- The paper introduces an end-to-end Instantiation-Net that reconstructs high-resolution 3D meshes of the right ventricle from a single 2D MRI image using a DCNN and GCN architecture.
- It achieves competitive performance with a mean 3D error near 2.2 mm, eliminating manual segmentation and complex parameter tuning.
- The study emphasizes real-time feasibilty for intra-operative use and suggests potential extensions to broader 2D-to-3D reconstructions.
Instantiation-Net: 3D Mesh Reconstruction from Single 2D Image for Right Ventricle
Introduction and Problem Statement
This work addresses the challenge of reconstructing detailed 3D cardiac anatomy from a single 2D projection, focusing on the right ventricle (RV) as captured by MRI. Traditional navigation in robot-assisted Minimally Invasive Surgery (MIS) relies on limited 2D images, complicating intra-operative 3D interpretation—especially for real-time dynamic navigation. Previous approaches, such as registration-based 3D-2D alignment and KPLSR-based shape instantiation, suffer from high computational cost, static modeling, and dependency on manual segmentation and parameter tuning. Recently, deep models with end-to-end learning for point cloud recovery were introduced, but lacked mesh connectivity, limiting their downstream clinical utility.
Instantiation-Net Architecture
This paper proposes Instantiation-Net, a fully automatic end-to-end system mapping a single 2D MRI image to a high-resolution, anatomically accurate 3D mesh suitable for surgical guidance. The framework integrates:
- DCNN feature extraction: A DenseNet-121 backbone (pre-trained on ImageNet) processes the 2D image and outputs a compact deep feature representation.
- Fully Connected transition: FC layers project and transform the DCNN features for mesh decoding.
- Graph Convolutional Network (GCN) mesh decoder: Four GCN layers perform hierarchical up-sampling and mesh vertex regression, preserving topological consistency and anatomical detail.
The workflow is visualized in (Figure 1).
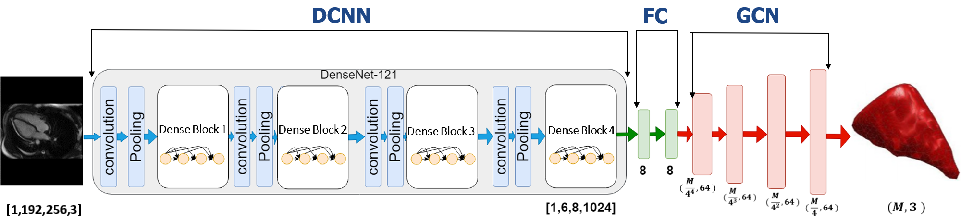
Figure 1: Intuitive block-wise architecture of Instantiation-Net, depicting DCNN, FC, and GCN components for end-to-end mesh reconstruction.
The structure enables learning both appearance-based features from images and graph-structured spatial transformations for mesh generation. Chebyshev polynomial-based GCNs enable computationally efficient spectral filtering on mesh topologies.
Experimental Design
Data Acquisition
The dataset comprises short-axis and long-axis MRI scans from 27 subjects (18 normal, 9 HCM), each sampled at 19–25 cardiac phases, yielding 609 paired 2D images and 3D meshes. Ground-truth meshes are obtained via manual semi-automated segmentation and meshing routines (e.g., Meshlab), ensuring vertex-level correspondence and consistent connectivity.
Training Protocol
The framework is evaluated using strict patient-specific leave-one-out cross-validation. The loss is defined as mean per-vertex L1 error, and optimization employs SGD with momentum and decaying learning rate. Inference time is minimized to approximately 0.5 seconds per case, fulfilling practical requirements for intra-operative deployment, although total training time and resource usage are higher than less expressive models.
Results
Vertex-wise Error Analysis
Vertex-level reconstruction errors are consistently distributed, with slightly higher error at the mesh apex (base of RV), attribut-able to topological sparsity and limited observational data for these regions in MR slices.
