- The paper presents a Hierarchical Discrete Diffusion Model that improves chemical validity while enhancing exploration in molecular graph generation.
- It introduces Decoupled Atom Encoding to precisely characterize atomic features, significantly increasing reconstruction fidelity and accurate motif generation.
- MolHIT achieves state-of-the-art performance on MOSES and GuacaMol benchmarks, offering robust conditional control over generated properties.
MolHIT: Advancing Molecular-Graph Generation with Hierarchical Discrete Diffusion Models
Introduction and Motivation
Molecular graph generation is a cornerstone task for AI-driven applications in drug discovery and materials design, where the challenge lies not only in structural novelty but also in rigorous chemical validity. Existing approaches based on graph discrete denoising diffusion models provide stronger inductive bias for structural generalization compared to 1D sequence-based models, but systematically trail in terms of chemical correctness and conditional property controllability. The MolHIT framework is introduced to explicitly address these limitations, offering a new paradigm leveraging a Hierarchical Discrete Diffusion Model (HDDM) and Decoupled Atom Encoding (DAE).
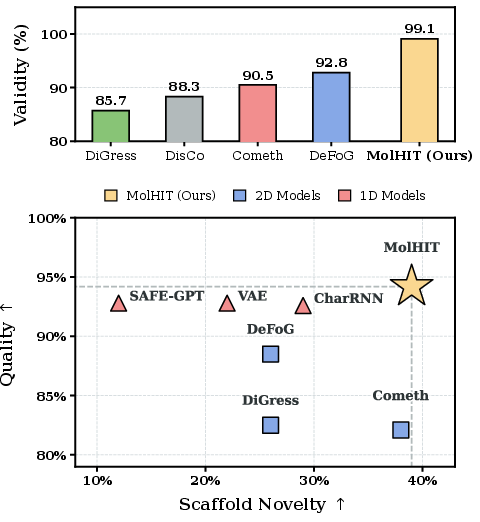
Figure 1: MolHIT achieves SOTA result on MOSES dataset. (Top) Near-perfect validity, outperforming existing graph diffusion models. (Bottom) Pareto-optimal in quality-novelty trade-off.
Hierarchical Discrete Diffusion Model (HDDM)
MolHIT introduces the HDDM, which generalizes the standard discrete graph diffusion process to operate over a coarse-to-fine discrete hierarchy, explicitly inducing mid-level semantic categories aligned with chemical prior knowledge. Instead of standard absorbing or uniform transitions, the process utilizes a deterministic, block-structured Markov chain: clean atomic types are mapped into semantically meaningful intermediate groups (e.g., grouping together all aromatic nitrogens, all halogens, etc.) before eventual masking, enabling the network to initially reason over abstracted chemical spaces prior to detailed atomic specification.
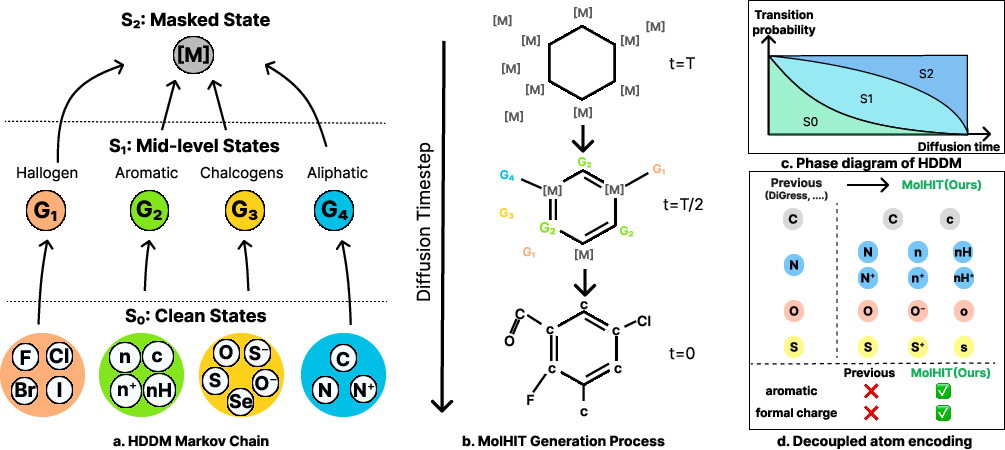
Figure 2: (a) Markov chain structure of HDDM; (b) Hierarchical denoising in generation; (c) Phase diagram for transition probabilities; (d) DAE representation separating aromatic and charged atoms.
