- The paper proposes the Perturb-and-Restore framework that synthesizes high-fidelity abnormal chromosome images using diffusion-based restoration and energy-guided sampling.
- The paper demonstrates substantial performance gains with sensitivity improvements up to 40% and F1-score increases of 37% under extreme imbalance across over 260,000 chromosome images.
- The paper validates the framework through detailed ablation studies and blinded user evaluations, confirming enhanced anomaly detection in clinical cytogenetics.
Simulation-driven Structural Augmentation for Imbalanced Chromosomal Anomaly Detection
Introduction
Structural chromosomal anomaly detection is a critical task in clinical genetics, underpinning the diagnosis of inherited disorders and cancer. The inherent rarity and diversity of abnormal chromosomes introduce extreme, long-tailed data imbalance, severely impeding the performance of deep learning–based automated karyotyping pipelines. The paper "Perturb-and-Restore: Simulation-driven Structural Augmentation Framework for Imbalance Chromosomal Anomaly Detection" (2604.00854) addresses this by introducing a simulation-based augmentation pipeline that leverages prior domain knowledge and generative diffusion models to synthesize diverse, high-fidelity abnormal chromosome images, coupled with an adaptive energy-based sampling strategy for effective anomaly detection under extreme class imbalance.
Limitations of Conventional Approaches
Existing deep anomaly detectors and augmentation frameworks are unsuitable for chromosomal imaging due to two interrelated constraints: (1) authentic structural abnormality samples are prohibitively rare and heterogeneous, resulting in highly skewed class distributions; (2) most generative augmentation pipelines are generic and fail to capture chromosomal banding patterns and nuanced structural variants, as they rely on fully supervised training and substantial abnormal data, which is unavailable in clinical cytogenetics. Consequently, standard resampling techniques and re-weighted loss functions yield models that overfit to prevalent normal patterns, while methods tailored to specific anomalies or simple synthetic procedures lack generalization and diversity.
The Perturb-and-Restore (P&R) Framework
The framework is designed around two core components: Structure Perturbation and Restoration Simulation (SPRS) and Energy-guided Adaptive Sampling (EAS).
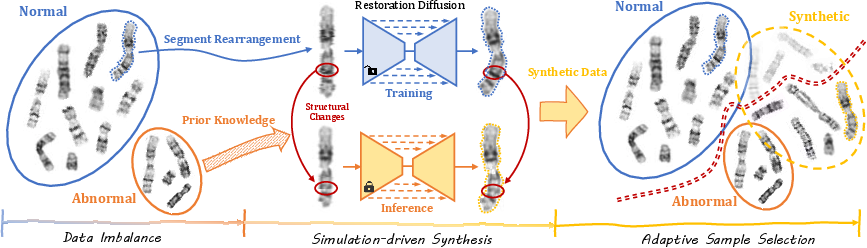
Figure 1: The P&R pipeline simulates diverse anomalies from normal chromosomes using guided perturbations and diffusion-based restoration, then selects high-quality synthetic anomalies with an energy-guided strategy.
Structure Perturbation and Restoration Simulation (SPRS)
SPRS exploits prior genetic knowledge to model abnormal chromosomes as perturbed rearrangements of normal banding patterns. A detailed segmentation of each chromosome is performed by extracting its skeleton and partitioning the axis into aligned rectangular patches. Structural variants—deletions, duplications, inversions, and translocations—are simulated through operations on this sequence of patches. To mitigate the introduction of unrealistic artifacts in highly curved or complex chromosomes, a medial axis cosine (MAC) score is computed, and only samples with high linearity (MAC > 85) are accepted for perturbation.
The synthetic rearranged chromosome images exhibit discontinuities and degraded content. Restoration is operationalized as an image-to-image mapping using a mean-reverting score-based diffusion model. The diffusion network is trained on paired (original, rearranged) chromosomes to learn the image manifold of realistic structures, allowing the system to restore perturbed outputs without abnormal labels. The restoration process enhances edge continuity and perceptual plausibility, yielding high-fidelity abnormal chromosomes.
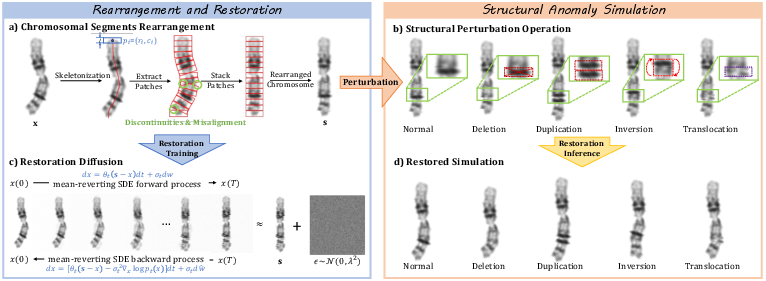
Figure 2: The SPRS module extracts and perturbs skeletal patches of chromosomes, then employs diffusion-based restoration for artifact-free abnormal sample synthesis.
Energy-guided Adaptive Sampling (EAS)
To ensure that only realistic, informative synthetic anomalies contribute to detector training, EAS computes an energy-based anomaly score for all samples: normals are mapped to low energy, abnormals (real/synthetic) to high energy. An energy margin loss enforces explicit separation. During training, EAS dynamically samples synthetic anomalies whose energy statistics closely align with real anomalies, using an adaptive recall threshold. This regime exposes the detector to difficult, high-quality abnormal patterns, improving robustness and generalization under severe imbalance.
