Disentangling Hippocampal Shape Variations: A Study of Neurological Disorders Using Mesh Variational Autoencoder with Contrastive Learning
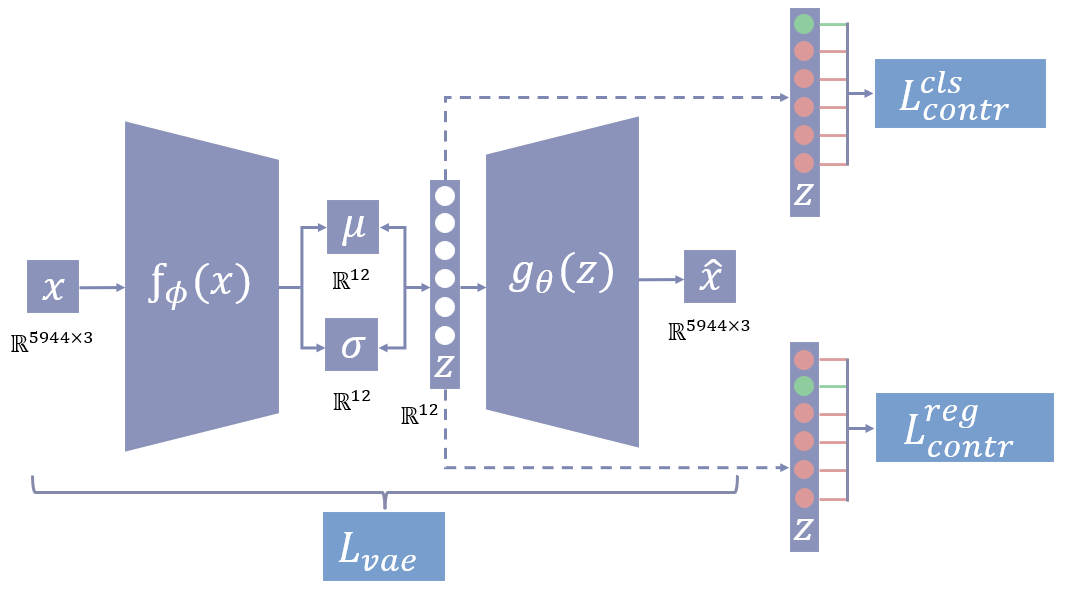
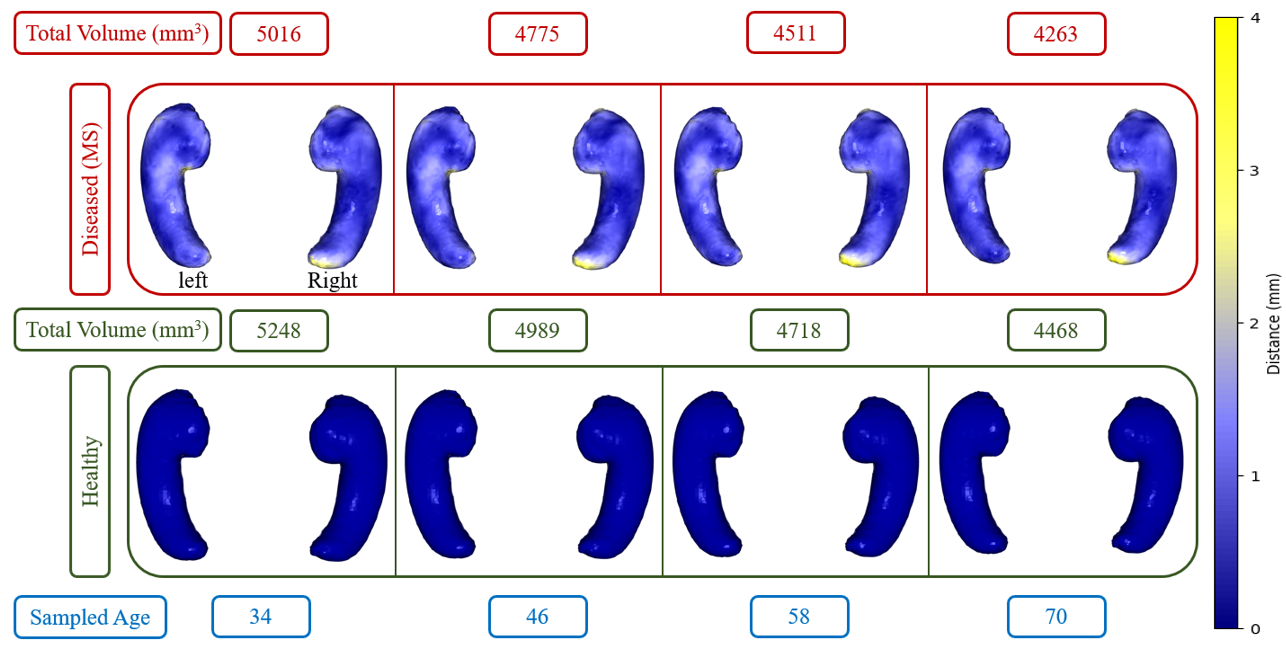
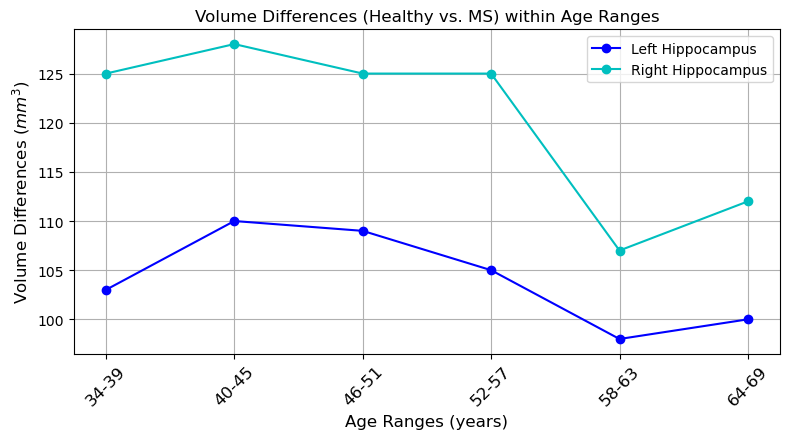
Abstract: This paper presents a comprehensive study focused on disentangling hippocampal shape variations from diffusion tensor imaging (DTI) datasets within the context of neurological disorders. Leveraging a Mesh Variational Autoencoder (VAE) enhanced with Supervised Contrastive Learning, our approach aims to improve interpretability by disentangling two distinct latent variables corresponding to age and the presence of diseases. In our ablation study, we investigate a range of VAE architectures and contrastive loss functions, showcasing the enhanced disentanglement capabilities of our approach. This evaluation uses synthetic 3D torus mesh data and real 3D hippocampal mesh datasets derived from the DTI hippocampal dataset. Our supervised disentanglement model outperforms several state-of-the-art (SOTA) methods like attribute and guided VAEs in terms of disentanglement scores. Our model distinguishes between age groups and disease status in patients with Multiple Sclerosis (MS) using the hippocampus data. Our Mesh VAE with Supervised Contrastive Learning shows the volume changes of the hippocampus of MS populations at different ages, and the result is consistent with the current neuroimaging literature. This research provides valuable insights into the relationship between neurological disorder and hippocampal shape changes in different age groups of MS populations using a Mesh VAE with Supervised Contrastive loss. Our code is available at https://github.com/Jakaria08/Explaining_Shape_Variability
- Optuna: A next-generation hyperparameter optimization framework. In Proceedings of the 25th ACM SIGKDD international conference on knowledge discovery & data mining, pages 2623–2631, 2019.
- Going deep in medical image analysis: concepts, methods, challenges, and future directions. IEEE Access, 7:99540–99572, 2019.
- A contrastive learning approach for training variational autoencoder priors. Advances in neural information processing systems, 34:480–493, 2021.
- James Dean Brown. Point-biserial correlation coefficients. Statistics, 5(3):12–6, 2001.
- Understanding disentangling in beta𝑏𝑒𝑡𝑎betaitalic_b italic_e italic_t italic_a-vae. arXiv preprint arXiv:1804.03599, 2018.
