- The paper introduces Automatic Prompt Generation (APG) to enhance cell instance segmentation by deriving robust prompt strategies without retraining.
- It empirically demonstrates that APG-augmented models consistently outperform standard SAM variants, particularly in challenging, high-density microscopy images.
- The study emphasizes that domain-specific adaptations and prompt refinement are critical for achieving robust segmentation across diverse imaging modalities.
Comprehensive Evaluation and Strategic Advancement of Foundation Models for Cell Instance Segmentation
Introduction
The paper "Revisiting foundation models for cell instance segmentation" (2603.17845) provides an exhaustive empirical evaluation of both general-purpose and domain-adapted segmentation foundation models for microscopy, focusing predominantly on adaptations and extensions of the SAM family (SAM, SAM2, SAM3). The authors benchmark specialized models (CellPoseSAM, CellSAM, μSAM, PathoSAM, CellViT) and propose a novel strategy, Automatic Prompt Generation (APG), designed to enhance SAM-based microscopy segmentation. The study systematically explores adaptation strategies, training data diversity/model size effects, and prompt-driven segmentation variants, yielding decisive insights into both the capabilities and limitations of current approaches.
Evaluation of Foundation Model Adaptations
Microscopy instance segmentation methods are largely derived from the SAM model architecture. General-purpose SAM variants are routinely adapted by three principal methods: 1) prompt automation, 2) custom decoders trained on domain data, and 3) architecture finetuning for promptable segmentation. Microscopy-focused models (μSAM, PathoSAM, CellSAM, CellPoseSAM) leverage various combinations of these strategies.
The paper confirms that naive use of SAM's Automatic Mask Generation (AMG) modes is insufficient for complex microscopy tasks, especially in settings with ambiguous cell boundaries or high object density (see the segmentation failures highlighted). SAM3, with DETR-style instance prediction and concept-based segmentation, improves performance but does not match domain-specialized models unless further finetuned.
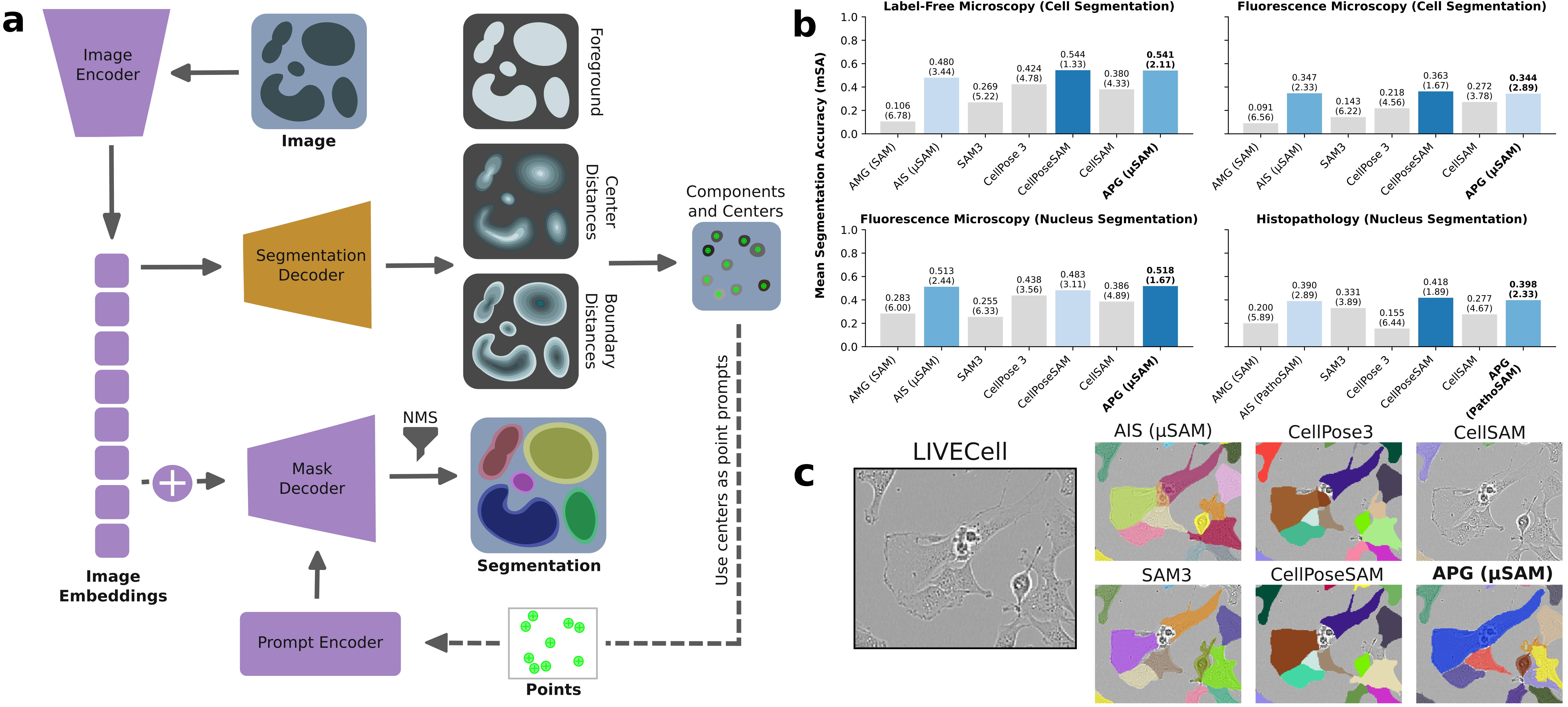
Figure 1: Panel a illustrates the APG framework layered atop μSAM, panel b summarizes modality-specific rankings across models, and panel c demonstrates APG's superior segmentation of challenging cell morphologies.
