- The paper introduces a complex-valued neural network with a novel cardioid activation function to improve MRI fingerprinting accuracy and computational efficiency.
- The network directly maps complex MRI signals to tissue parameters like T1, T2, and B0, eliminating the need for time-consuming dictionary matching.
- Experimental results show reduced error rates and significant computational gains under noisy conditions, outperforming traditional methods.
Complex-Valued Neural Nets for MRI Fingerprinting
Introduction
Magnetic Resonance Fingerprinting (MRF) is a cutting-edge technique that enables the simultaneous quantification of multiple tissue parameters from complex-valued MRI signals using a single scan. Traditionally, these tissue parameters—such as T1 and T2 relaxation times—are identified using dictionary-based methods, which involve matching an observed MRI signal against a pre-computed set of signals with known parameters. However, this approach does not scale well with increased parameter dimensionality due to its computational inefficiency. The paper introduces a novel application of deep learning that leverages complex-valued neural networks to directly solve the inverse mapping problem in MRI fingerprinting, demonstrating improved accuracy over traditional methods through the adoption of complex calculus and a new activation function.
MRI fingerprinting can be understood as an inverse mapping problem, where the task is to deduce the original tissue parameters from observed MRI signals. The authors propose using a deep learning framework to model this inverse mapping as a nonlinear function. Unlike conventional methods, the proposed neural network approach does not require a pre-defined dictionary, thus significantly enhancing computational efficiency and scalability.
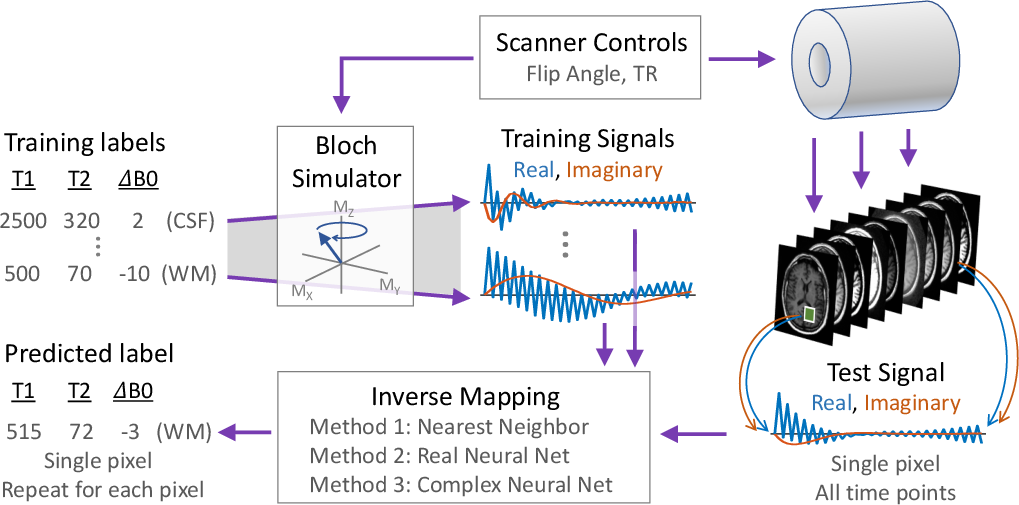
Figure 1: MRI fingerprinting as an inverse mapping problem, illustrating the process of inferring tissue parameters from MRI signals.
Complex-Valued Neural Network Design
The core contribution of the paper lies in developing a complex-valued neural network equipped with a new cardioid activation function, engineered to accommodate the unique characteristics of complex-valued MRI data. Despite the fact that complex-valued signals could be represented as two-channel real signals, this loses the intricate phase information inherent in complex algebra. The cardioid activation function respects this phase aspect of complex numbers by modulating the output based on the input phase, unlike conventional real-valued activation functions that operate independently on real and imaginary components.