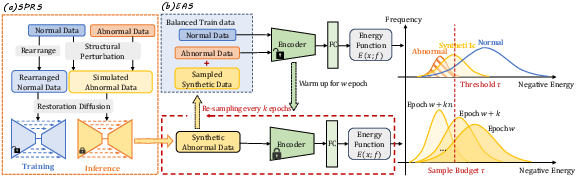
Figure 3: Illustration of the complete P&R training framework, combining simulation-driven synthetic data with energy-driven dynamic sampling.
Experimental Evaluation
Dataset Construction
An extensive benchmark is introduced, including over 260,000 chromosome images spanning 24 categories, with only 4,242 annotated abnormal samples (imbalance ratio averaging 179.87:1). The synthetic pipeline generates hundreds of thousands of high-fidelity abnormal chromosomes as augmentation data, substantially enriching the minority class.
Across both ResNet18 and ResNet50 backbones, P&R consistently achieves large gains over classical and recent long-tailed learning and cytogenetics-specific baselines. Key improvements include:
- Average sensitivity (Sen) improvement of +8.92 (ResNet18) and +7.11 (ResNet50).
- Mean F1-score improvements exceeding +13.4.
- Substantial gains in abnormal precision (+4 to +8.9).
Radar chart analysis across all chromosome classes demonstrates that P&R achieves improved detection in almost every category, including those with imbalance ratios >200.
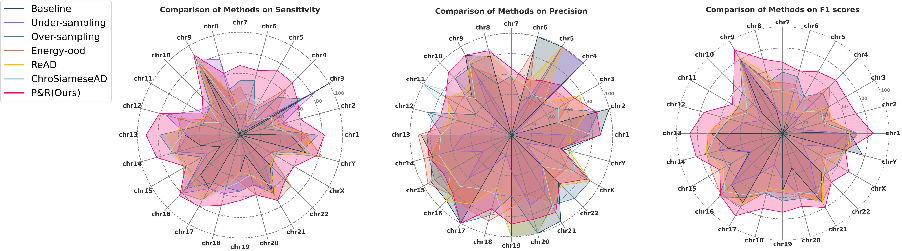
Figure 4: Radar chart showing that P&R provides superior sensitivity, precision, and F1-scores across 24 chromosome categories.
Ablation studies confirm that synthetic samples without restoration contribute little value, while the diffusion-based restoration module is crucial for utilitarian synthetic anomaly distribution. The energy-guided sampling schedule consistently further enhances all key metrics, particularly when source synthetic images are imperfect.
Generalization analyses show that introducing diffusion-restored synthetic data (SYN*) considerably boosts the performance of a range of long-tailed learning and anomaly detection baselines. Sensitivity improvements of nearly 40% and F1 increases of 37% are observed for standard cross-entropy-trained detectors.
Qualitative and Distributional Analyses
Energy score visualizations during training illustrate that EAS gradually sharpens the abnormal–normal class boundary, even under extreme imbalance. By the end of training, clear separation is consistently achieved for all data partitions.
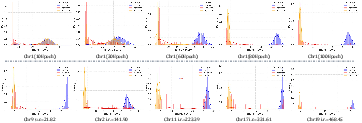
Figure 5: Evolution of energy score distributions, revealing the network's increasing capacity to distinguish between normal and synthetic/real anomalous data.
Diffusion-based restoration is quantitatively validated with both fidelity (PSNR/SSIM) and distributional alignment (KID) to normal and abnormal chromosome manifolds. The resulting synthetic images are frequently labeled as real by both domain experts and non-experts in blinded user studies, confirming the visual and structural realism of the generative process.
Practical and Theoretical Implications
The P&R framework provides a generalizable, annotation-efficient architectural paradigm for rare event modeling in clinical imaging domains characterized by high intra-class diversity and sparse anomaly labels. It demonstrates that simulation-guided augmentation, when regularized by explicit energy-driven alignment, can induce robust anomaly detection models even at imbalance ratios exceeding 200:1. Integrating prior domain knowledge with SDE-based diffusion decouples synthetic data quality from annotation bottlenecks, while dynamic fitness screening (EAS) ensures continued sample utility throughout training.
Practically, P&R enables automated karyotyping and cytogenetic analysis systems to generalize beyond common abnormalities, facilitating improved diagnostics and potentially enabling rare disease discovery from limited cohorts. Theoretically, these results suggest that label-free perturbation–restoration and energy-aligned sampling comprise a potent approach for general imbalanced anomaly detection, particularly where label scarcity precludes conventional generative or discriminative schemes.
Future Directions
Potential extensions include more refined integration of biological priors—such as chromosome-specific abnormality frequencies, spatial constraints, or disease semantics—into the perturbation simulation phase. Incorporation of multi-modal data (e.g., FISH or sequence-based features) may further expand model capability. The general paradigm can be adapted to other rare abnormality detection problems in medical imaging, genomics, and industrial inspection, especially where structurally coherent synthetic augmentation is essential for model generalization.
Conclusion
The Perturb-and-Restore framework constitutes a significant advancement for imbalanced structural anomaly detection in cytogenetics. By fusing explicit simulation of abnormality with diffusion-driven restoration and energy-guided adaptive sample selection, it establishes a template for robust, label-efficient learning under extreme class imbalance. The methodology is generalizable and offers a foundation for further AI-driven innovations in clinical genomics and rare disease detection.