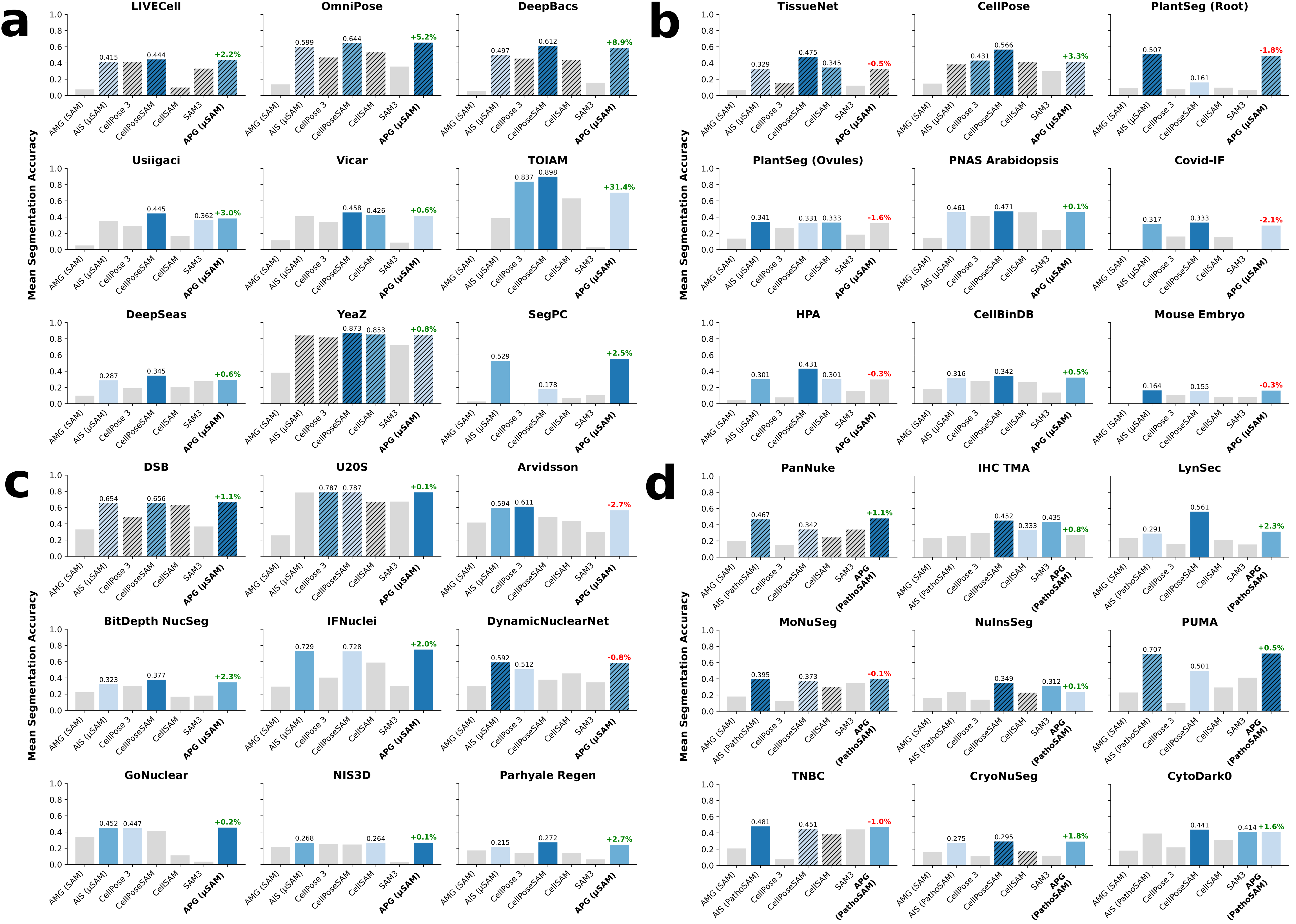
Figure 2: Modality-stratified mean segmentation accuracy for 36 datasets, with explicit model ranking, training split status, and APG's incremental improvements over prior methods.