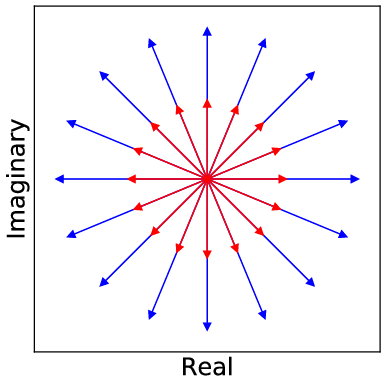
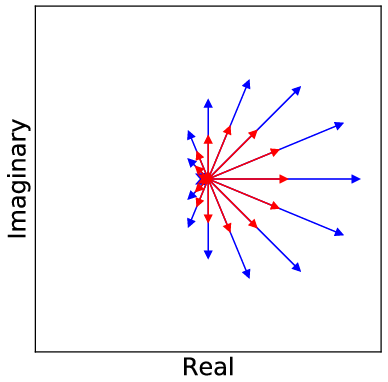
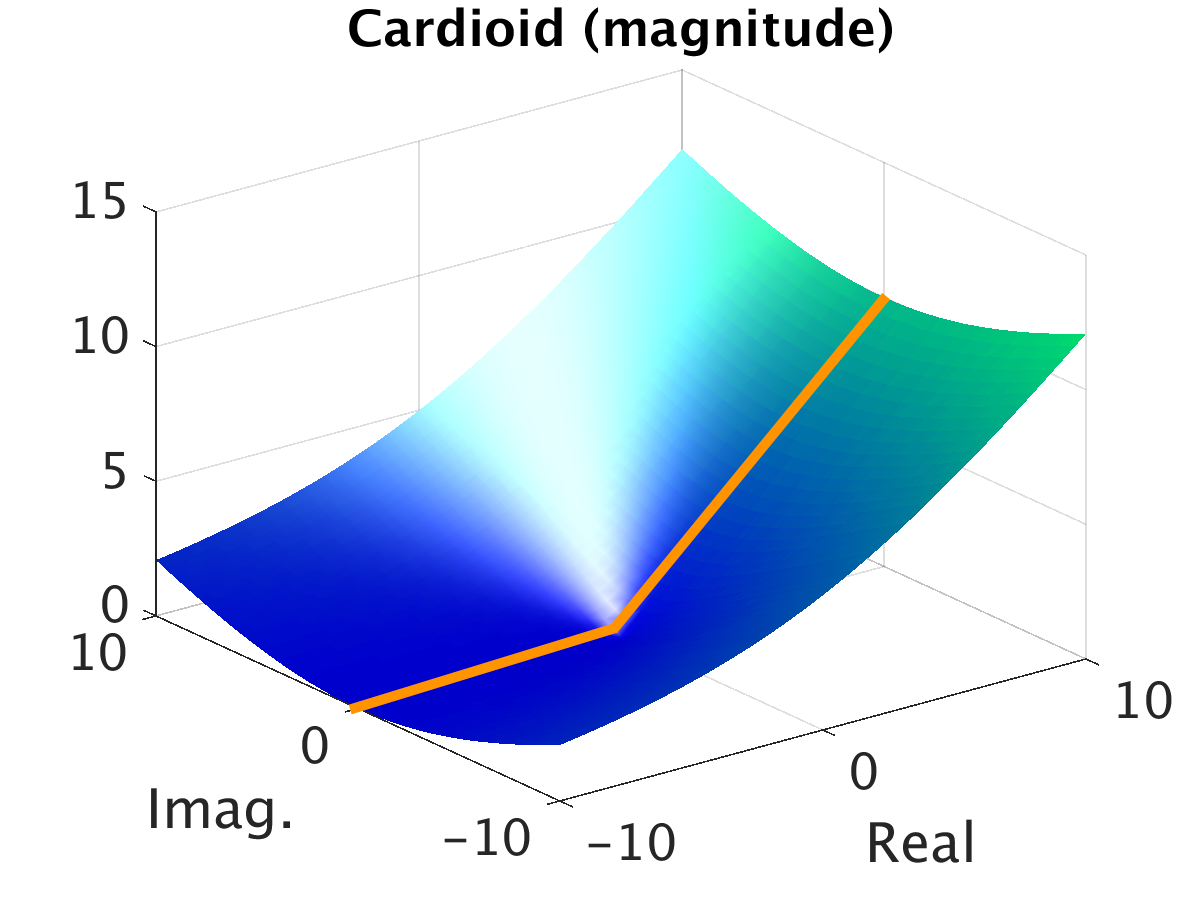
Figure 2: Visualization of the cardioid activation function on the complex plane, demonstrating its phase sensitivity.