- Attri-vae: Attribute-based interpretable representations of medical images with variational autoencoders. Computerized Medical Imaging and Graphics, 104:102158, 2023.
- Isolating sources of disentanglement in variational autoencoders. Advances in neural information processing systems, 31, 2018.
- The qt interval in patients with covid-19 treated with hydroxychloroquine and azithromycin. Nature medicine, 26(6):808–809, 2020.
- Pearson correlation coefficient. Noise reduction in speech processing, pages 1–4, 2009.
- Disentangled and controllable face image generation via 3d imitative-contrastive learning. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, pages 5154–5163, 2020.
- Guided variational autoencoder for disentanglement learning. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, pages 7920–7929, 2020.
- Emilien Dupont. Learning disentangled joint continuous and discrete representations. Advances in neural information processing systems, 31, 2018.
- Morphometry of anatomical shape complexes with dense deformations and sparse parameters. NeuroImage, 101:35–49, 2014.
- Hippocampus segmentation on high resolution diffusion mri. In 2021 IEEE 18th International Symposium on Biomedical Imaging (ISBI), pages 1369–1372. IEEE, 2021.
- Dava: Disentangling adversarial variational autoencoder. arXiv preprint arXiv:2303.01384, 2023.
- 3d shape variational autoencoder latent disentanglement via mini-batch feature swapping for bodies and faces. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, pages 18730–18739, 2022.
- Analyzing and improving representations with the soft nearest neighbor loss. In International conference on machine learning, pages 2012–2020. PMLR, 2019.
- Spiralnet++: A fast and highly efficient mesh convolution operator. In Proceedings of the IEEE/CVF international conference on computer vision workshops, pages 0–0, 2019.
- beta-vae: Learning basic visual concepts with a constrained variational framework. In International conference on learning representations, 2016.
- Contrastive masked autoencoders are stronger vision learners. IEEE Transactions on Pattern Analysis and Machine Intelligence, 2023.
- Memory impairment in multiple sclerosis: relevance of hippocampal activation and hippocampal connectivity. Multiple Sclerosis Journal, 21(13):1705–1712, 2015.
- Application of k-nearest neighbor (knn) approach for predicting economic events: Theoretical background. International journal of engineering research and applications, 3(5):605–610, 2013.
- Explaining anatomical shape variability: Supervised disentangling with a variational graph autoencoder. In 2023 IEEE 20th International Symposium on Biomedical Imaging (ISBI), pages 1–5. IEEE, 2023.
- Disentangling by factorising. In International Conference on Machine Learning, pages 2649–2658. PMLR, 2018.
- An introduction to variational autoencoders. Foundations and Trends® in Machine Learning, 12(4):307–392, 2019.
- Variational inference of disentangled latent concepts from unlabeled observations. arXiv preprint arXiv:1711.00848, 2017.
- Marching cubes: A high resolution 3d surface construction algorithm. In Seminal graphics: pioneering efforts that shaped the field, pages 347–353. 1998.
- Voxel structure-based mesh reconstruction from a 3d point cloud. IEEE Transactions on Multimedia, 24:1815–1829, 2021.
- Representation learning with contrastive predictive coding. arXiv preprint arXiv:1807.03748, 2018.
- Structural and functional hippocampal changes in multiple sclerosis patients with intact memory function. Radiology, 255(2):595–604, 2010.
- High resolution diffusion tensor imaging of the hippocampus across the healthy lifespan. Hippocampus, 31(12):1271–1284, 2021.
- Information bottlenecked variational autoencoder for disentangled 3d facial expression modelling. In Proceedings of the IEEE/CVF Winter Conference on Applications of Computer Vision, pages 157–166, 2022.
- High-resolution diffusion tensor imaging and t2 mapping detect regional changes within the hippocampus in multiple sclerosis. NMR in Biomedicine, 36(9):e4952, 2023.
- Explainable artificial intelligence (xai) in deep learning-based medical image analysis. Medical Image Analysis, 79:102470, 2022.
- Pointflow: 3d point cloud generation with continuous normalizing flows. In Proceedings of the IEEE/CVF international conference on computer vision, pages 4541–4550, 2019.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Collections
Sign up for free to add this paper to one or more collections.