Path Sampling for Rare Events Boosted by Machine Learning
Abstract: The study by Jung et al. (Jung H, Covino R, Arjun A, et al., Nat Comput Sci. 3:334-345 (2023)) introduced Artificial Intelligence for Molecular Mechanism Discovery (AIMMD), a novel sampling algorithm that integrates machine learning to enhance the efficiency of transition path sampling (TPS). By enabling on-the-fly estimation of the committor probability and simultaneously deriving a human-interpretable reaction coordinate, AIMMD offers a robust framework for elucidating the mechanistic pathways of complex molecular processes. This commentary provides a discussion and critical analysis of the core AIMMD framework, explores its recent extensions, and offers an assessment of the method's potential impact and limitations.
Summary
- The paper introduces a novel AIMMD framework that integrates machine learning with transition path sampling to automate reaction coordinate discovery and enhance rare event sampling.
- It demonstrates efficient transfer learning across systems and robust identification of complex mechanistic pathways with improved acceptance rates and committor prediction.
- Methodological challenges such as validation of the learned committor and data imbalance in non-transition regions are critically evaluated, providing insight for future improvements.
Machine Learning-Enhanced Path Sampling for Rare Events: An Analysis of the AIMMD Framework
Introduction
The challenge of reliably sampling rare events in molecular simulations is central to computational chemistry and biophysics. Traditional molecular dynamics (MD) simulations are often unable to access the timescales relevant to such transitions, necessitating the development of specialized rare event sampling techniques. The paper "Path Sampling for Rare Events Boosted by Machine Learning" (2602.05167) offers a detailed discussion and critique of Artificial Intelligence for Molecular Mechanism Discovery (AIMMD), a framework that marries ML with transition path sampling (TPS) to automate and enhance the identification of reaction coordinates (RCs) and transition path ensembles. This essay offers an expert analysis of the AIMMD method, examines its technical innovations, evaluates its extensions, and considers its implications for the field.
Theoretical Background and Current Limitations
Rare, reactive events—such as chemical reactions, nucleation, and large conformational changes—are challenging for brute-force MD due to timescale separation. Enhanced sampling techniques, both biased (e.g., umbrella sampling, metadynamics) and unbiased (e.g., TPS, weighted ensemble), have been developed to address this, but their efficiency strongly depends on the choice of reaction coordinate or collective variable (CV). The committor probability, pB(x), is theoretically optimal as a RC but is computationally intensive to calculate directly, and its lack of physical interpretability limits mechanistic insight.
TPS sidesteps explicit RCs by generating unbiased ensembles of reactive trajectories, yet its efficiency still depends critically on the ability of a chosen CV to demarcate states A and B. Poor CV choices result in inefficient sampling and low acceptance rates. AIMMD circumvents this issue by integrating ML to iteratively infer the committor function directly from the TPS data, offering a data-driven approach to RC discovery and improved sampling in the critical transition region.
The AIMMD Framework: Algorithmic Structure and Innovations
AIMMD operates by initiating standard TPS to generate initial reactive trajectories, from which it collects shooting points as data for ML training. A feed-forward neural network, parameterized by θ and using system-specific CVs as input features, is trained to estimate the logit-committor, with the committor probability output as
pB(x)=1+e−q(x∣θ)1
The network is optimized using a negative log-likelihood loss based on the observed outcomes of shooting moves, leveraging both successful (AB) and unsuccessful (AA/BB) trajectories.
TPS moves are then guided by a Lorentzian selection probability centered around the predicted transition state, focusing sampling near the transition state ensemble (TSE) to maximize the efficiency of discovering new reactive trajectories. Upon convergence, symbolic regression is used to express the neural network's committor as an explicit analytical function of the original CVs, enhancing interpretability.
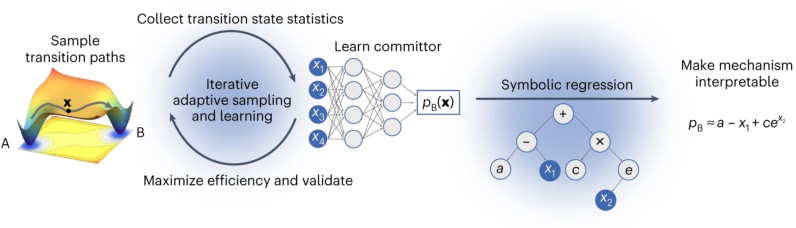
Figure 1: AIMMD alternates between TPS sampling and neural network training, ultimately expressing the learned committor via symbolic regression on physical collective variables.
AIMMD’s loop iterates: paths are sampled, the model is trained on new data, and TPS moves are biased by updated estimates until convergence. This tight integration of learning and sampling is a defining strength of AIMMD, making it adaptive to the true mechanistic coordinates of the system.
Empirical Results and Numerical Performance
AIMMD was tested on diverse systems—ion association/dissociation, gas hydrate nucleation, polymer folding, and membrane protein assembly—demonstrating efficient route discovery and robust RC identification even in high-dimensional or highly diffusive regimes [Hummer2023NCS]. For ion association/dissociation, the model trained on LiCl was rapidly adapted to other salts via few-shot transfer learning involving only the last neural network layer, evidencing strong transferability of the learned mechanisms. In the case of complex systems like Mga2 assembly, pooling data from parallel TPS runs enabled identification of multiple distinct mechanistic pathways, with the resulting committor landscape accurately reflecting mechanistic heterogeneity.
These results underscore the method’s capacity for efficient and generalizable mechanism discovery with a high degree of interpretability by mapping the neural RCs onto explicit CV expressions. Quantitative performance metrics, such as acceptance rates and committor prediction accuracy, have shown significant gains over standard TPS procedures.