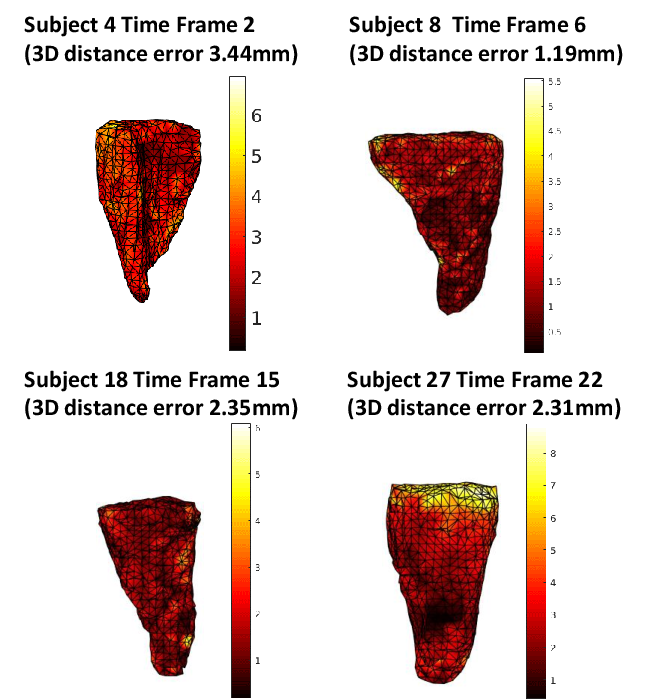
Figure 2: Vertex-level spatial error maps for four mesh reconstructions; intensity encodes mm error, highlighting spatially homogeneous performance except at mesh extremities.
Temporal Error Robustness
Frame-wise analysis over 12 patients demonstrates that Instantiation-Net maintains mean errors near 2 mm throughout the cardiac cycle. Outlier time-points (e.g., end-systole/diastole) exhibit higher error due to underrepresentation in training and boundary effects in temporal dynamics.
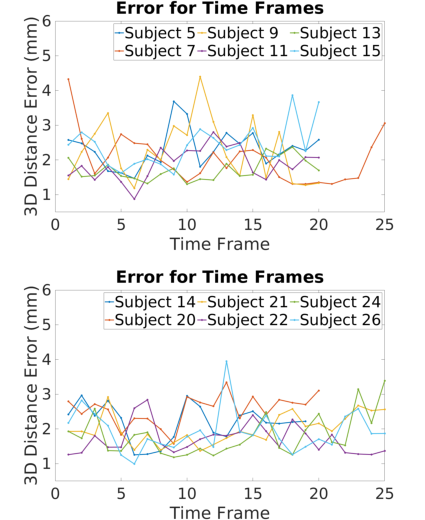
Figure 3: Mean 3D distance error per cardiac frame across 12 subjects, demonstrating stability and highlighting frames affected by "boundary" phenomena.
Comparison with Baseline Methods
Instantiation-Net is directly compared with previous KPLSR and PLSR frameworks. On the full patient cohort, mean 3D error for Instantiation-Net is 2.21 mm, outperforming PLSR (2.38 mm) and closely approaching the performance of KPLSR (2.01 mm). The model yields consistent results across all subjects—without need for hand-crafted features or manual segmentation at inference, unlike the baselines.
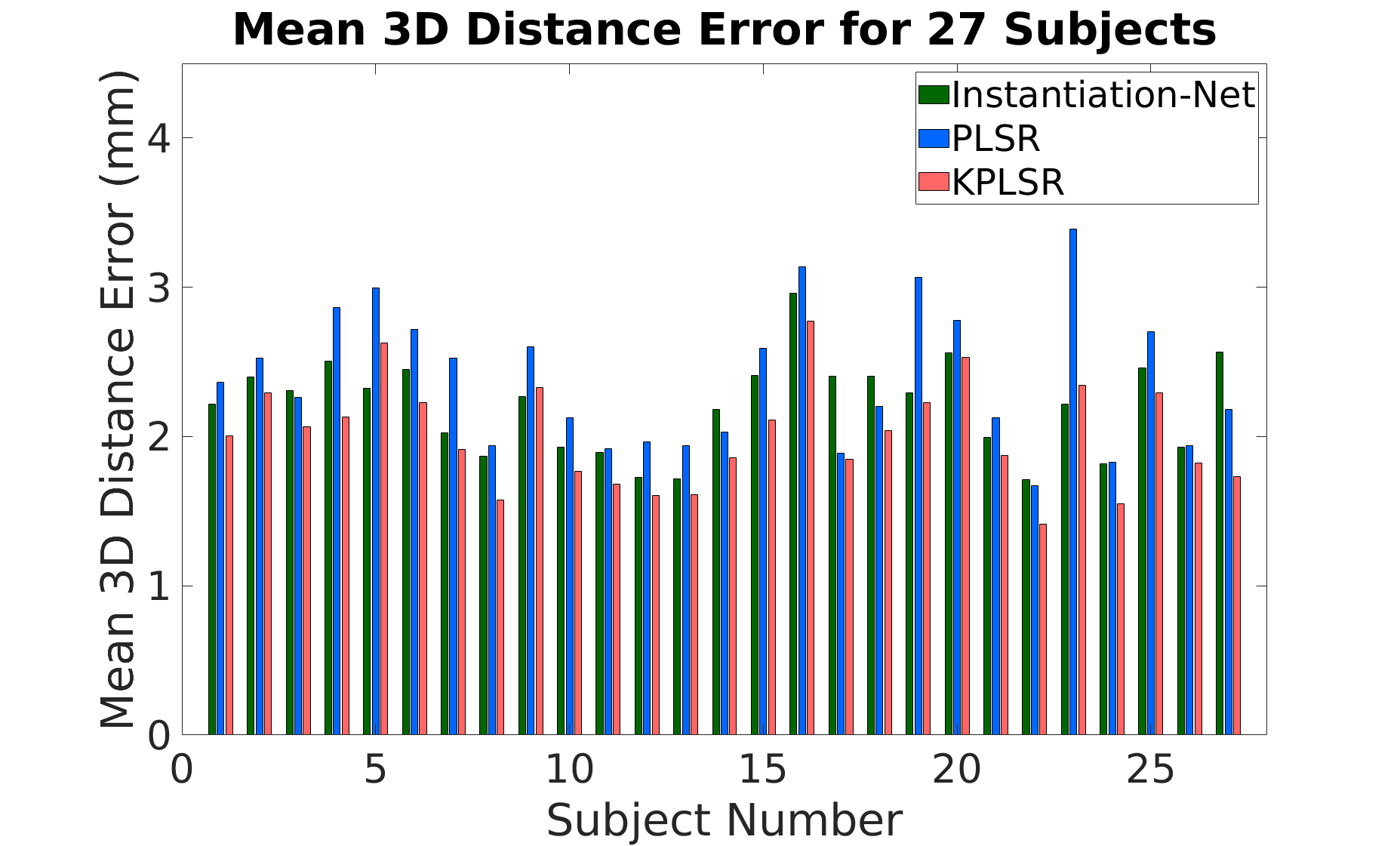
Figure 4: Subject-wise mean 3D mesh reconstruction error for Instantiation-Net, PLSR, and KPLSR approaches; Instantiation-Net exhibits competitive accuracy and lower variance.
Discussion
The findings underscore several practical and theoretical implications:
- End-to-end trainability and automation: Instantiation-Net eliminates dependency on manual segmentation and parameter tuning, streamlining deployment and reducing operator bias.
- High-resolution mesh output: In contrast to models limited to unstructured point clouds or low-res volumes, the output is a topologically consistent mesh with vertex correspondence—vital for intra-operative navigation and simulation.
