- The paper introduces a novel MMM mediation framework that jointly models high-dimensional exposures, mediators, and outcomes using a multivariate LSEM approach.
- It employs elastic net regularization and cross-fitted prediction techniques to ensure consistency, asymptotic normality, and effective variable selection.
- Empirical studies, including Alzheimer’s disease applications, demonstrate accurate recovery of mediation paths and strong out-of-sample predictive performance.
Introduction
The paper "High-dimensional Many-to-many-to-many Mediation Analysis" (2604.02886) introduces and formalizes the many-to-many-to-many (MMM) mediation framework. This model generalizes classical mediation analysis by simultaneously considering multivariate exposures, mediators, and outcomes—each of which may be high-dimensional. The MMM framework enables rigorous estimation, inference, and variable selection in causal pathways linking multiple exposures to multiple mediators and, in turn, to multiple outcomes. It provides both theoretical guarantees (consistency, asymptotic normality, and error bounds for estimators) and empirical validation through comprehensive simulation studies and a substantive application to genetic-neuroimaging-cognitive relationships in Alzheimer's disease (AD).
The MMM mediation setting is modeled by a multivariate linear structural equation model (LSEM), allowing exposures x∈Rq, mediators m∈Rp, and outcomes y∈RT, with both q and p potentially exceeding sample size n. The model is:
mi=α⊤xi+ζ⊤zi+ϵi yi=β⊤mi+γ⊤xi+η⊤zi+ξi
where (α,β,γ,ζ,η) specify path-coefficient matrices, and zi denotes covariates. This configuration yields a matrix-valued global indirect (mediation) effect αβ mapping exposures to outcomes via all mediators.
A joint penalized estimation procedure is employed, leveraging elastic net regularization in each LSEM stage to enforce sparsity and enable scalable estimation and selection in high-dimensional regimes. The estimation procedure is summarized in Algorithm 1 of the paper and includes cross-fitted out-of-sample prediction, where mediation parameters allow outcome prediction using only exposures and covariates, even when mediators are unavailable.
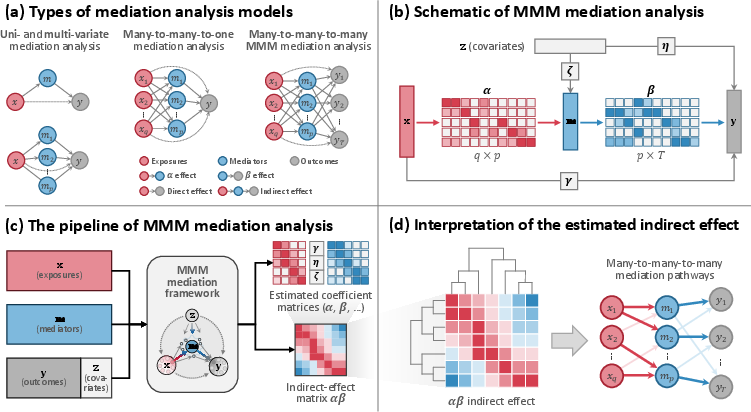
Figure 1: Schematic of the MMM mediation framework: multivariate exposures, mediators, and outcomes linked via structured paths, with pipeline for coefficient estimation and effect interpretation.
Theoretical Properties
The MMM estimators for direct and indirect effects are shown to be consistent and asymptotically normal under mild regularity conditions. Error bounds for mean squared estimation error are derived explicitly. The theoretical analysis extends elastic net model selection and sign consistency theorems to the high-dimensional multivariate pathway context, using the Elastic Irrepresentable Condition (EIC) adapted to simultaneous high-dimensional settings for exposures and mediators.
Explicit formulas for the identification and estimation of the matrix-valued natural indirect and direct effects are provided under potential outcomes notation, with extensions of sequential ignorability to the MMM causal setting. Entrywise asymptotic normality is established for estimated indirect (mediation) effects, supporting large-sample inference at the individual path level.
Simulation Studies
A comprehensive set of simulation experiments evaluates finite-sample parameter recovery, stability, Type I error behavior, and robustness to noise and sample size. These studies demonstrate:
- Highly accurate recovery of coefficient and indirect-effect matrices under block-structured ground truths, attaining clear separation of true nonzero and null mediation pathways.
- Bootstrap-based stability indices indicating reproducibility of results and resilience to sampling variability.
- Robustness to elevated noise and stability across varying sample sizes; indirect effect estimation is more sensitive due to the compounded nature of m∈Rp0.
- Empirical convergence of normalized estimation errors and parameter correlations to the ground truth at moderate-to-large m∈Rp1.
- Empirical distributions of selected indirect effects closely approximating normality, corroborating the limiting theory.
