- The paper introduces a novel JEPA-DNA framework that integrates joint-embedding predictive loss with token-level generative pre-training to capture global biological functions.
- It employs span-based masking, EMA-updated target encoders, and VICReg-inspired regularization to balance nucleotide-level detail and long-range semantic understanding.
- Experimental results demonstrate significant AUROC gains in promoter prediction and variant effect interpretation, underscoring its practical impact on genomic analysis.
JEPA-DNA: Grounding Genomic Foundation Models with Joint-Embedding Predictive Architectures
Motivation and Limitations of Generative Objectives in Genomic Foundation Models
Genomic Foundation Models (GFMs) have predominantly relied on Masked Language Modeling (MLM) and Next Token Prediction (NTP), adapting LLM paradigms for nucleotide sequences. Despite success in modeling local syntax and motif structure, these objectives bias models toward reconstructing individual nucleotides, over-allocating capacity to non-informative high-frequency noise. Thus, existing GFMs exhibit poor grounding in global biological semantics and insufficient modeling of regulatory, structural, and functional relationships across distant genomic intervals.
JEPA-DNA is motivated by the need to break this "granularity trap" by grounding pre-training in objectives that force acquisition of global, functionally meaningful representations. The method injects inductive bias toward high-level biological logic rather than mere sequence syntax.
The JEPA-DNA Architecture
JEPA-DNA introduces a hybrid framework combining joint-embedding predictive architectures (JEPA) with conventional token-level generative pre-training. The architecture is composed of a context encoder, a target encoder updated via EMA, and a predictor head responsible for mapping context representations in the masked input to the predicted latent embedding of the unmasked target sequence.
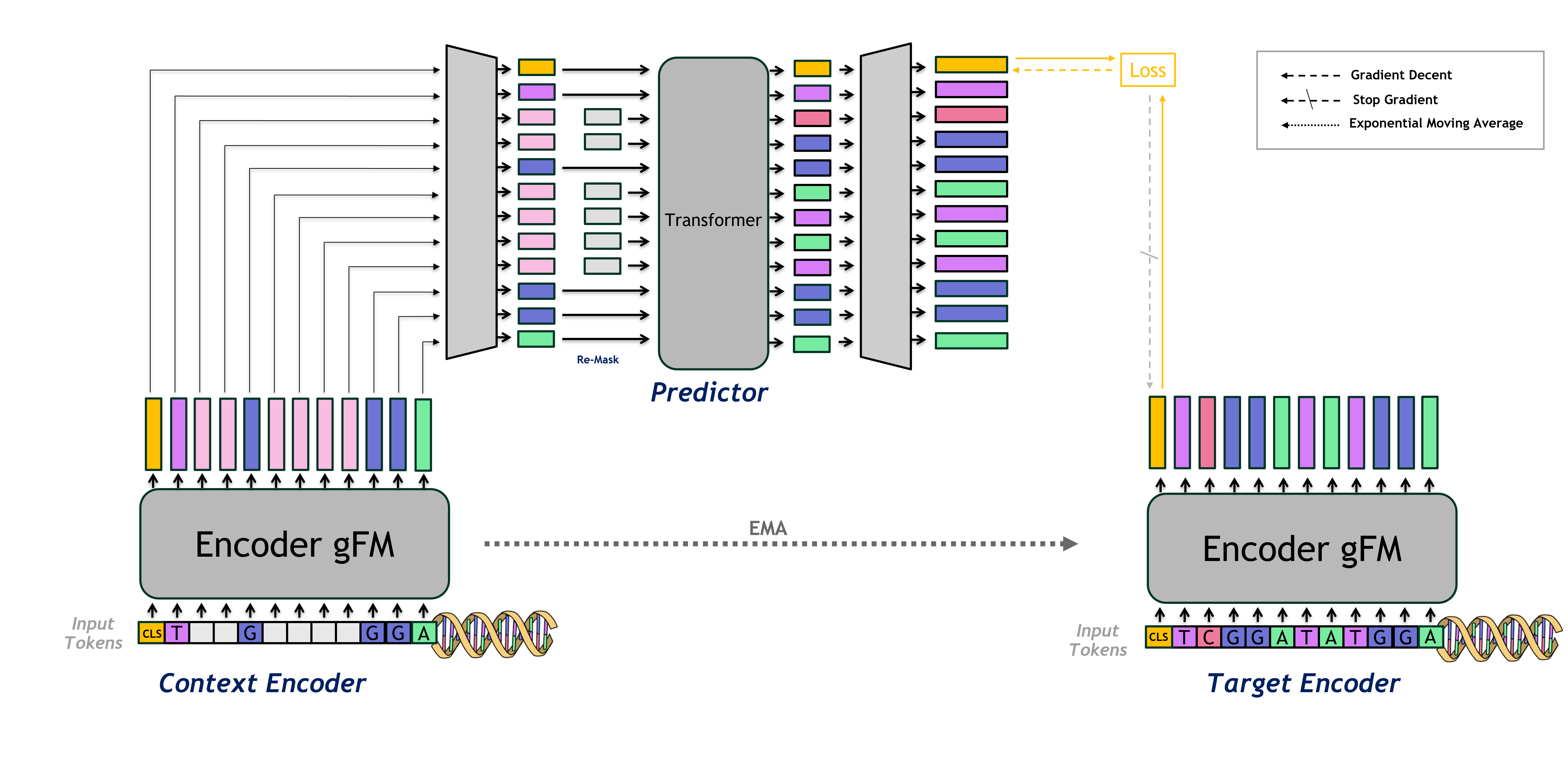
Figure 1: Diagrammatic overview of the JEPA-DNA architecture, depicting dual masking, context/target encoders, and predictor head supervising the global CLS latent.