Critical Evaluation and Methodological Considerations
While AIMMD succeeds in automating RC discovery and improving TPS efficiency, the method's validation of the committor model remains insufficiently rigorous within the framework. The lack of integration of the histogram test, which checks if configurations assigned the same committor indeed yield statistically indistinguishable shooting outcomes, leaves a gap in the quantitative assessment of model accuracy. As such, there is a risk that the learned committor (and thus the RC) is not truly optimal, especially in regions away from the TSE where training data are sparse.
A second limitation concerns the imbalance of training data distribution, as the Lorentzian selection biases most shooting points toward the TSE. While this improves sampling efficiency for transition states, it reduces committor accuracy in non-TSE regions, potentially limiting the method’s ability to predict off-pathway behaviors. Broader or adaptive selection strategies, or explicit inclusion of exploratory shooting moves, may be required to reconstruct accurate committor landscapes over the full configuration space.
Finally, AIMMD inherits TPS’s notorious difficulty in sampling multiple, well-separated mechanistic pathways, especially in systems presenting branching reaction networks. While parallel TPS and data pooling (as in the Mga2 assembly application) mitigate this to an extent, the completeness of pathway exploration still depends on initialization and statistical overlaps between pathway ensembles.
Methodological Extensions and Future Directions
Several enhancements have been proposed to address AIMMD’s limitations. Waste-recycling TPS [Bolhuis2023JCTC] reuses trajectories typically discarded in TPS (non-reactive paths), incorporating them into ML training to provide a more representative sample distribution, thus improving global committor estimation.
The integration of AIMMD with Transition Interface Sampling (AIMMD-TIS) [Bolhuis2025arXiv] represents a significant advance. AIMMD-TIS combines initial AIMMD-inferred committor surfaces with a TIS protocol to generate broader, nearly isocommittor-distributed interface ensembles, leading to denser and less biased coverage of configuration space and committor values. The method yields improved rate and free energy estimates and enables quantitative feature importance analysis along the reaction pathway.
AIMMD-TIS also more adequately samples alternative mechanistic channels by virtue of its interface-ensemble structure, bridging the gap towards comprehensive pathway space exploration. Extending this to replica exchange TIS protocols further promises to address branching mechanisms and slow inter-pathway transitions.
The growing software ecosystem (e.g., OpenPathSampling, PyRETIS, AIMMD’s own package) is facilitating the broader application and further evolution of these methods.
Implications and Prospects
AIMMD represents a convergence of ML and statistical mechanics, delivering an interpretable, efficient, and adaptive framework for mechanistic discovery in complex molecular systems. By automating RC discovery and enhancing sampling, AIMMD helps to alleviate a central computational bottleneck, thereby extending the applicability of path sampling techniques.
The broader practical impact will hinge on the method’s ability to maintain efficiency for highly diffusive and slow transitions seen in nucleation or large biomolecular assemblies. Its few-shot learning capabilities, as demonstrated in transfer learning experiments, highlight its promise for rapid adaptation to related chemical systems—a critical capability for data-driven chemical discovery and dynamic mechanism mapping.
From a theoretical perspective, AIMMD’s approach to RC discovery may inform new classes of hybrid ML/statistical mechanics methods, especially if supplemented with robust RC validation protocols and pathway space exploration tools. Deeper integration with on-the-fly interface optimization, adaptive sampling, and generalized committor learning is a conceivable next step.
Conclusion
AIMMD offers a principled, algorithmically sophisticated solution to reaction coordinate discovery and rare event sampling in molecular simulations. Its fusion of ML and TPS yields both efficient and interpretable models of mechanistic transitions, establishing a blueprint for further advances in data-driven rare event techniques. While certain methodological challenges persist—notably around model validation, data imbalance, and pathway completeness—the rapidly evolving ecosystem of AIMMD and its variants promises substantial future impact, both practical and theoretical, for the computational study of complex molecular systems.
Paper to Video (Beta)
No one has generated a video about this paper yet.
Whiteboard
No one has generated a whiteboard explanation for this paper yet.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Continue Learning
- How does the AIMMD framework integrate machine learning with traditional transition path sampling to improve rare event detection?
- What specific advantages does transfer learning offer in adapting the AIMMD method to different molecular systems?
- In what ways does symbolic regression contribute to the interpretability of the learned reaction coordinates in AIMMD?
- How do the challenges of training data imbalance and non-transition state sampling affect the overall performance of AIMMD?
- Find recent papers about machine learning-enhanced path sampling in molecular simulations.
Related Papers
- Transition Path Sampling with Boltzmann Generator-based MCMC Moves (2023)
- Combining transition path sampling with data-driven collective variables through a reactivity-biased shooting algorithm (2024)
- Supervised Learning and the Finite-Temperature String Method for Computing Committor Functions and Reaction Rates (2021)
- Doob's Lagrangian: A Sample-Efficient Variational Approach to Transition Path Sampling (2024)
- Everything everywhere all at once: a probability-based enhanced sampling approach to rare events (2024)
- Iterative variational learning of committor-consistent transition pathways using artificial neural networks (2024)
- Molecular free energies, rates, and mechanisms from data-efficient path sampling simulations (2023)
- AI-guided transition path sampling of lipid flip-flop and membrane nanoporation (2025)
- Optimal Rejection-Free Path Sampling (2025)
- Following the Committor Flow: A Data-Driven Discovery of Transition Pathways (2025)
Authors (2)
Collections
Sign up for free to add this paper to one or more collections.
Tweets
Sign up for free to view the 1 tweet with 0 likes about this paper.