Numerical results (mean segmentation accuracy, mSA) demonstrate that APG-enhanced μSAM and CellPoseSAM consistently outperform SAM/AMG and even the recently released SAM3 across four domains (label-free cells, fluorescent cells, fluorescent nuclei, histopathology nuclei). APG delivers pronounced gains—even on out-of-domain datasets—without retraining and achieves competitive results with state-of-the-art models. For example, APG improves μSAM's segmentation quality in all label-free microscopy cases, and exhibits substantial improvements on datasets such as TOIAM and DeepBacs.
Automatic Prompt Generation (APG)
APG fundamentally alters the instance segmentation regime for SAM-based models by deriving point prompts from decoder outputs—specifically from intersections of thresholded foreground, boundary, and center distance predictions—and using SAM’s prompt encoder to predict masks, followed by NMS to filter overlaps. Unlike CellSAM, which is dependent on accurate box detection, APG’s prompt strategy allows multiple masks per object, enhancing object recovery under domain shift. This approach sidesteps the trade-offs inherent in previous watershed-based strategies for seed determination, enabling robust segmentation of complex morphologies and ambiguous boundaries.
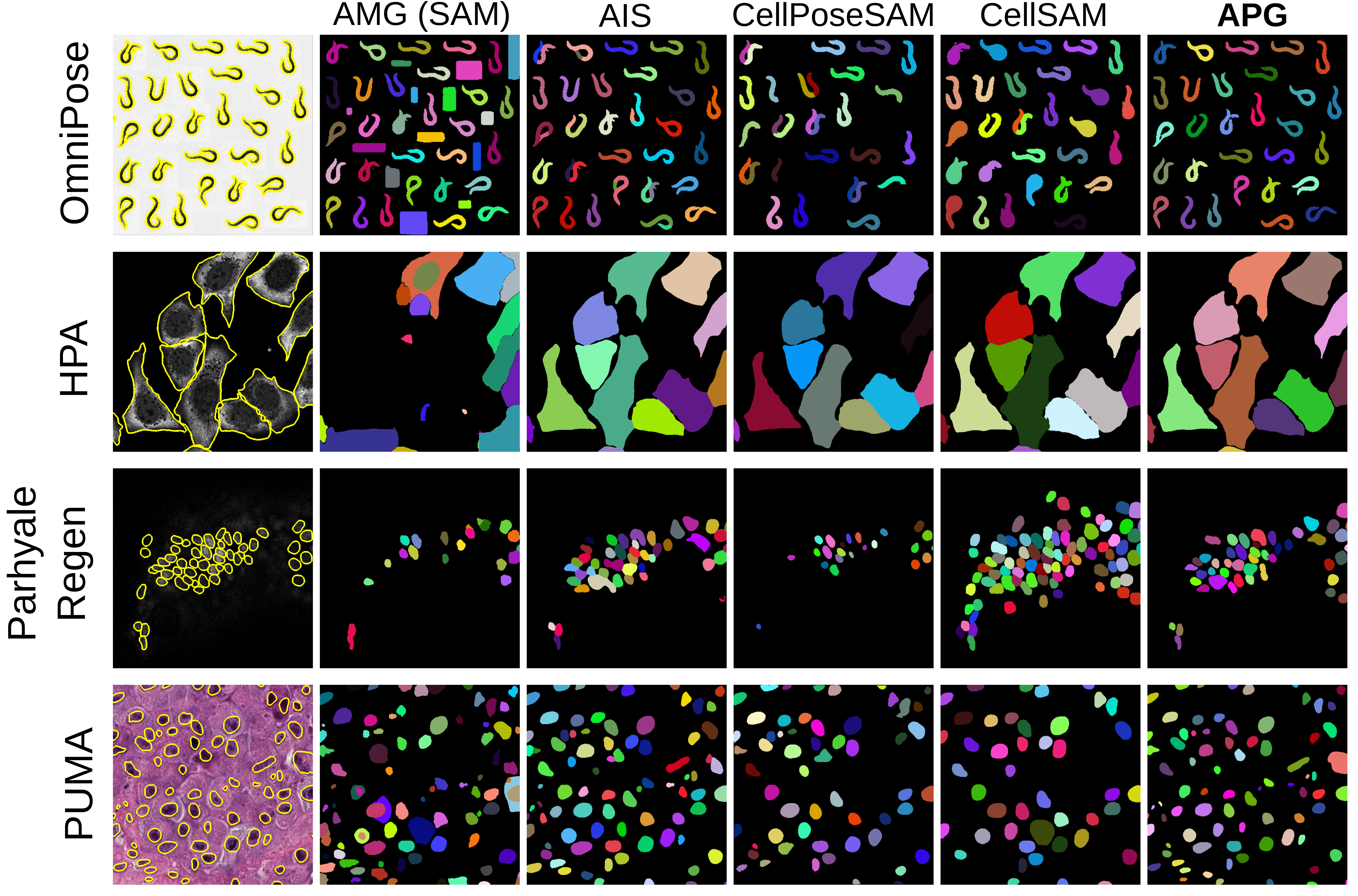
Figure 3: Qualitative segmentation examples comparing models on representative datasets across all tasks and modalities.
