- The paper introduces a novel framework that decouples spatial and temporal dynamics, enabling fully parallelizable spiking state-space models with biologically inspired constraints.
- It partitions state-space dynamics into orthogonal neuron and synapse layers using a Multi-Transmission Loop to emulate realistic neural microcircuits.
- Empirical results on benchmark datasets demonstrate superior accuracy and parameter efficiency, highlighting the benefits of integrating biological regularization.
Parallelized Hierarchical Connectome: A Spatiotemporal Recurrent Framework for Spiking State-Space Models
Introduction and Motivation
The Parallelized Hierarchical Connectome (PHC) framework introduces a fundamentally novel approach to recurrent sequence modeling by reconciling two previously irreconcilable properties: spatiotemporal recurrence with lateral feedback and O(log T) temporal parallelizability. Conventional State-Space Models (SSMs), including S4, S5, LRU, and Mamba, leverage diagonal state transition structures to enable parallel associative scans; this design sacrifices intra-step spatial connectivity, limiting sequence models to purely temporal recurrence. Biologically inspired Spiking Neural Networks (SNNs) and RNNs support complex lateral and feedback communication but are non-parallelizable due to their dense recursive dependencies.
The PHC framework departs from both paradigms. It explicitly partitions state-space dynamics into orthogonal temporal and spatial components, enabling the integration of biologically grounded constraints such as Dale's Law, adaptive spiking (ALIF), short-term plasticity (STP), strict connectome topology, and reward-modulated spike-timing-dependent plasticity (R-STDP) within a scalable, parallelizable architecture suitable for mainstream sequence modeling benchmarks.
Architectural Framework: Spatiotemporal Decoupling and the Multi-Transmission Loop
At the core of PHC is the Intra-Step Spatiotemporal Decoupling principle, which separates conventional state-space evolution into a strictly diagonal, cell-autonomous Neuron Layer (NL) and a sparse, structured Synapse Layer (SL). Whereas conventional SSMs replicate layerwise depth (L) via stacking independent blocks and MLPs (Figure 1A), PHC collapses this into a hierarchical, region-partitioned neuron population where all communication—feedforward, lateral, and feedback—takes place via a shared, biologically constrained connectome.
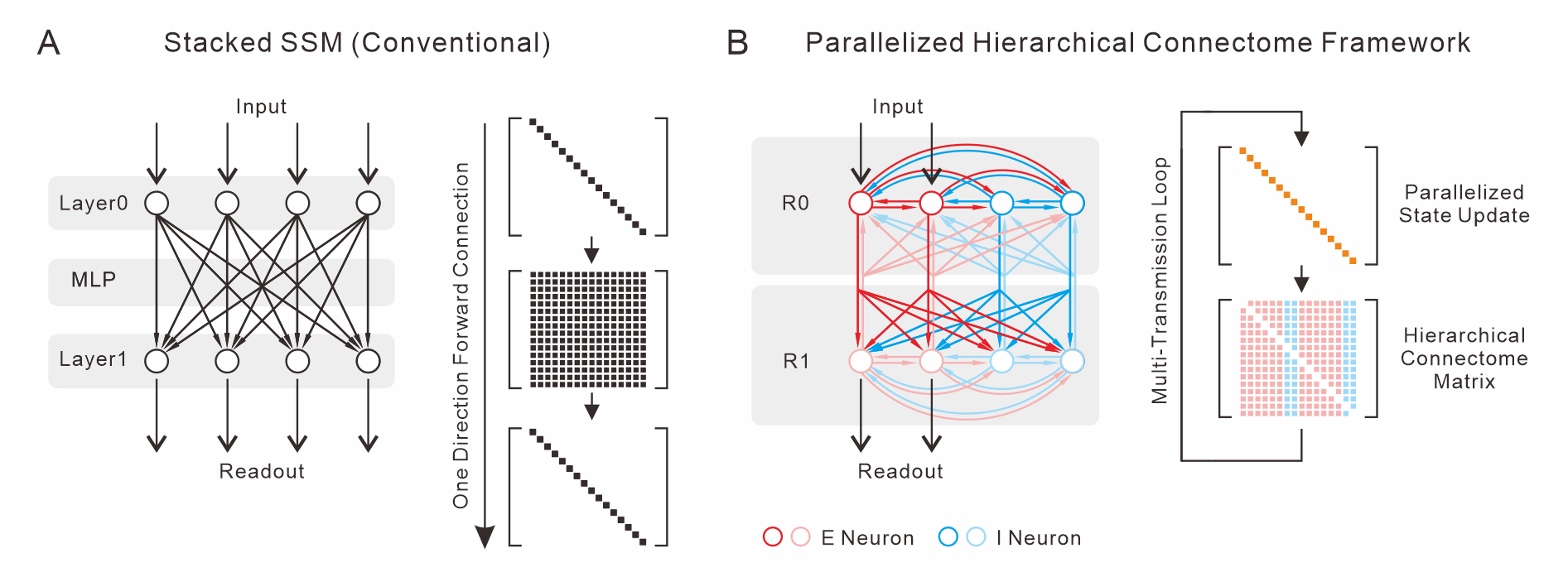
Figure 1: Structural comparison of conventional stacked SSMs (A) with PHC (B), highlighting lateral/feedback connectivity and hierarchical neuron partitioning.
