- The paper introduces a robust 3-step method combining pre-alignment, parametric, and non-parametric registration, achieving a median MrTRE of 0.19%.
- It utilizes the NGF distance measure and Gauss-Newton iterative optimization in the affine registration step to refine image alignment effectively.
- Application on the ANHIR dataset demonstrates significant error reduction and runtime efficiency while ensuring smooth deformations without grid foldings.
Robust, Fast and Accurate: A 3-Step Method for Automatic Histological Image Registration
Introduction
The paper presents a robust and efficient three-step pipeline for registering histological images, particularly those that are differently stained. This method is crucial for digital pathology where precise alignment is necessary for analyzing serial tissue sections. The pipeline's efficiency is underscored by its runtime of 4 seconds and an impressive median relative target registration error (MrTRE) of 0.19%. This solution integrates a combination of pre-alignment, parametric registration, and non-parametric registration, optimizing the Normalized Gradient Fields (NGF) distance measure throughout the process.
Pipeline Description
The proposed registration pipeline processes histological images through three distinct steps, each building upon its predecessor to refine the registration accuracy.
Step 1: Pre-Alignment
The initial step involves automatic rotation alignment (ARA) to address arbitrary positioning of tissue slices post-manual processing. By applying a rigid transformation, the method focuses on translation and rotation adjustments. This preliminary transformation is derived using the center of mass of the images, optimized over several potential rotations within the [0,2π] range to minimize the NGF distance measure.
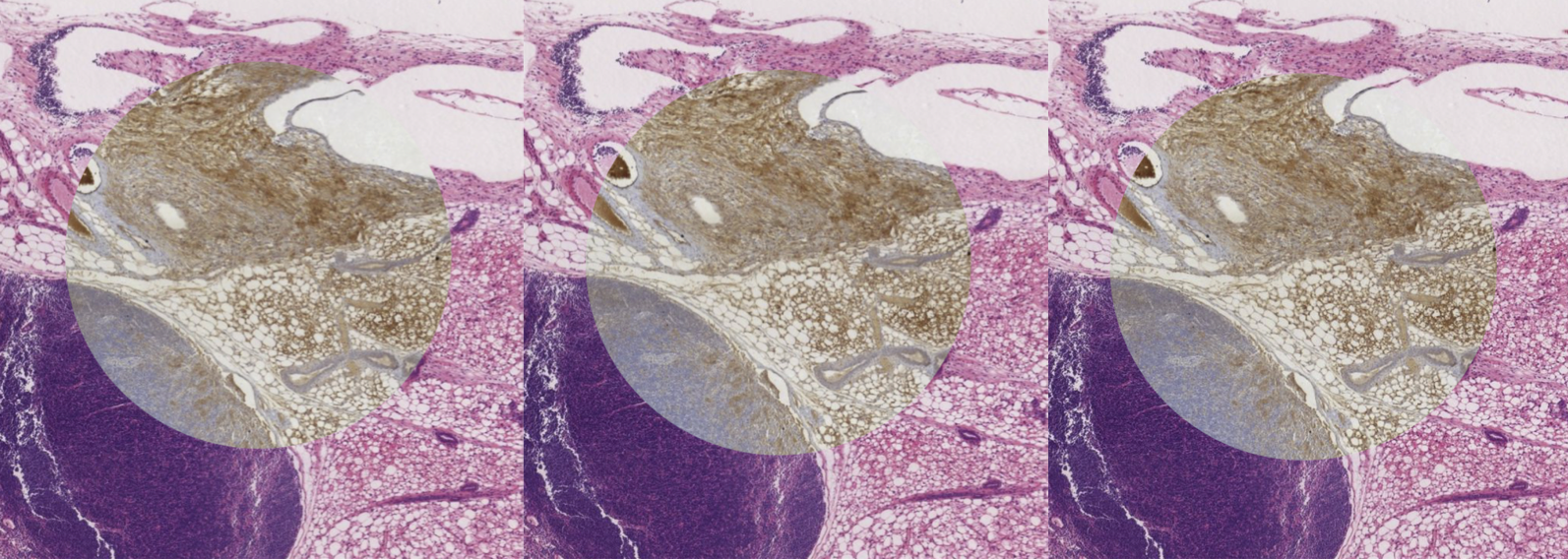
Figure 1: Spy-view of an image pair after pre-alignment, parametric and non-parametric registration (left to right).
Step 2: Parametric Registration
Building on the pre-alignment, the second stage utilizes affine transformations characterized by six degrees of freedom. The NGF distance measure is again minimized to adjust affine parameters using a Gauss-Newton iterative method, which begins with initial guesses derived from pre-alignment outputs, emphasizing further refining of the alignment on a coarse level.
Step 3: Non-Parametric Registration
The final step involves non-parametric registration, introducing additional flexibility through a displacement field. A regularization term is introduced to ensure smoothness and prevent topological changes. The optimization of the registration continues using a multi-level approach, employing curvature regularization to finely tune deformations. An L-BFGS approach further aids in minimizing local minima and computational demands.