APG is applied as a post-process atop μSAM (and PathoSAM), requiring no retraining. Parameterization is simple and robust to default values, significantly simplifying deployment.
Comparative Prompt Strategies and SAM3 Prompt Sensitivity
The authors contrast APG’s connected component-based prompt derivation against a boundary distance maxima alternative. The former is decisively superior in segmentation accuracy across all modalities, reinforcing the utility of integrating domain-specific post-processing into prompt selection.
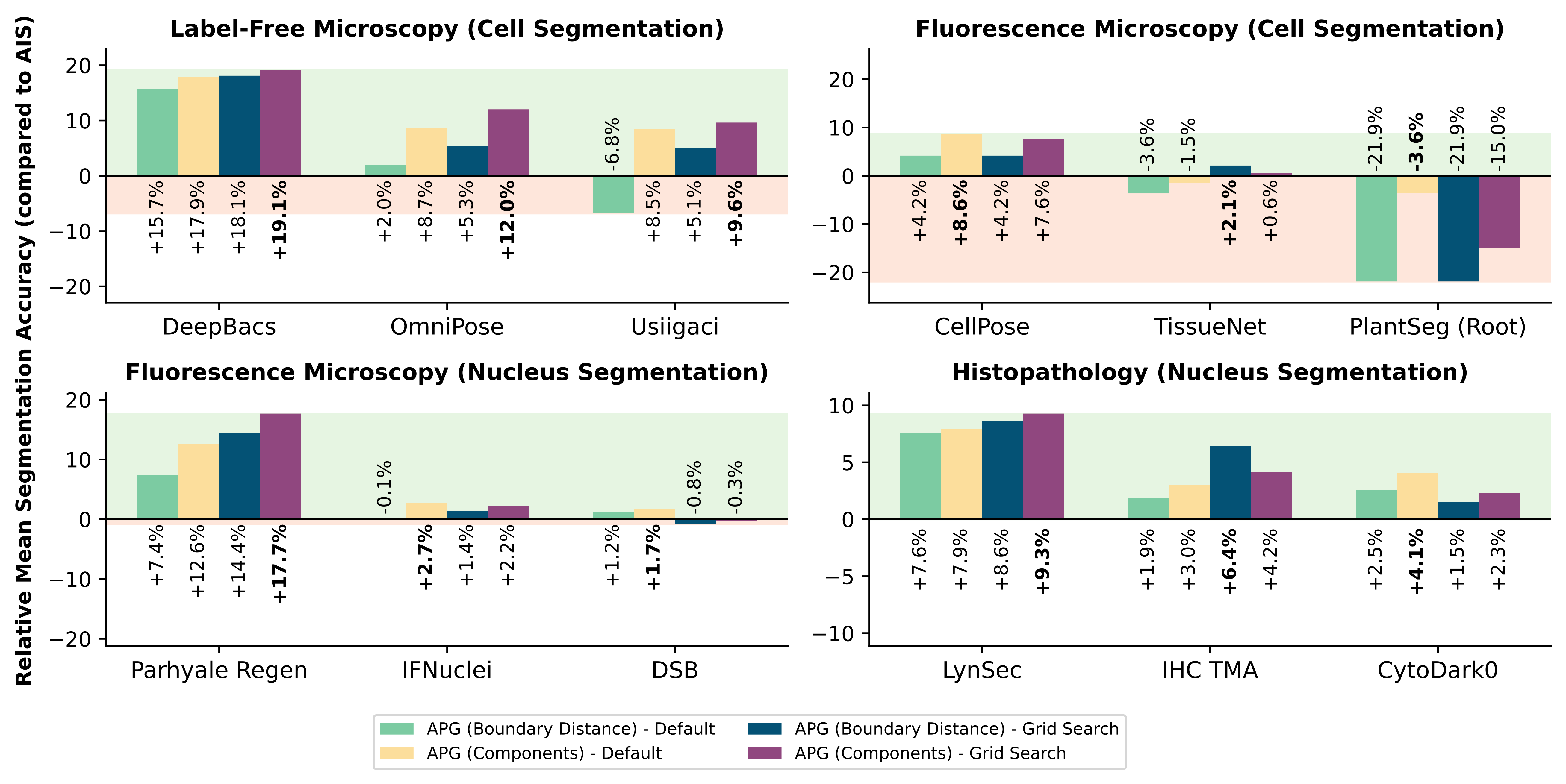
Figure 4: Accuracy gain of APG (Components) versus APG (Boundary) relative to AIS baseline across four imaging modalities.