The pre-training pipeline involves a span-based masking of the input, with multiple contiguous blocks masked to drive the model toward harder reconstruction problems. The predictor head receives the (re-masked) context encoder outputs, including the global [CLS] representation, and is trained to predict the [CLS] latent vector from the target encoder. This dual-objective setup ensures the model learns nucleotide-level syntax and long-range, function-driven embeddings.
Loss Functions and Regularization
The total training objective is a composite sum of:
- LLM Loss (LLLM): Standard MLM or NTP applied over masked tokens for preserving base-level resolution.
- Latent Predictive Loss (Ljepa): Cosine similarity between the predictor output and the target encoder [CLS] embedding; enforces global semantic prediction.
- VICReg-Inspired Regularization: Variance and covariance losses (Lvar, Lcov) applied to batch embeddings to prevent latent space collapse (degenerate constant solutions) and promote feature diversity.
Hyperparameters control the mixture of these components, and the framework is agnostic to underlying backbone (Transformer, SSM, LongConv).
Experimental Results: Linear and Zero-Shot Evaluation
Extensive experiments establish the empirical advantage of JEPA-DNA over standard generative-only pre-training regimes. DNABERT-2 is used as the primary backbone, with JEPA-DNA introduced through continual pre-training.
Linear probing protocols (encoder frozen, linear classifier trained) yield consistent performance enhancement across standard genomic benchmarks—promoter prediction, transcription factor binding, splice site detection, and variant effect interpretation. For example, AUROC increases of up to +5.98% (VB Coding Pathogenicity) are reported, directly attributable to the improved feature expressivity brought by latent grounding.
In zero-shot evaluation, where classifier heads are not fine-tuned and function relies solely on cosine similarity of sequence embeddings, JEPA-DNA exhibits notable gains in functional variant ranking (BEND Expression Effect, +6.94%), and pathogenicity annotation (Songlab ClinVar, +3.03%)—highlighting its superior capture of high-level semantics.
Architectural Generalization and Practical Considerations
The architecture and objectives of JEPA-DNA are explicitly designed for compatibility across backbone types as well as with either generative pre-training protocol. [CLS] or [EOS] tokens are used as targets for the predictive latent loss, and the predictor can be attached either to MLM- or NTP-trained models.
Key practical details include the importance of predictor warmup to avoid instability, careful normalization in variance regularization, and mitigation of variance artifacts arising from input length heterogeneity.
Theoretical and Practical Implications
The adoption of JEPA-style latent objectives addresses not only structural deficiencies of local generative models but also suggests a new paradigm for grounding sequence models in global biological function. This explicit operationalization of "world modeling" in genomics parallels similar trends in computer vision and NLP, where predictive latent architectures yield better semantic alignment.
Practically, the improvements in downstream linear and zero-shot performance portend better annotation, interpretation, and variant ranking in both research and clinical contexts, especially as GFMs are deployed for candidate variant prioritization, regulatory genomics, and trait prediction.
Future Directions
The work points to several important next steps:
- Pre-training from scratch using JEPA-DNA, versus its current formulation as continual pre-training for already-trained GFMs.
- Extending to alternative architectures (SSMs, Long-Conv, HyenaDNA) and tokenizations.
- JEPA-centric paradigm shift: Investigating the theoretical limit where the generative loss becomes auxiliary and latent prediction is primary, possibly completely supplanting MLM/NTP.
- Rigorous ablation and statistical significance reporting, critical for benchmarks with regulatory or clinical importance.
- Integration of biologically-informed masking strategies and aggregation operators for even tighter grounding in regulatory grammar and chromatin organization.
- Comparative benchmarking against contrastive and alignment-based self-supervision approaches, to map the tradeoffs unique to joint-embedding latent objectives.
Conclusion
JEPA-DNA marks a substantial methodological advance in the pre-training of Genomic Foundation Models by integrating explicit latent grounding with conventional token-level objectives. Empirical results support its efficacy for producing more robust, globally meaningful representations, yielding downstream performance improvements in both supervised and zero-shot tasks. The approach is likely to influence the trajectory of foundation model pre-training in genomics, with implications for annotation, interpretation, and trait elucidation at scale (2602.17162).