The proposed method was evaluated using the ANHIR challenge dataset, which includes 230 image pairs. Post-registration, the method achieved an average MrTRE of 0.49% and a median MrTRE of 0.19%, demonstrating robust performance in reducing registration errors across nearly all cases. The comprehensive application of this pipeline results in significantly improved alignment without any grid foldings, validating its utility in a real-world setting.
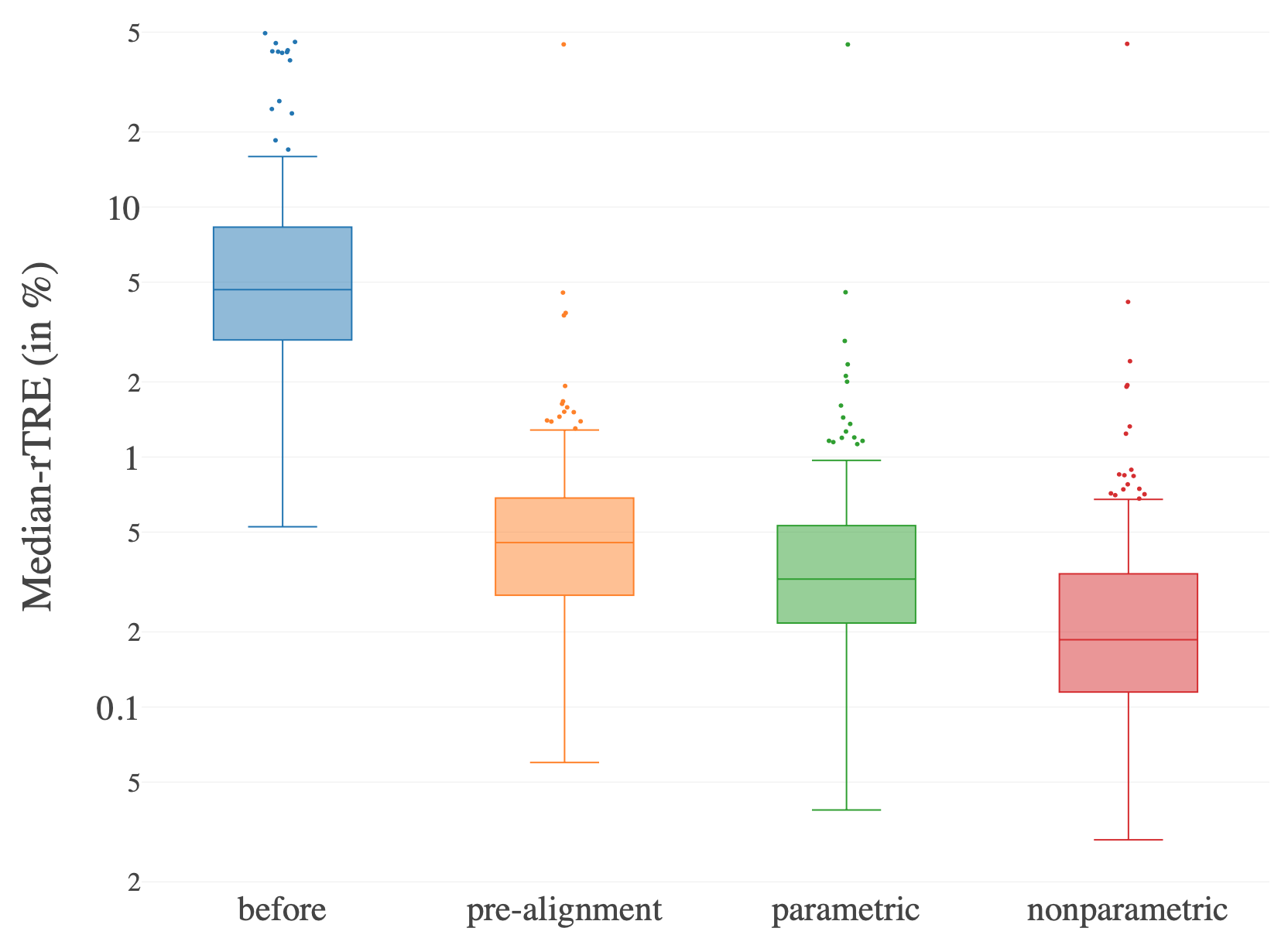
Figure 2: Histogram of the Median-rTRE measured on the training data $(N_{\text{pairs}=230)$ before alignment and after each registration step.
Conclusion
The 3-step histological image registration pipeline achieves robustness, speed, and high accuracy, effectively applying NGF distance measures throughout the process. This method stands out for its remarkable runtime and precise alignment capabilities, addressing computational challenges in multi-modal registration tasks extensively. Future developments could focus on further optimizing computational efficiency or exploring deeper learning integration to enhance robustness in more complex scenarios, potentially expanding its utility across other domains requiring image registration.