SAM3’s text-prompt-driven segmentation is empirically shown to be highly sensitive to prompt phrasing. Biological terms such as "nucleus" are often unrecognized unless specifically present in training data, with shape descriptors ("blob", "dot", "irregular shape") sometimes yielding higher accuracy than canonical object names. This exposes a bottleneck in concept transfer and prompt generalization, suggesting that example-based prompting or explicit domain adaptation/fine-tuning is necessary for robust microscopy segmentation by concept-driven models.
Statistical Evaluation and Qualitative Insights
Paired Wilcoxon signed-rank tests corroborate the ranking of methods across datasets—APG, CellPoseSAM, and AIS dominate the top spots, with statistically significant improvements over standard SAM and SAM3. Visual assessment further confirms APG’s ability to resolve complex morphologies and maintain segmentation fidelity across diverse modalities and conditions.
Implications, Limitations, and Future Directions
Key claims substantiated by the study include:
- Domain-adapted foundation models (CellPoseSAM, μSAM + APG/PathoSAM + APG) consistently outperform general-purpose segmentation models (SAM/SAM3) for microscopy tasks.
- APG delivers substantial improvement in segmentation quality without additional model retraining.
- Model performance scales with domain-specific training data size and diversity; out-of-domain generalization is greatly enhanced by robust prompt generation strategies and post-processing.
- SAM3’s concept-driven architecture is highly sensitive to prompt formulation, limiting its practical utility in microscopy without further adaptation.
Practical implications include reduced technical burden for end-users, increased robustness in deployment across diverse datasets, and minimal inference-time tuning requirements. Theoretically, the work underlines the necessity of domain-specific adaptation even for large-scale, pre-trained foundation models and advocates for strategic fusion of automated prompt generation with finetuned architectures.
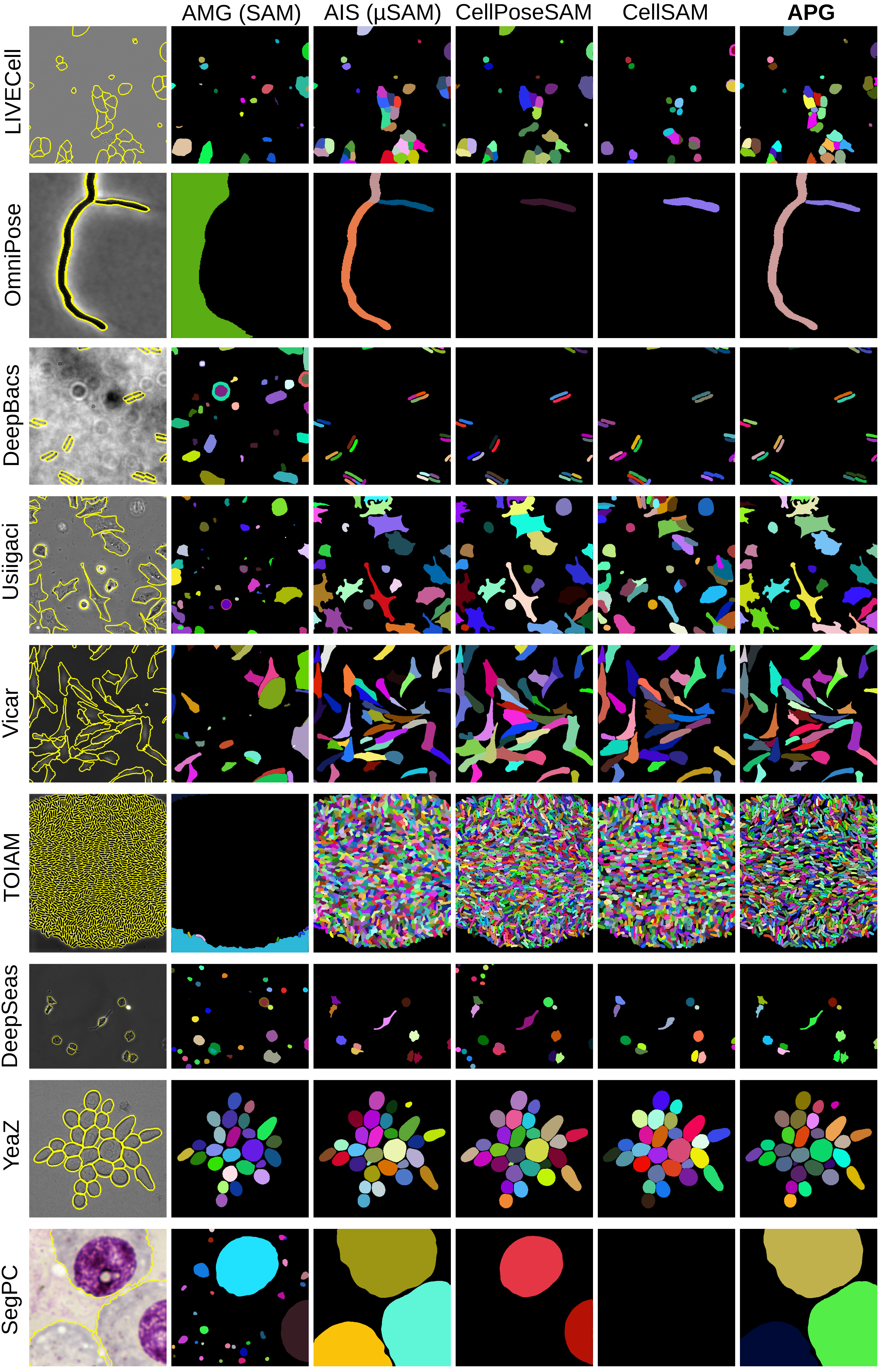
Figure 5: Qualitative results for all label-free microscopy datasets for cell instance segmentation.
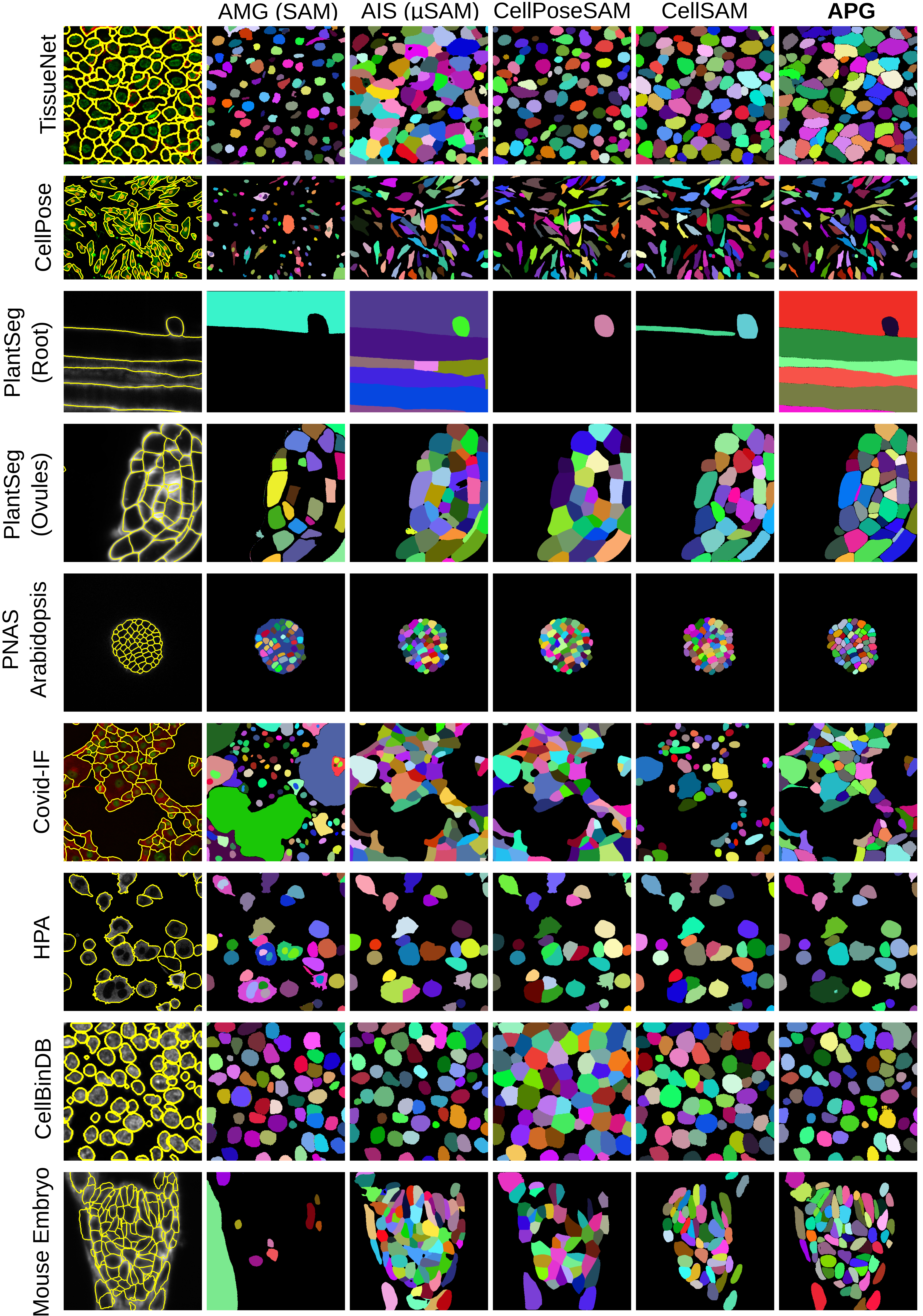
Figure 6: Qualitative results for all fluorescence microscopy datasets for cell instance segmentation.
