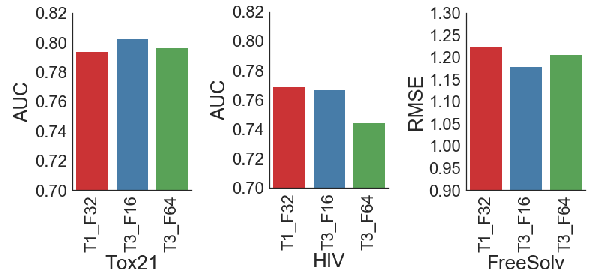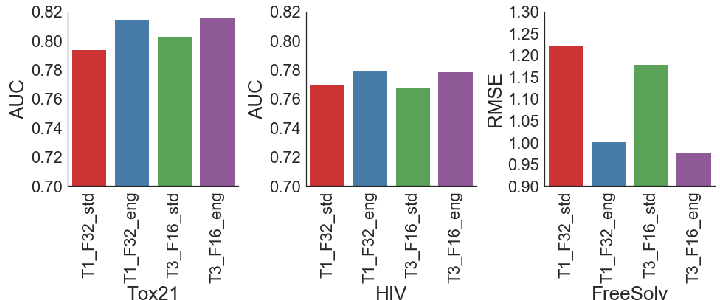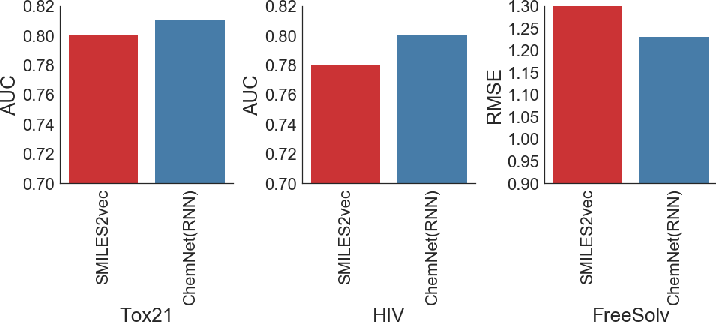- The paper introduces ChemNet, leveraging rule-based molecular descriptors for weak supervised learning to pre-train on large unlabeled chemical databases.
- It demonstrates significant improvements in predicting toxicity, activity, and solvation energy compared to models relying on engineered features.
- Experiments reveal ChemNet’s robustness across CNN and RNN architectures, indicating its potential in industrial and research applications.
Using Rule-Based Labels for Weak Supervised Learning: A ChemNet for Transferable Chemical Property Prediction
Introduction
This paper outlines the development and application of ChemNet, a deep neural network trained through weak supervised learning using rule-based knowledge. ChemNet aims to address the challenges inherent in chemical data, which tend to be fragmented and small in volume compared to other domains such as computer vision. Traditional DNNs in chemistry rely on engineered features, which might limit their potential representation capabilities, especially when domain knowledge is insufficient or inadequately developed.
Limitations of Current Approaches
The chemical sciences often face the challenge of limited labeled data, with large databases such as ChEMBL being only sparsely labeled. This stands in stark contrast to domains like computer vision where datasets like ImageNet offer extensive labeled data, enabling effective representation learning using raw data inputs in CNN models. Thus, leveraging representation learning in chemistry remains constrained by the available labeled data, necessitating alternative approaches.
ChemNet Design and Rule-Based Learning
ChemNet leverages rule-based knowledge encapsulated in molecular descriptors for weak supervised learning, facilitating the training of neural networks on large unlabeled chemical databases. These descriptors reflect historical research and encode chemical knowledge, making them suitable for generating inexpensive, consistent labels. ChemNet was pre-trained using the ChEMBL database and was designed to be network architecture agnostic, demonstrating efficacy across both CNN and RNN models.