Signals propagate within each temporal window by circulating multiple times through the NL-SL pipeline—a Multi-Transmission Loop—allowing recurrent spatial aggregation along the connectome before proceeding to the next input token. This recursion emulates biological microcircuits, where information routes through a fixed physical connectome, in contrast to the deep stacking paradigm.
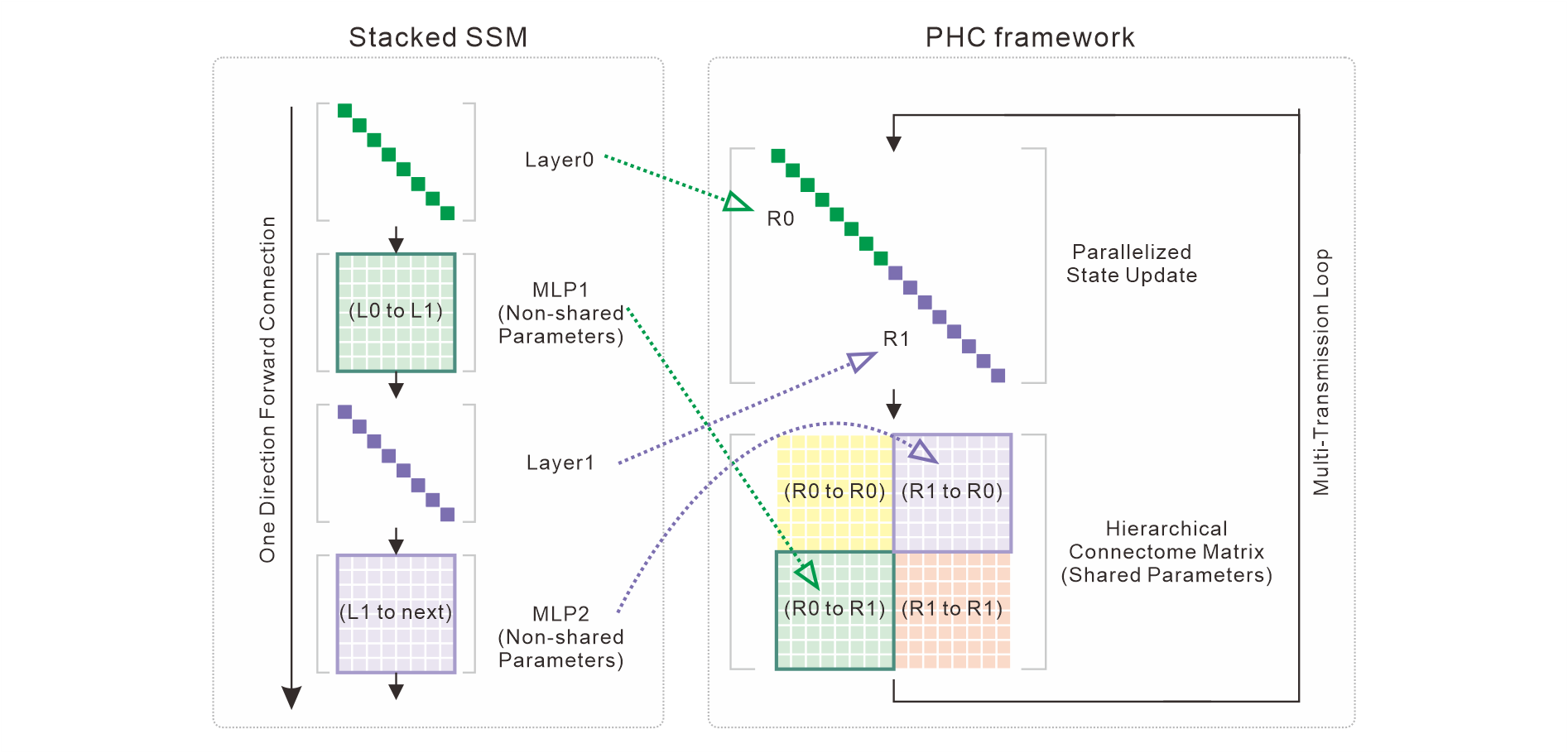
Figure 2: Isomorphism of standard stacked SSMs and the PHC architecture. Layer stacking is replaced by region partitioning, MLPs by connectome sub-blocks with Dale's Law and topology constraints.