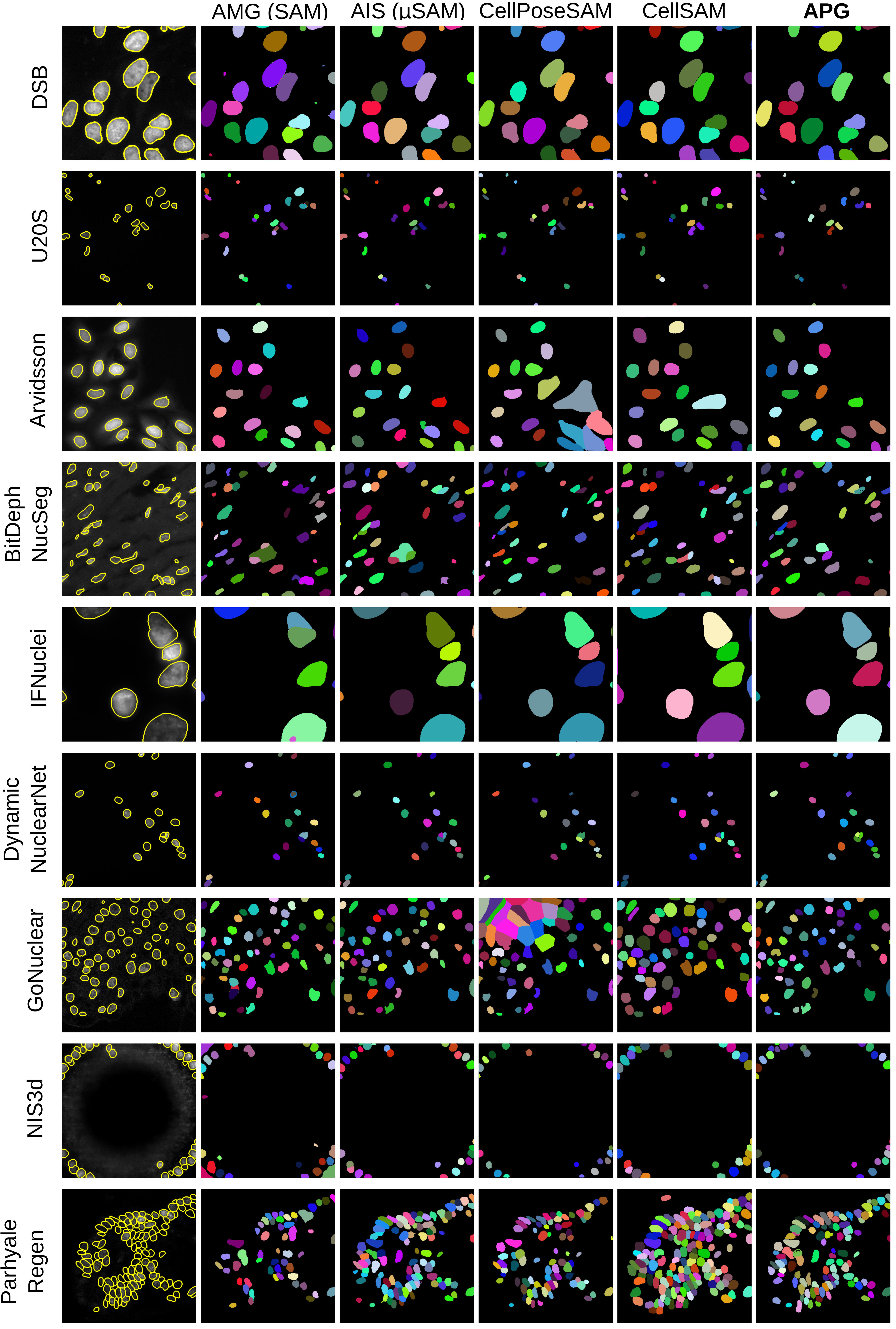
Figure 7: Qualitative results for all fluorescence microscopy datasets for nucleus instance segmentation.