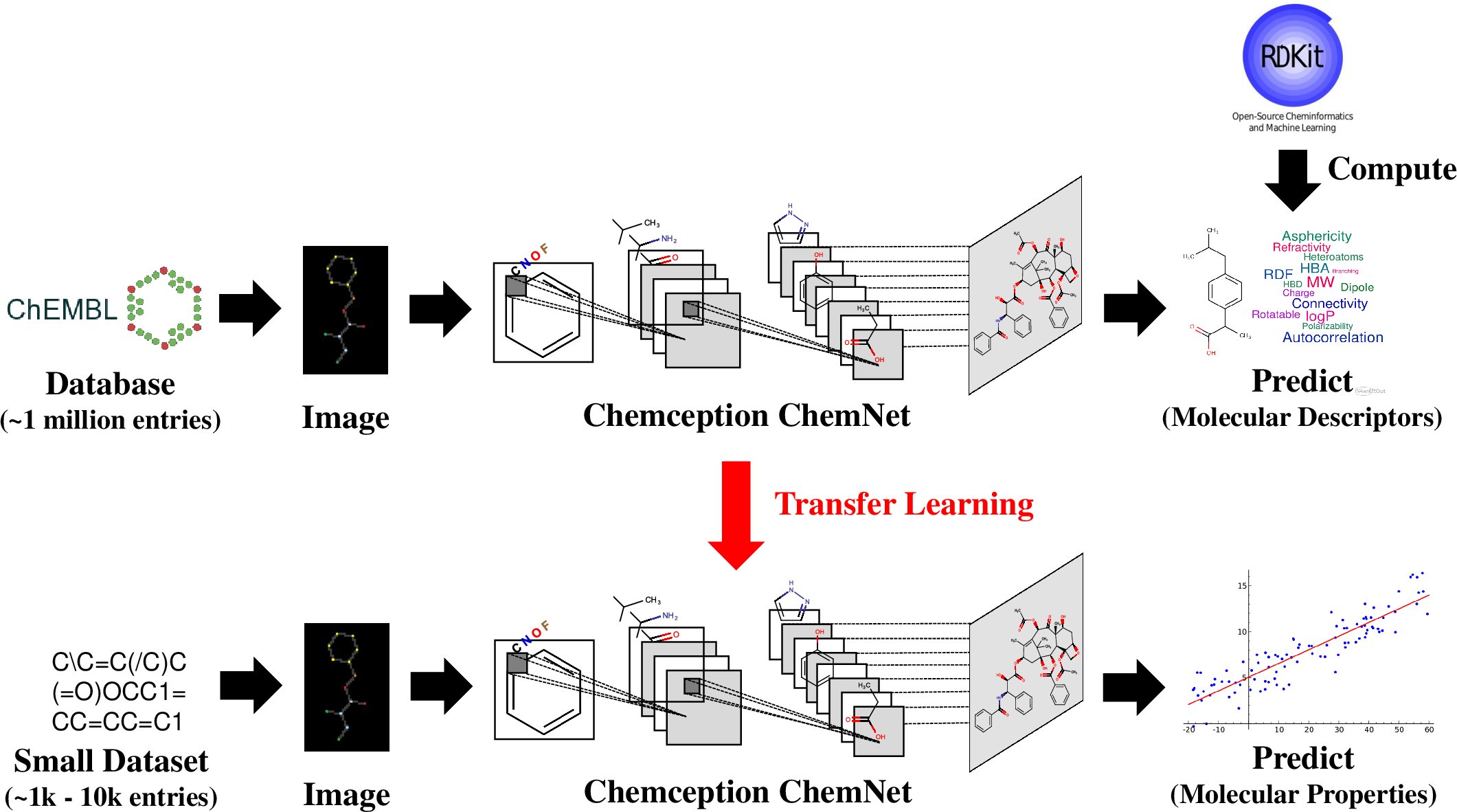
Figure 1: Schematic illustration of ChemNet pre-training on the ChEMBL database using rule-based molecular descriptors, followed by fine-tuning on smaller labeled datasets on unseen chemical tasks.
Experimental Evaluation
ChemNet exhibited superior performance in predicting chemical properties such as toxicity, activity, and solvation energy compared to traditional DNN models leveraging engineered features. Evaluations on datasets such as Tox21, HIV, and FreeSolv confirmed its generalizability to other chemical properties and emphasized its industrial applicability across sectors like pharmaceuticals and biotechnology.
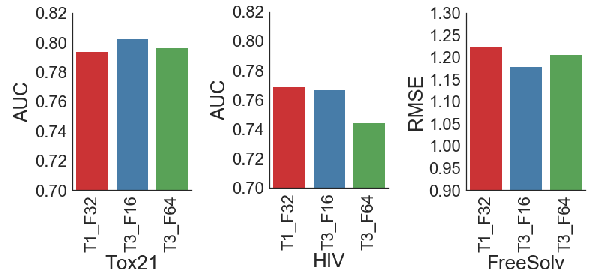
Figure 2: The ChemNet T3_F16 architecture generally had consistently better performance on the validation AUC/RMSE for toxicity, activity and solvation energy predictions.
Enhancements through Image Augmentation and Transfer Learning
The integration of augmented images in ChemNet training proved beneficial, consistently enhancing model performance across chemical tasks. Fine-tuning experiments revealed that freezing half of ChemNet's layers did not degrade performance, indicating the robustness of its learned chemistry-relevant representations.
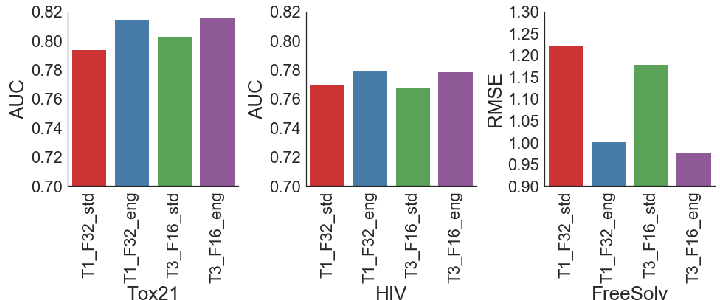
Figure 3: Using augmented images results in consistently better performance on the validation AUC/RMSE for toxicity, activity and solvation energy predictions.