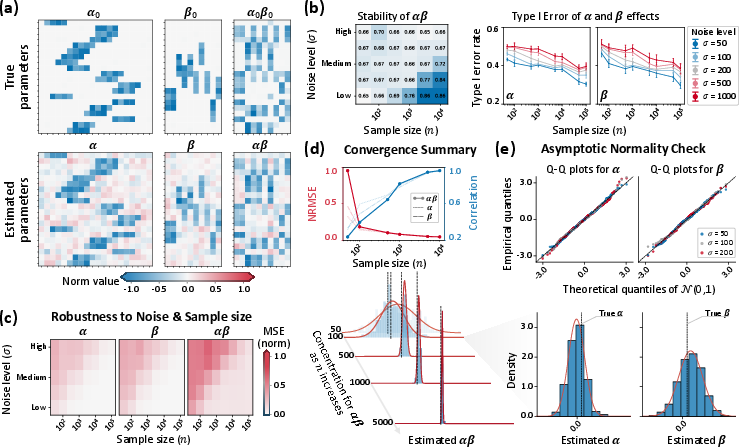
Figure 2: Heatmaps of true and estimated m∈Rp2, m∈Rp3, and m∈Rp4, with error convergence, stability, and empirical normality analyses across simulated regimes.
The MMM framework is applied to the Alzheimer's Disease Neuroimaging Initiative (ADNI) dataset to elucidate polygenic–brain–cognition pathways. The exposures are 688 genome-wide significant SNPs, mediators are 202 regional cortical thickness measures, and outcomes are 11 AD-related diagnosis and cognitive/behavior endpoints. The framework is able to:
- Recover interpretable and biologically coherent genetic–brain (m∈Rp5) maps, with dominant pathways spanning Default Mode, Control, Dorsal Attention, and Visual cortical networks, consistent with known AD vulnerability loci.
- Localize brain-to-cognition mediator effects (m∈Rp6) to DMN, temporo–parietal, and prefrontal circuits, matching areas repeatedly implicated in cognitive deterioration.
- Uncover a compact, structured genetic–neural–cognitive mediation network, identifying both convergent and divergent SNP–ROI–cognition pathways, notably funneling polygenic signals through AD-relevant cortical hubs to diverse cognitive outcomes.
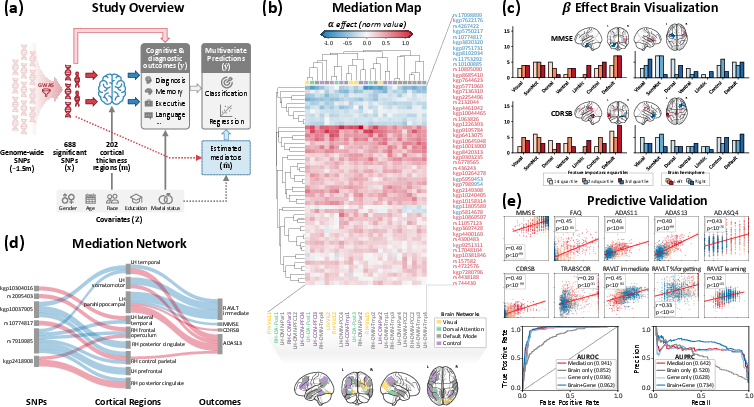
Figure 3: AD application—(a) experimental design, (b) estimated exposure–mediator effects, (c) spatial localization of mediator–outcome effects, (d) mediation network of strongest SNP–brain–outcome links, and (e) multivariate out-of-sample prediction performance.
In addition, out-of-sample prediction experiments reveal that mediator representations and path coefficients identified by MMM enable strong prediction of cognitive and diagnostic outcomes, even when only exposures and covariates are available at test time, thus validating reproducibility.
Implications and Future Directions
The MMM methodology directly addresses critical unmet needs in high-dimensional pathway analysis, where multiple correlated exposures, mediators, and outcomes interact. Practically, it yields interpretable variable selection and pathway identification, supports outcome prediction under real-world data constraints, and integrates seamlessly for scientific inference in complex biological systems such as imaging genetics.
The framework claims consistency, asymptotic normality, and statistical efficiency for indirect effect estimators; empirical results strongly support these claims. Compared to prior work restricted to univariate or separate multivariate exposures/outcomes, the MMM framework enables estimation and hypothesis testing on the entire path matrix in a single, coherent model.
Theoretically, future extensions include nonlinear and nonparametric MMM path models, incorporation of prior knowledge for biomarker or network selection (e.g., Bayesian variants), and longitudinal MMM mediation for dynamic pathway inference. Technically, integration with generalized estimating equations or PDE-based mixed-effects models could further expand MMM utility.
Conclusion
The MMM mediation framework provides a robust, theoretically justified, and empirically validated methodology for high-dimensional, multilayer mediation analysis. It facilitates variable selection, effect estimation, and outcome prediction in settings with multiple exposures, mediators, and outcomes. The approach is effective both in recovering interpretable structures in scientific data (as shown in complex neuroimaging-genomics-disease settings) and in rigorous statistical inference, opening avenues for more granular study of multivariate causal pathways in diverse high-dimensional scientific domains.