This design supports fully parallelized training and inference via log-domain prefix scans for all dynamical components, including adaptive thresholding, membrane dynamics, and STP.
PHCSSM: A Biophysically Constrained Spiking State-Space Model
The instantiation of the PHC framework, PHCSSM, realizes the above architectural principles in a fully spiking SSM that enforces all of the following constraints:
- ALIF neural dynamics: Intrinsic adaptation is realized via learnable, elementwise leaky integration and adaptive spike thresholds using sequential parallel-scan recurrences.
- Short-term plasticity (STP): Implemented using the Tsodyks–Markram model, supporting both facilitation and synaptic depression through additional affine recurrences.
- Dale's Law and E/I partitioning: Synaptic weights are sign-constrained, segregating excitatory and inhibitory populations, and further pruned by a region-wise topology mask.
- Hierarchical Connectome Topology: Each neuron is assigned to a region, and the sparsity/connectivity is controlled at the block level, with explicit feedforward and feedback projections.
- Online R-STDP: Genuine binary spikes enable native spike-timing-dependent plasticity modulated by a scalar reward signal derived from downstream loss/accuracy, operating external to gradient descent.
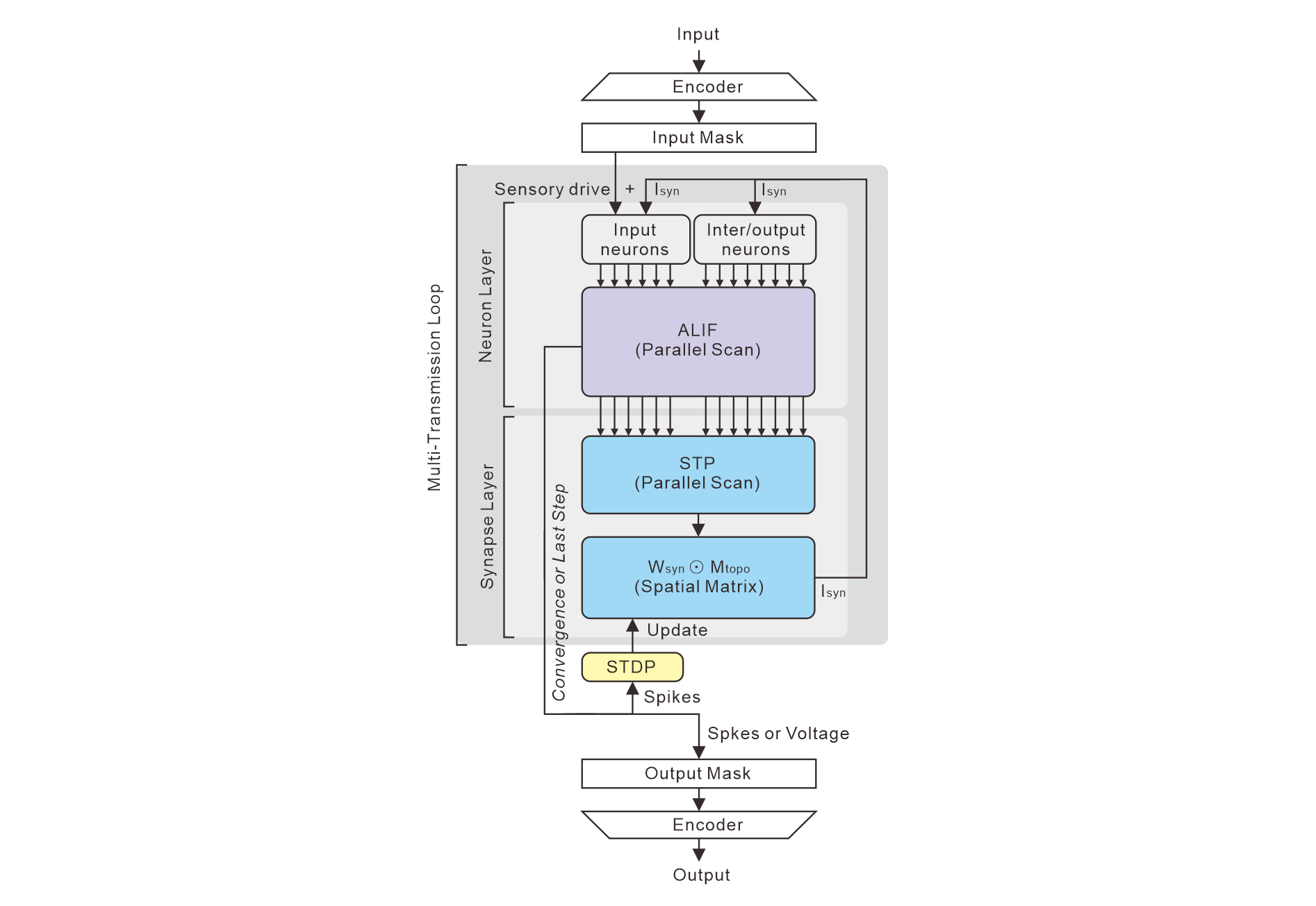
Figure 3: Detailed PHCSSM signal flow in the forward pass, showing encoding, masked sensory injection, iterative ALIF and STP scans, and biologically constrained matrix transmission.
