency for scalability; more efficient penalized or stepwise model selection could further improve inference.
- Demographic covariates (age, gender) are presently omitted but can be naturally incorporated into the mixed-effects framework.
Future lines of research include:
- Extension to source-localized or deep-brain regions.
- Exploration of alternative neural mass or mean field models within the NccDCM architecture.
- Application to broader neuropathologies and perturbational regimes (e.g., pharmacological, stimulation).
Conclusion
The NccDCM framework provides a rigorously justified, computationally tractable, and biophysically interpretable approach for constructing, identifying, and comparing differential dynamic causal networks from EEG data. It enables new empirical insights into the mechanism of epileptic and other neural disorders at the network level, leveraging modern optimization and hierarchical modeling techniques. The open-source release of code and supplementary analysis enhances reproducibility and future applicability to large-scale clinical neuroimaging studies.




