The neural network architecture consists of a fully connected setup tailored for each parameter prediction (T1, T2, B0), ensuring robust learning of the inverse mapping from simulator-generated synthetic MRI data. The network is implemented within the Caffe framework, with adaptations to accommodate complex arithmetic through the use of CR calculus for optimization.
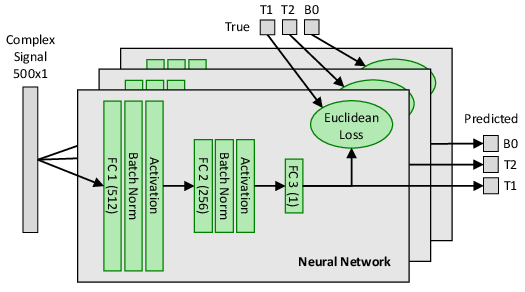
Figure 3: Fully connected neural network architecture used for mapping MRI signals to tissue parameter outputs.
Experimental Evaluation
Experiments utilize synthetic training data generated from Bloch equations and evaluate the proposed methods against a numerical MRI phantom. A noise model with peak signal-to-noise ratio (pSNR) 40 is used to simulate real-world conditions, enabling robust assessments of network performance against noise. The complex neural network with cardioid activation consistently outperforms the baseline nearest neighbor method and real-valued networks, particularly in the accurate recovery of T1 and T2 parameters.
The analysis of floating point operations (FLOPs) reveals significant computational gains, with complex networks requiring markedly fewer resources relative to dictionary-based approaches.
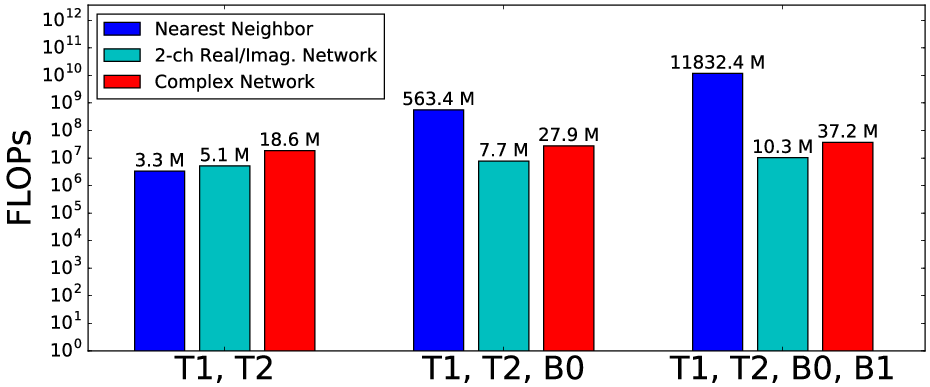
Figure 4: Comparison of floating point operations across different methods, highlighting the efficiency of the complex-valued neural net approach.
Results
The neural network approach substantially improves the prediction of T1 and T2 parameters, with a notable reduction in error rates compared to the nearest neighbor baseline. Complex-valued networks leverage the novel cardioid activation to enhance prediction accuracy, surpassing traditional methods and real-valued network counterparts, thereby underscoring the efficacy of respecting the complex nature of MRI signals.
For clean signals, the normalized root mean square error (NRMSE) results indicate significant precision improvements, with complex networks outperforming in terms of both accuracy and computational demand. Under noisy conditions, the trend persists, further validating the robustness of the proposed methodology.
Conclusion
This research delineates a novel approach for MRI fingerprinting using complex-valued neural networks, capitalizing on a tailored cardioid activation function. The results, reinforced by detailed experimental evaluations, indicate substantial advancements over existing methods in both accuracy and computational efficiency. The application of complex neural networks to MRI opens new horizons for efficient, scalable medical imaging solutions. Future research may explore further deployment of such networks, potentially extending towards full image reconstructions and accommodating additional MRI parameters.