The full NL and SL computation is executed in parallel across the temporal axis, with spatial interaction depth decoupled from sequence length.
Empirical Evaluation
PHCSSM is evaluated on the six-dataset UEA Multivariate Time Series Classification Archive under standard protocols. Key results include:
- SCP2: PHCSSM achieves 59.3% mean test accuracy, surpassing the previous best SSM, LinOSS-IMEX (58.9%), and exceeding all baseline models, including Mamba and S5.
- MotorImagery: Achieves 53.7%, outperforming Mamba (+6.0pp), S6, and Transformer.
- EigenWorms: Attains 83.9% accuracy with only 2,701 parameters, surpassing five strong SSM baselines.
Across all datasets, PHCSSM matches or exceeds several unconstrained models while requiring one to two orders of magnitude fewer parameters (1,748–9,485 vs. tens/hundreds of thousands), confirming the parameter efficiency conferred by weight sharing and biological regularization.
Ablation experiments substantiate the non-redundant contribution of each biological constraint. Disabling ALIF, STP, Dale's Law, or R-STDP each degrades accuracy (e.g., –12.3pp on SCP2 for ALIF removal), and removal of Dale's Law/online learning increases training variance, demonstrating strong regularization.
Theoretical and Practical Implications
The PHC paradigm demonstrates that biological constraints are not a performance bottleneck for parallel SSMs and, in fact, act as stabilizing inductive biases, enhancing both optimization regularity and parameter efficiency. By decoupling spatial recurrence from temporal propagation, PHCSSM introduces a new architectural axis—transmission depth M—enabling deep logical processing over fixed parameter budgets without layer stacking.
Pragmatically, these design principles permit:
- Integration of realistic microcircuit phenomena (STP, Dale's Law) in efficient GPU-compatible sequence models.
- Seamless transition from parallel training to sequential (RSNN) simulation without post-hoc conversion.
- Incorporation of empirical connectome data (mouse/human brain atlases) for functional digital twins.
- Natively supporting reward-modulated plasticity unavailable to continuous models, which may facilitate lifelong/online learning.
This framework directly supports applications in physiological signal decoding, neuroscience simulation, and energy-efficient AI via neuromorphic deployment.
Limitations and Future Directions
Current evaluation is restricted to sequence classification and region sizes up to R=2 (D=16-64). Future work should address:
- Scaling PHC to larger connectomes and more complex input-output tasks (e.g., language, speech, multivariate regression).
- Automatic connectome/region structure discovery from biological or task-specific priors.
- Quantitative analysis of parallel-to-sequential mapping error to support faithful neuromorphic implementation.
Incorporating additional biological mechanisms such as chemotactic modulation, dendritic nonlinearity, or synaptic homeostasis is also viable within this framework.
Conclusion
The PHC framework and its PHCSSM instantiation establish that biologically principled architecture, explicit spatiotemporal recurrence, and full temporal parallelizability are simultaneously achievable in SSMs (2604.01295). The resultant models offer state-of-the-art accuracy on benchmarks while maintaining extreme parameter parsimony and full support for structured, interpretable connectivity and online plasticity. This work uncovers a systematic route for leveraging biological priors as architectural design principles in mainstream deep learning.


