- The paper reveals that NAP clustering accumulates stress which, upon abrupt release, drives the stepwise nucleoid expansion observed in simulations.
- It employs a 3D coarse-grained polymer model that integrates dynamic NAP binding and explicit replication processes to mimic bacterial chromosome behavior.
- Findings indicate an optimal range of NAP interactions that maximizes segregation efficiency without the need for dedicated segregation machinery.
Nucleoid Clustering Regulates Stepwise Expansion and Segregation of Replicating Bacterial Chromosomes
Introduction
Accurate spatial organization and segregation of the bacterial chromosome occur robustly in the absence of a spindle apparatus. This paper elucidates how the interplay between DNA replication and nucleoid-associated protein (NAP) mediated clustering constitutes a non-equilibrium mechanism driving nucleoid dynamics, stepwise expansion, and chromosome segregation. Employing a coarse-grained, three-dimensional polymer model (PolyRep) that incorporates explicit replication mechanics and adjustable NAP-mediated interactions, the authors provide compelling evidence that clustering-mediated stress accumulation and release naturally result in experimentally observed phenomena such as stepwise nucleoid expansion and segregation during replication (2603.29719).
Model Specification and Implementation
The bacterial chromosome is modeled as a flexible ring polymer confined within a cylindrical geometry, representing the bacterial cell (Figure 1). NAP binding sites, which undergo stochastic post-translational switching between ON/OFF states, mediate local DNA bridging. The model allows systematic tuning of NAP-mediated attraction to interrogate the interplay between bridging strength, nucleoid compaction, and replication-driven dynamics. Replication initiation and fork progression are implemented via mobile firing factors that engage with defined origin sites, generating two replication forks that produce daughter strands as they traverse the template.
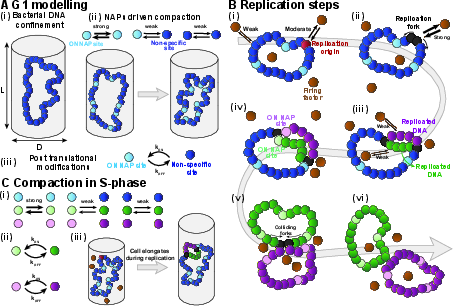
Figure 1: Coarse-grained polymer model of bacterial DNA, NAP-mediated clustering, and the dynamic ON/OFF regulation of NAP binding sites.
Cellular elongation is mimicked via time-dependent expansion of the confining cylinder, and the framework allows for detailed observation of nucleoid morphology and density both before and during replication.
Dynamics of G1 Nucleoid Clustering
Simulations of the unreplicated (G1) nucleoid reveal a highly dynamic entity. Even in the absence of replication, the nucleoid displays pronounced density fluctuations, spatial rearrangements, and transient clustering of NAP-binding sites. For strong NAP–NAP attraction cases, large, dynamic clusters drive drastic longitudinal nucleoid extension fluctuations, which are well-correlated with abrupt changes in cluster integrity (Figure 2). No persistent helical ordering emerges, in agreement with the absence of imposed loop extrusion or architectural constraints in the model.
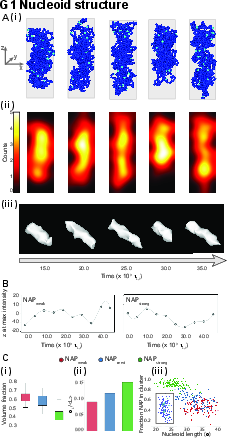
Figure 2: Time-lapse nucleoid evolution, density fluctuations, and cluster statistics in G1 phase for varying NAP interaction strengths.
Stepwise Nucleoid Expansion During Replication
Upon initiation of replication, the model recapitulates the experimentally-observed stepwise expansion dynamic. Rather than continuous elongation, the nucleoid exhibits intermittent contractions followed by rapid expansion events (Figure 3). These are temporally correlated with abrupt dissolution of large NAP-mediated clusters. This stick–release mechanism—where nucleoid contraction owing to clustering accumulates internal stress, which is subsequently relieved upon cluster breakup—accounts for the non-linear, jump-driven nucleoid extension observed in vivo. The statistics of extension jumps are invariant across a range of NAP binding strengths, indicating that replication progression, rather than cluster equilibrium, predominantly governs these events.
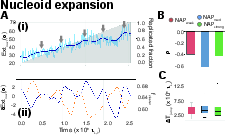
Figure 3: Representative time series illustrating stick–release nucleoid extension and correlation with NAP cluster fraction during replication.
Chromosome Segregation as an Emergent Non-equilibrium Phenomenon
The stick–release dynamics inherent to the replication-perturbed, clustering-driven nucleoid produce effective demixing of the two daughter chromosomes, quantified using a global segregation order parameter (Figure 4). The segregation exhibits non-monotonic time dependence: early replication yields partial demixing, which transiently decreases at mid-replication (correlated with fork dissociation and cluster reorganization), before recovering as nucleoid expansion and structural rearrangement resume. Segregation efficiency is maximized within a finite window of NAP interaction strengths; excessive compaction hinders segregation, while insufficient clustering precludes demixing. Consistently, the model predicts a positive correlation between the probability of dual-fork replication factory formation and large-scale nucleoid segregation.
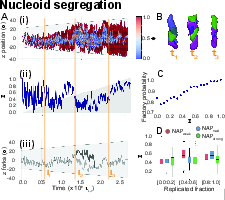
Figure 4: Kymograph of local segregation, time evolution of global segregation parameter, and snapshots illustrating demixing during stepwise nucleoid expansion.
Implications and Theoretical Consequences
This work demonstrates that the bacterial nucleoid constitutes an active, non-equilibrium polymer system in which internal protein-mediated attractions and replication-driven growth co-define spatial organization. The principal mechanism underpinning nucleoid stepwise expansion and segregation is a cycle of stress accumulation via NAP-mediated bridging and stochastic stress release upon cluster dissolution. Segregation arises without explicit segregation machinery, but only under a finely balanced interaction regime.
The results position the bacterial chromosome within the broader paradigm of actively-driven soft matter systems, where internally-generated forces (replication) and controlled cohesion (NAP bridging) dictate mesoscale structure and organization. Notably, the model predicts that replication factory formation and segregation are emergent, rather than imposed, phenomena and directly arise from the interplay of clustering and internal DNA synthesis.
Limitations and Future Directions
The model's minimal ingredients capture essential nucleoid physical phenomena but do not reproduce experimentally observed helical nucleoid ordering, likely due to the omission of SMC complex-driven loop extrusion and supercoiling. Incorporation of such features, along with multifork replication and macromolecular crowding, is expected to enhance predictive power and biological fidelity. The framework provides a generic foundation for simulation-driven investigation of chromosome behavior in both wild-type and mutant cells lacking molecular segregation machinery.
Conclusion
Nucleoid-associated protein-mediated clustering, modulated by post-translational ON/OFF switching and perturbed by ongoing replication, can account for stepwise nucleoid expansion and robust segregation of bacterial chromosomes in the absence of dedicated segregation motors. The results substantiate a theoretical foundation for chromosome organization as an emergent, non-equilibrium phenomenon controlled by protein–DNA interactions and replication-driven stress dynamics, with far-reaching implications for the understanding of active polymer architectures in cellular contexts (2603.29719).