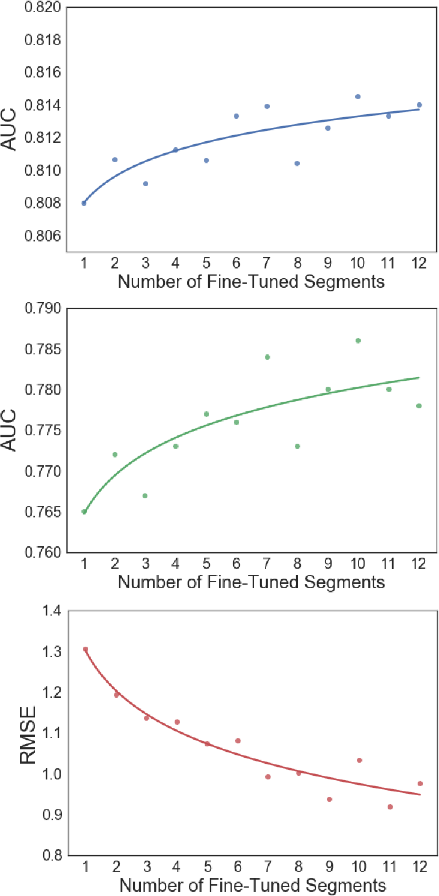
Figure 4: Fine-tuning beyond the 7th segment of the ChemNet T3_F16 architecture yield diminishing returns in performance improvement.
ChemNet demonstrated substantial improvements over earlier Chemception models and displayed competitive or superior performance against contemporary state-of-the-art models such as the ConvGraph algorithm. This highlights ChemNet's effectiveness in utilizing chemical domain knowledge combined with deep learning techniques for enhanced predictive power.
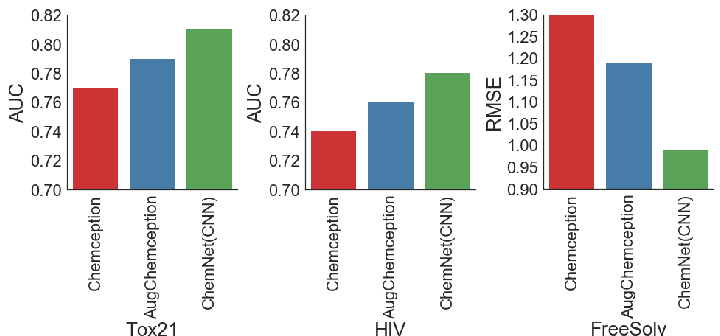
Figure 5: ChemNet (based on the Chemception CNN model) provides consistently better performance compared to earlier Chemception models.
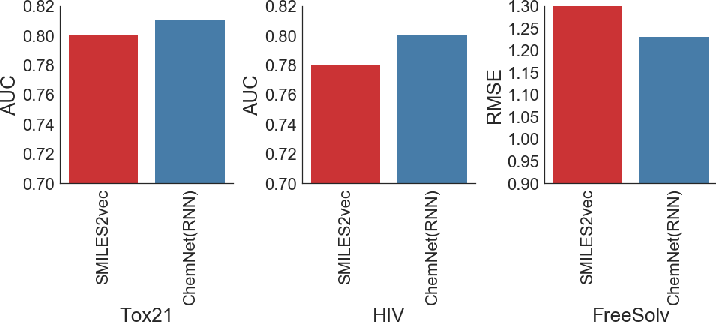
Figure 6: ChemNet (based on the SMILES2vec RNN model) provides consistently better performance compared to earlier SMILES2vec models.
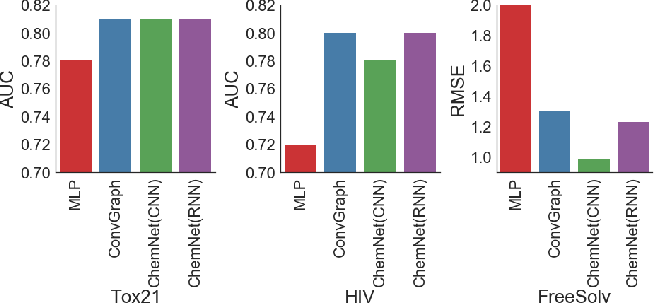
Figure 7: ChemNet consistently outperforms MLP models trained on engineered features and matches the performance of ConvGraph.
Conclusions
ChemNet represents a critical advancement in bridging the gap between traditional feature-engineering approaches and modern deep learning methodologies in chemistry. Its ability to yield superior performance across various tasks implies promising applications in industry-centric chemical predictions, potentially revolutionizing how chemical properties are predicted and utilized in research and development contexts. The rule-based weak supervised learning approach offers a template for analogous application in other domains, leveraging existing domain knowledge to enhance neural network training and predictions.
This work emphasizes the potential for incorporating domain-specific rule-based knowledge into the training frameworks of neural networks, suggesting pathways for future research that further capitalizes on these strategies across diversified fields.