Mathematically, the transition kernel supports rigorous NELBO-based training, and the design is compatible with flexible state space partitions, as proven in the appended lemmas. This allows MolHIT to outclass models that rely on uniform noising, and avoid the pitfalls of information-averse full masking at each step. The HDDM forward process can be tailored per-dataset by clustering atom types based on underlying chemical distributions.
Decoupled Atom Encoding (DAE)
A fundamental bottleneck of prior graph diffusion works is the use of coarse atomic encodings. Classical encodings collapse variants such as formal charge, aromaticity, and hydrogenation into a single identifier, making the denoising task ill-posed. MolHIT resolves this with DAE, where each atomic token decomposes into a tuple: elemental class, aromaticity, hydrogen count, and charge state. This greatly expands the state vocabulary, but allows for precise and unambiguous graph decoding and reconstruction.
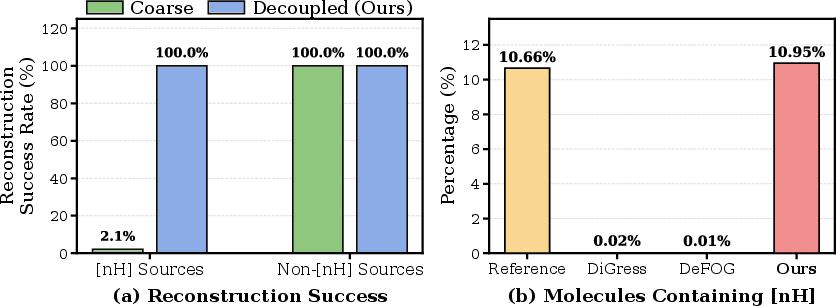
Figure 3: (Left) DAE achieves nearly 100% reconstruction for complex motifs on the Moses dataset versus prior approaches. (Right) DAE allows accurate generation of structures containing pyrrolic nitrogen.
DAE not only leads to a drastic increase in reconstruction fidelity (including complex moieties such as [nH], charged, and hypervalent atoms), but also enables the model to match training set distributions for rare but drug-relevant structures, as further confirmed by ablation on GuacaMol.
Sampling and Conditional Generation
MolHIT introduces a Project-and-Noise (PN) sampler, which projects denoised predictions onto the discrete manifold and immediately re-samples noise in the Markov chain. This avoids restrictions inherent to posterior-based updates, facilitating improved exploration, essential for complex chemical spaces. The framework also incorporates temperature and nucleus/top-p sampling, which gives robust control of the quality-diversity trade-off in unconditional and conditional generation.
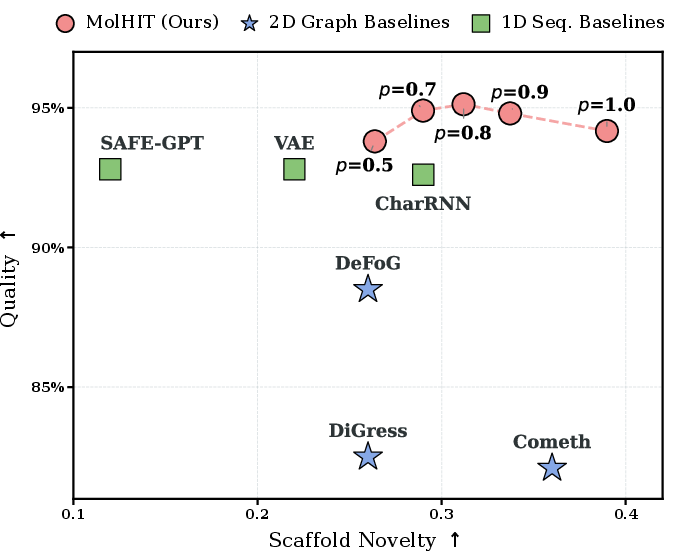
Figure 4: Effect of top-p sampling in MolHIT: quality-diversity trade-off modulation.
