- The paper presents Neural Latent Aligner (NLA), a framework that aligns neural representations using a sequential autoencoder with contrastive loss.
- It introduces a differentiable time warping model (TWM) that effectively resolves temporal misalignments in intracranial ECoG data for improved behavioral decoding.
- Empirical results demonstrate that NLA achieves higher cross-trial consistency and decoding performance compared to baseline methods, highlighting its potential for robust brain-computer interfaces.
Neural Latent Aligner: Cross-trial Alignment for Learning Representations of Complex, Naturalistic Neural Data
Introduction
The paper introduces a framework called Neural Latent Aligner (NLA) for learning representations from complex, naturalistic neural data. Addressing the challenges of high-dimensional behaviors and low signal-to-noise ratios, the NLA is designed to align neural data representation across trials using unsupervised methods. The framework introduces a differentiable time warping model (TWM) to resolve temporal misalignment between trials. This method is applied to intracranial electrocorticography (ECoG) data of natural speaking, proving superior to baseline models in decoding behaviors, particularly in lower dimensional settings.
Proposed Framework
Neural Latent Aligner (NLA)
The NLA framework aims to find consistent neural data representations across repeated trials by learning cross-trial alignment. This is achieved through a sequential autoencoder with 1D CNNs to encode temporal factors (Figure 1). The architecture leverages InfoNCE-based contrastive loss to maximize mutual information between signals, specifically focusing on behaviorally relevant features.
Time Warping Model (TWM)
To address trial misalignments, the framework proposes a novel neural time warping model (TWM) that provides a differentiable solution for alignment (Figure 2). This model postulates unimodal and monotonic probabilistic distribution over time indexing, ensuring one-to-one mappings without inversions in alignment sequences.
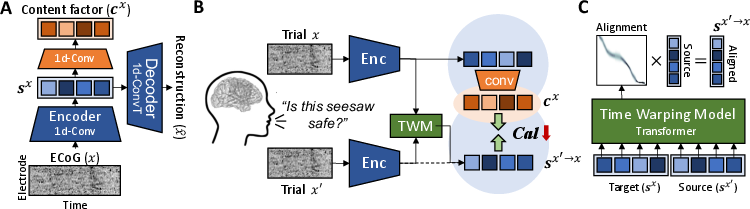
Figure 1: Overview of the proposed framework. A) Sequential autoencoder with 1D CNNs for encoding sx and cx. B) Diagram of the proposed cross-trial alignment minimizing the contrastive alignment loss. C) Time warping model for alignment distribution from source to target.
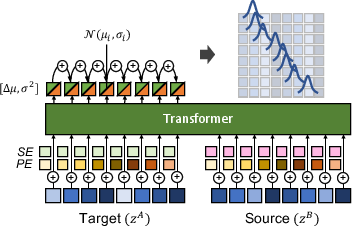
Figure 2: Architecture of the proposed neural time warping model.
Results
Behavioral Relevance
The paper evaluates the NLA by decoding articulatory kinematic trajectories (AKTs) using linear models. The framework outperforms baseline methods in both full and reduced dimensional settings, showing greater behavioral relevance and efficiency with lower dimensional representations (Figure 3).
Alignment Accuracy
The TWM is validated empirically by assessing the phonetic coherence between warped and target trials, showing a high true positive rate (TPR) for proper phoneme alignment (Figure 4). It outperforms traditional dynamic time warping methods applied post hoc on other representations.
Cross-trial Consistency
The content factor from NLA exhibits greater cross-trial consistency compared to baselines, with visualizations from t-SNE manifolds revealing coherent shared neural trajectories across trials (Figure 5).
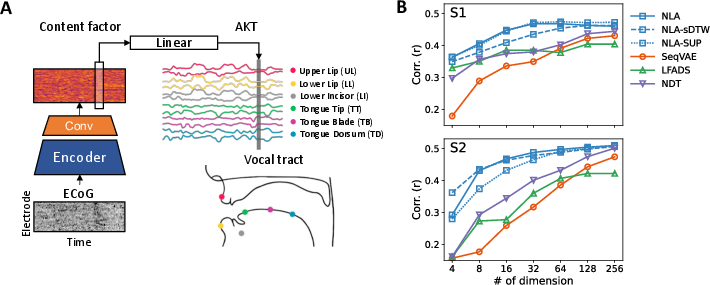
Figure 3: A) Overview of behavioral decoding analysis using a linear model. B) Behavioral decoding performance across different dimensionalities.
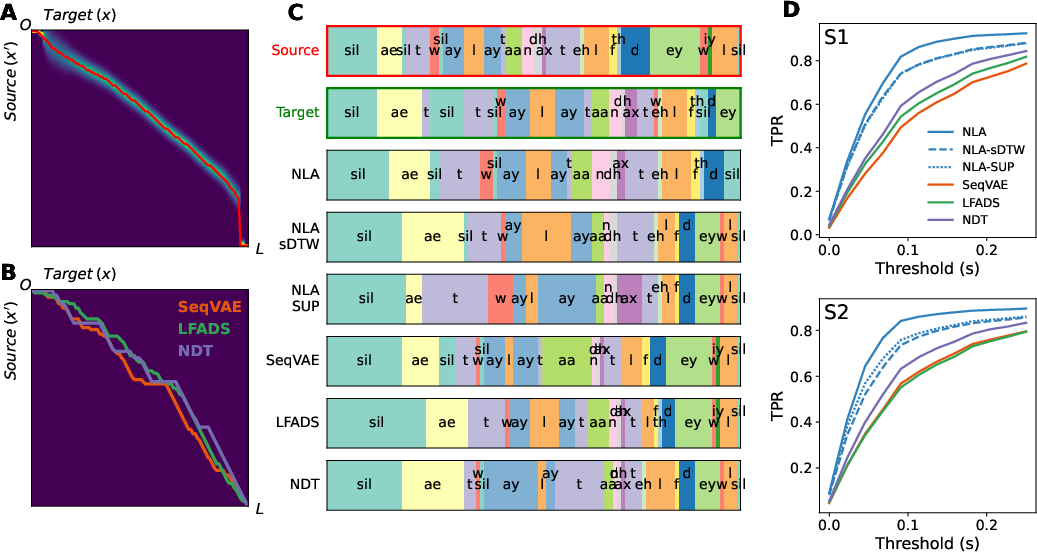
Figure 4: A) Alignment distribution inferred from TWM. Red line indicates warping path. B) Alignments from DTW applied to baseline models. C) Frame-wise phoneme labels across trials.

Figure 5: Manifolds visualized by t-SNE showing shared neural trajectories across different models.
Discussion and Conclusion
The NLA framework demonstrates its potential to uncover behaviorally relevant representations amid complex neural data without supervision. Particularly, the content factor's compactness and correlation with behaviors suggest potential applications in enhancing robustness and efficiency in brain-computer interfaces (BCIs). The framework's design is flexible enough to be applied to various domains, encouraging future exploration in handling diverse neural dynamics and behaviors across subjects or conditions.
In closing, the proposed NLA framework establishes a novel approach to neural data alignment, providing a foundation for further advancements in the field of neural representation learning. With its distinguished performance in handling naturalistic, high-dimensional neural datasets, NLA opens avenues to deeper insights into neural computation and the development of more reliable neurotechnologies.