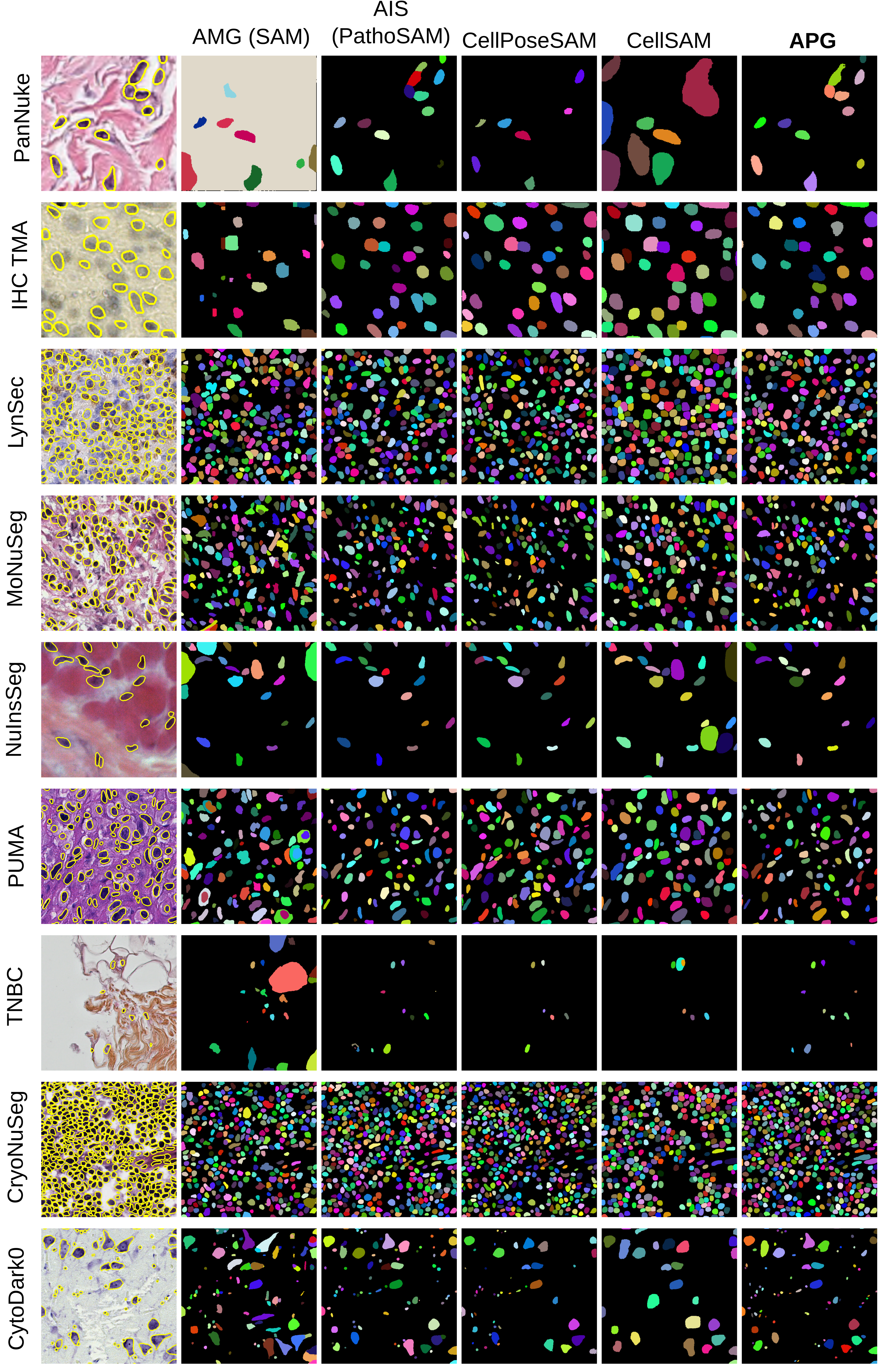
Figure 8: Qualitative results for all histopathology datasets for nucleus instance segmentation.
Limitations highlighted include exclusive evaluation in 2D (while several models support true 3D segmentation), and yet-unexplored strategies for box prompt derivation within APG. The paper suggests iterative prompt refinement and inclusion of example-based prompting in SAM3 as promising avenues for next-generation microscopy foundation models.
Conclusion
This work provides a rigorous, modality-diverse benchmark of foundation model adaptations for cell instance segmentation in microscopy, demonstrating that automatic prompt generation significantly enhances SAM-based model performance and rivals state-of-the-art dedicated models. The analysis establishes the need for continued domain adaptation and strategic prompting in foundation models, setting a clear direction for future advances in robust, generalized bioimage segmentation and practical deployment of vision foundation models in biological research.