For conditional generation, MolHIT modifies the baseline graph transformer with adaptive layer normalization and classifier-free guidance, supporting multi-property conditioning (e.g., drug-likeness, SA, logP, MW). This design achieves high compositionality and property alignment.
Empirical Results
MolHIT achieves new state-of-the-art results on MOSES and GuacaMol benchmarks. On MOSES, MolHIT sets a new upper bound across most standard metrics (quality, validity, FCD, scaffold novelty) and attains near-perfect validity (>99%), distinctly surpassing strong 1D and 2D baselines. Importantly, its scaffold novelty and recovery scores indicate generalization beyond memorization-heavy sequence-based models.
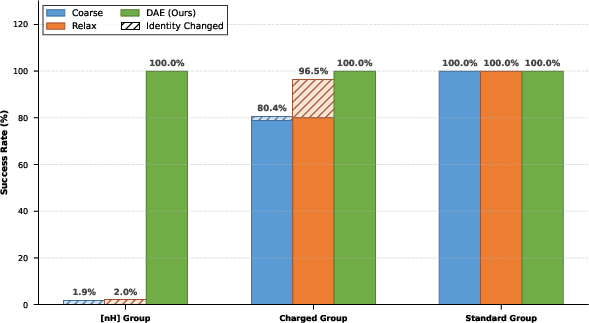
Figure 5: Ratios of generated molecules with formal charge – only DAE-based approaches recover the empirical statistics.
In ablation, DAE is shown to be necessary (but not sufficient) for full reconstruction and valid motif generation (including charged and aromatic heteroatoms). The HDDM and PN-sampler further boost sample quality and chemical realism. In rigorous conditional tests (multi-property guidance, scaffold extension), MolHIT exhibits lower MAEs and higher Pearson r relative to all graph diffusion comparators, while maintaining high validity.
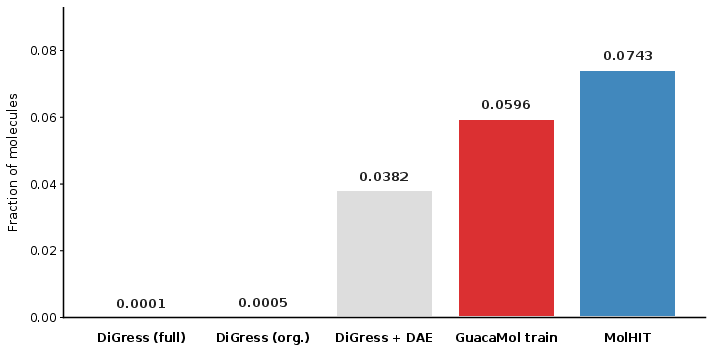
Figure 6: Fidelity and identity preservation for formal charge motifs – DAE enables near-perfect retention.
Theoretical and Practical Implications
From a theoretical standpoint, MolHIT demonstrates that hierarchical, domain-knowledge-infused noise schedules lead to provable advantages in discrete generative graph modeling: more effective exploration, better inductive bias, and no loss of closed-form ELBO guarantees. Practically, this means that ML systems can now correctly generate charged, aromatic, and complex heteroatom-containing molecules, which are otherwise systematically underrepresented in the output of all prior diffusion architectures.
Beyond molecular design, the HDDM framework can be directly extended to any discrete structured domain where latent hierarchical priors exist (e.g., macromolecules, language modeling with semantic token grouping). Moreover, DAE establishes that explicit, chemically motivated decompositions are essential for accuracy in generative chemistry.
Future Directions
There is clear headroom for further innovation, including model scaling, incorporating functional group motifs as mid-level HDDM states, application to 3D structures, and extension to protein design and multi-modal generative tasks. Exploration of more advanced conditional sampling and posterior inference routines is also warranted.
Conclusion
MolHIT establishes a new methodological and empirical standard for molecular graph generation, demonstrating that hierarchical discrete diffusion models, coupled with chemically-plausible atom encoding, close the validity-quality gap while retaining exploratory power. This work points toward general strategies for structured generation in scientific domains where categorical state spaces and strong priors are present.





