- The paper demonstrates a novel autonomous agent, Latent-Y, that executes de novo drug design campaigns with lab-validated nanomolar affinities.
- It integrates subagents, bioinformatics tools, and generative models to perform epitope discovery, structural analysis, candidate generation, and rigorous quality filtering.
- The approach achieves up to a 56-fold acceleration over traditional workflows, showcasing adaptive reasoning and cross-species binder design capabilities.
Latent-Y: Autonomous Agentic Framework for De Novo Drug Design
Overview and Key Contributions
"Latent-Y: A Lab-Validated Autonomous Agent for De Novo Drug Design" (2603.29727) presents an end-to-end agentic platform capable of autonomously executing full biologics design campaigns—covering literature extraction, structural analysis, candidate generation, and computational and wet-lab validation—initiated from natural-language prompts. Latent-Y orchestrates complex workflows typically requiring domain experts, leveraging a tight integration with both the Latent Labs Platform and the Latent-X2 generative model for atomic-level antibody design. Distinctively, the study demonstrates experimental validation, with hit rates and nanomolar affinities across diverse setting, and evaluates the capacity for adaptive reasoning, tool extension, and collaboration.
Agent Architecture and Workflow Design
Latent-Y is architected around a general-purpose, provider-agnostic agent harness leveraging frontier LLMs for long-context planning, tool invocation, and dynamic reasoning. The system manages context transfer through an orchestrator-subagent paradigm: specialized subagents execute analytical tasks such as epitope identification, hotspot analysis, and quality filtering. Central to the platform is model context protocol (MCP), which orchestrates tool access (bioinformatics databases, sequence analysis utilities, structural alignment modules) and Latent-X2 inference calls.
Upon receiving a prompt—ranging from open-ended functional goals to structured design constraints—Latent-Y autonomously (or semi-autonomously, with human inputs) executes a workflow comprising:
- Extraction of biological context (e.g., from literature databases)
- Target and epitope inference through multimodal data integration and reasoning
- Design space exploration using Latent-X2, followed by iterative refinement and prioritized exploitation of productive regions
- Quality assurance: diversity clustering, liability assessment, cross-database sequence filtering
- Transparent trace logging for all reasoning steps, tool invocations, and decisions
Together, these enable highly parallelizable, expert-equivalent campaign execution, and support on-the-fly adaptation based on both intermediate results and user steering.
Autonomous Epitope Discovery and De Novo Antibody Design
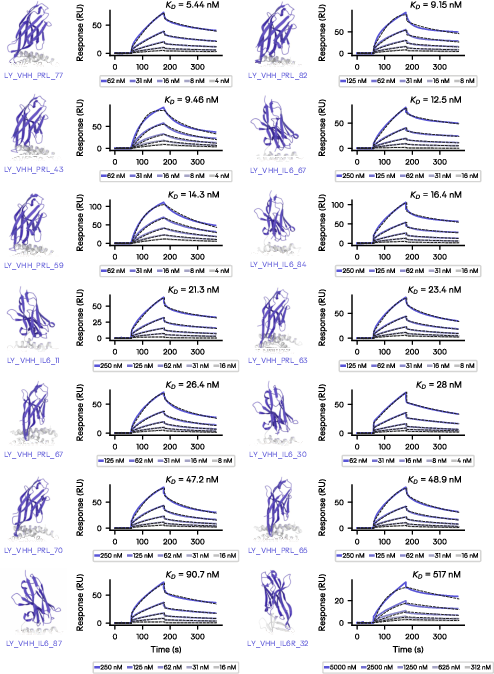
Latent-Y was evaluated on fully autonomous epitope discovery campaigns against six therapeutically significant targets, including IL-6, IL-6R, and PRL. Campaigns were initiated with minimal prompts specifying only the high-level objective. For all validated targets, the agent successfully identified functionally relevant epitopes consistent with desired mechanisms of action, sometimes choosing structurally unconstrained or previously uncharacterized regions.
Experimentally, the system achieved binders with robust hit rates and impressive affinities:
The average per-target hit rate ranged from 1–17%. The best binders from each campaign displayed diverse frameworks and CDR lengths, reflecting effective navigation of the combinatorial design space.
Agent Reasoning and High-Fidelity Traceability
A critical aspect of Latent-Y's design is comprehensive reasoning traceability. Each campaign maintains a full log of the agent's reasoning, tool usage, subagent calls, and feedback integration, with the capacity for both interruption and redirection by human users. For example, in the IL-6 campaign, the system iteratively rerouted based on computational screening feedback, accentuating productive epitope configurations.

Figure 2: Condensed trace of Latent-Y's autonomous campaign to design disruptive IL-6 binders, illustrating complex tool chains and iterative logic steps.
Autonomous Capability Extension: Cross-Species Binder Design
A unique demonstration involved cross-species binder design against TNFL9 homologues (human and cynomolgus). Here, Latent-Y was tasked with solving a translational challenge: generating a VHH capable of dual reactivity across species with divergent mutations and incomplete structural data. Notably, the agent autonomously developed a custom generative method, implemented from a one-line natural language description, with only high-level expert steering for biological plausibility and logic verification (e.g., to avoid reward hacking in inaccessible trimer configurations).
Experimentally, Latent-Y identified three cross-reactive VHHs (out of 40 tested) passing high-throughput SPR, with hits observed for both human and cyno targets.
