- The paper introduces a global non-greedy optimization framework that jointly adjusts all tree splits to enhance classification accuracy.
- It employs a convex-concave surrogate loss function optimized via SGD, effectively reducing overfitting and tree complexity.
- Experimental results on benchmarks like MNIST and Protein validate improved generalization and efficiency over traditional greedy methods.
Efficient Non-greedy Optimization of Decision Trees
Introduction
The paper "Efficient Non-greedy Optimization of Decision Trees" presents an innovative approach to optimize decision trees by moving away from traditional greedy algorithms. Instead of optimizing each node locally and independently, the proposed methodology optimizes split functions across all tree levels simultaneously, utilizing a global objective. This approach is aligned with the principles of structured prediction with latent variables, aiming to form decision trees that generalize better and offer superior classification performance.
The research focuses on non-greedy decision tree induction, with binary classification trees as a primary example. Traditional decision tree algorithms operate in a local optimization mode, where nodes are split without regard for subsequent tree structure. This paper introduces a global optimization framework, linking decision tree formulation with latent structured prediction techniques, where each split in the tree is treated as a function that considers subsequent potential splits.
The key variables are the weight matrices and log-probability vectors at each node, which are optimized collectively rather than independently. This setup is framed as a quadratic programming problem whose complexity scales quadratically with tree depth — allowing feasible training of deeper trees.
Non-greedy Learning Methodology
One of the distinguishing features of this work is the use of a convex-concave upper bound on empirical loss to guide the optimization of the tree parameters. This surrogate loss function is efficiently optimized using Stochastic Gradient Descent (SGD). The authors mitigate the complexity arising from the exponential number of potential split combinations by modifying the surrogate objective for more manageable computational requirements.
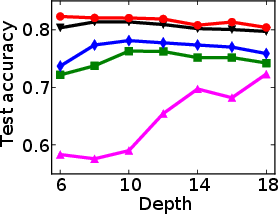
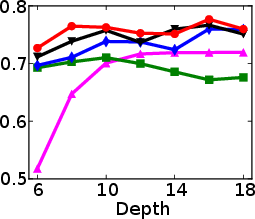
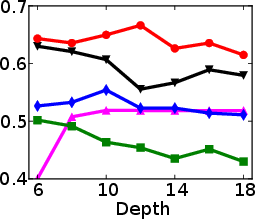
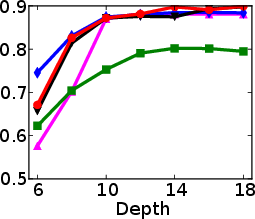
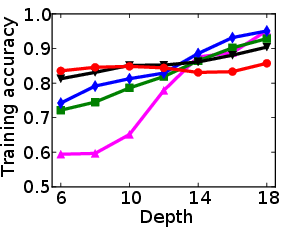
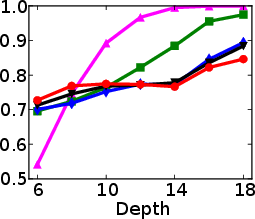
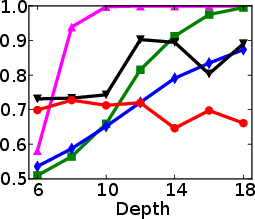
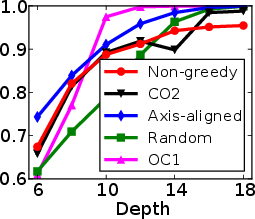
Figure 1: Test and training accuracy of a single tree as a function of tree depth for different methods. Non-greedy trees achieve better test accuracy throughout different depths. Non-greedy exhibit less vulnerability to overfitting.
This method's robustness is evident from its reduced susceptibility to overfitting compared to traditional greedy algorithms. By jointly optimizing the decision tree parameters, the paper demonstrates a decrease in final tree complexity and an enhancement in test accuracy.
Implementation
The implementation utilizes SGD with specific adjustments to ensure efficient traversal and optimization of leaf parameters. Implementation details include regularization techniques to prevent overfitting and measures to handle computational overhead associated with deep trees. The paper provides a foundation for adapting decision tree learning to incorporate kernel methods for potential applications in higher-order splits, expanding the utility of the proposed approach.
Experiments
The authors perform extensive experiments on benchmark datasets such as MNIST, Connect4, and Protein, proving the superiority of non-greedy induced trees over traditional greedy ones. The results underscore the enhanced generalization capacity and efficiency of the proposed algorithm, particularly for deeper trees and larger datasets. Notably, the method's efficacy is not limited by tree depth, showcasing consistent performance improvements across varying tree depths as depicted in the summary of their results.
Conclusion
This research provides a significant contribution to decision tree learning methodologies in machine learning. By pivoting from local, greedy optimizations to a more holistic loss function encompassing all tree levels, the authors furnish a path to deeper, more accurate, and computationally efficient decision trees. Future work could explore extending these techniques with kernel methods and applying them to a wider range of datasets and applications. This paper's insights could further drive advancements in domains where decision trees are the algorithm of choice due to their interpretability and efficiency.