- Computational/Resource Trade-offs: The main limitation lies in training resource demands (GPU memory and runtime), a trade-off for higher model expressivity and automation. However, inference speed is compatible with intra-operative requirements.
- Boundary and sparsity effects: Performance is attenuated in regions with limited imaging support (apical/basal mesh), necessitating either more comprehensive imaging or auxiliary regularization in these regimes.
- Generalizability: While validated on right ventricle MRI, the DCNN+GCN architecture is potentially extensible to 3D reconstruction tasks from limited 2D data in other anatomical or even non-medical settings—where mesh correspondence is crucial.
Future Directions
Key extensions include multi-view/multi-modal integration, domain adaptation for robustness across scanner protocols, and scaling to whole-organ reconstructions in complex anatomical regions. Incorporation of temporal priors or shape constraints may further ameliorate boundary effects and enable temporally consistent mesh sequences.
Conclusion
Instantiation-Net advances the state of the art for 3D mesh instantiation from minimal 2D imaging in cardiac applications, merging DCNN-based image feature extraction with GCN-based mesh decoding. Yielding mean errors on par with prior semi-automatic methods—and achieving full end-to-end differentiability and automation—this architecture facilitates accurate, efficient, and robust 3D reconstruction for clinical navigation and surgical planning. Its design paradigm is likely generalizable to broader 2D-to-3D instantiation challenges within medical AI and beyond.