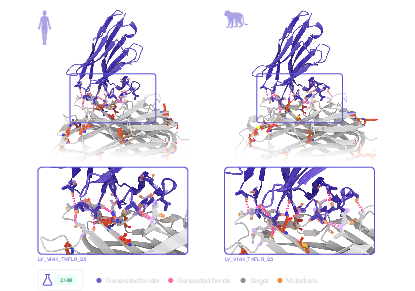
Figure 3: Structures of autonomously designed cross-species VHHs binding both human and cynomolgus TNFL9, with sequence divergence highlighted.

Figure 4: Condensed reasoning trace for collaborative cross-species binder design, evidencing agent-initiated tool augmentation and campaign diversification.
This capability for self-modification and real-time toolset extension is critical for therapeutic settings requiring rapid customization and surrogate targeting.
Literature-Inferred Design and Prompt Benchmarking
Latent-Y was systematically benchmarked across 21 therapeutic targets selected from peer-reviewed literature, requiring extraction of site and mechanism details solely from the provided publications. With a fixed budget (10,000 samples), the agent achieved epitope identification accuracy of 100%, with ≥24 computationally passing binders generated in 76% of campaigns post-QA.
The agent's effective reasoning strategy entailed initial high-diversity exploration, followed by targeted exploitation as productive frameworks were discovered. On the blood-brain barrier crossing (hTfR) campaign, experimental validation yielded a hit rate of 28% (11/40 designs).
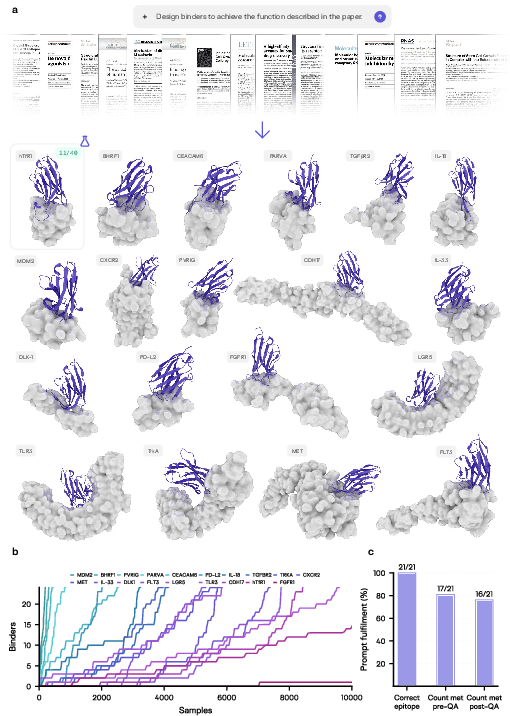
Figure 5: Latent-Y-generated de novo VHH complexes across 19 therapeutic targets, all derived from only publication context; wet-lab hit rates reported for hTfR.
Acceleration of Expert Workflows and Practical Impact
Across all settings, Latent-Y delivered a 56-fold acceleration over expert-alone workflows, as established by structured interviews with computational biologists (median experience: 9.8 years). The greatest acceleration was observed in reasoning and information gathering stages, compressing weeks to hours per campaign.
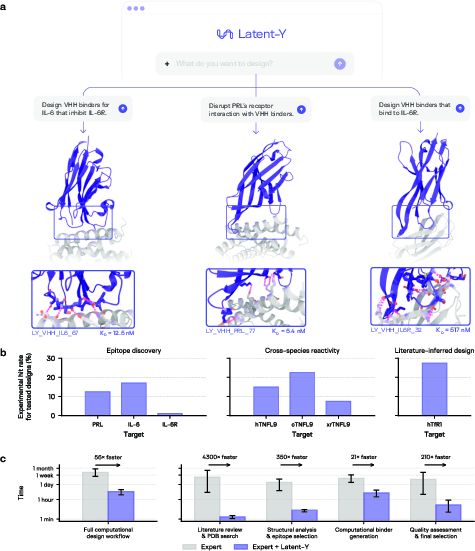
Figure 6: Latent-Y summary of epitope discovery and cross-species design campaigns, including workflow acceleration analysis and per-stage time breakdown.
These metrics underscore the practical impact: by abstracting the expert workflow, the platform substantially increases the achievable throughput and parallelism of discovery organizations.
Theoretical Implications and Future Directions
Latent-Y establishes a concrete realization of an AI scientist paradigm in molecular biology—capable not only of following pre-scripted pipelines but of enabling autonomous hypothesis generation, adaptation, and, crucially, extension of its own computational methods. The strong performance validates agentic architectures for multi-modal scientific discovery and underscores the feasibility of fully AI-driven candidate generation workflows.
The work also surfaces multiple open research directions:
- Full experimental closure (e.g., real-time feedback retraining and closed-loop laboratory automation)
- Modality expansion—Latent-Y is extensible to macrocyclic peptides and mini-binders, with additional experimental validation in progress
- Autonomous discovery—the agent’s ability to uncover novel mechanisms or propose entirely new biology remains nascent; further integration with wet-lab robotics and large-scale screening is requisite for this advance
Integrating robust traceability and safe human override, Latent-Y provides a blueprint for responsible deployment of highly autonomous agents in therapeutic design.
Conclusion
Latent-Y demonstrates that LLM-driven agentic systems, with tight integration of domain-specific generative models and bioinformatics toolchains, can directly produce lab-validated de novo antibody binders from free-text prompts. The agent operates in both purely autonomous and collaborative settings, extending its own computational repertoire as contexts demand. Reporting hit rates, nanomolar affinities, and substantial acceleration of expert workflows across various campaign types—including demanding scenarios such as cross-species binder design and literature-only epitope inference—this work marks a significant step toward automated, scalable, and auditable AI scientists for drug discovery and protein engineering.