- The paper introduces a multi-agent system that uses GPT-4 to decompose clinical trial problems into specialized tasks for accurate outcome predictions.
- The paper demonstrates significant improvements in ROC-AUC and PR-AUC metrics over baseline methods, validating its superior predictive performance.
- The paper integrates external knowledge bases and few-shot learning to overcome traditional LLM limitations in clinical trial analytics.
ClinicalAgent: A Multi-Agent System for Clinical Trial Reasoning via LLMs
Introduction
The ClinicalAgent system proposes a LLM-oriented multi-agent framework designed for complex clinical trial tasks, particularly clinical trial outcome prediction and process decomposition. Leveraging GPT-4 as the underlying reasoning engine and augmenting its capabilities via multi-agent organization, external tool integration, and explicit advanced reasoning paradigms, the system addresses limitations of previous LLM-only solutions in medical informatics—namely, lack of access to up-to-date domain knowledge, insufficient actionable prediction capacity, and inability to systematically decompose and solve clinical tasks that require multi-perspective analysis.
The framework is motivated by the necessity to fully operationalize LLMs for real-world clinical trial analytics, extending their role from conversational support to genuine reasoning, retrieval, and synthesis in multi-factorial tasks. Central technical contributions include agent specialization, function calling to medical databases (e.g., DrugBank, Hetionet), integration of few-shot learning with specialized system prompts, and amalgamation of ReAct and Least-to-Most reasoning patterns.
Architecture and Agent System
ClinicalAgent (CT-Agent) is architected as a tightly orchestrated, expert-inspired multi-agent system. The core methodology is depicted below.
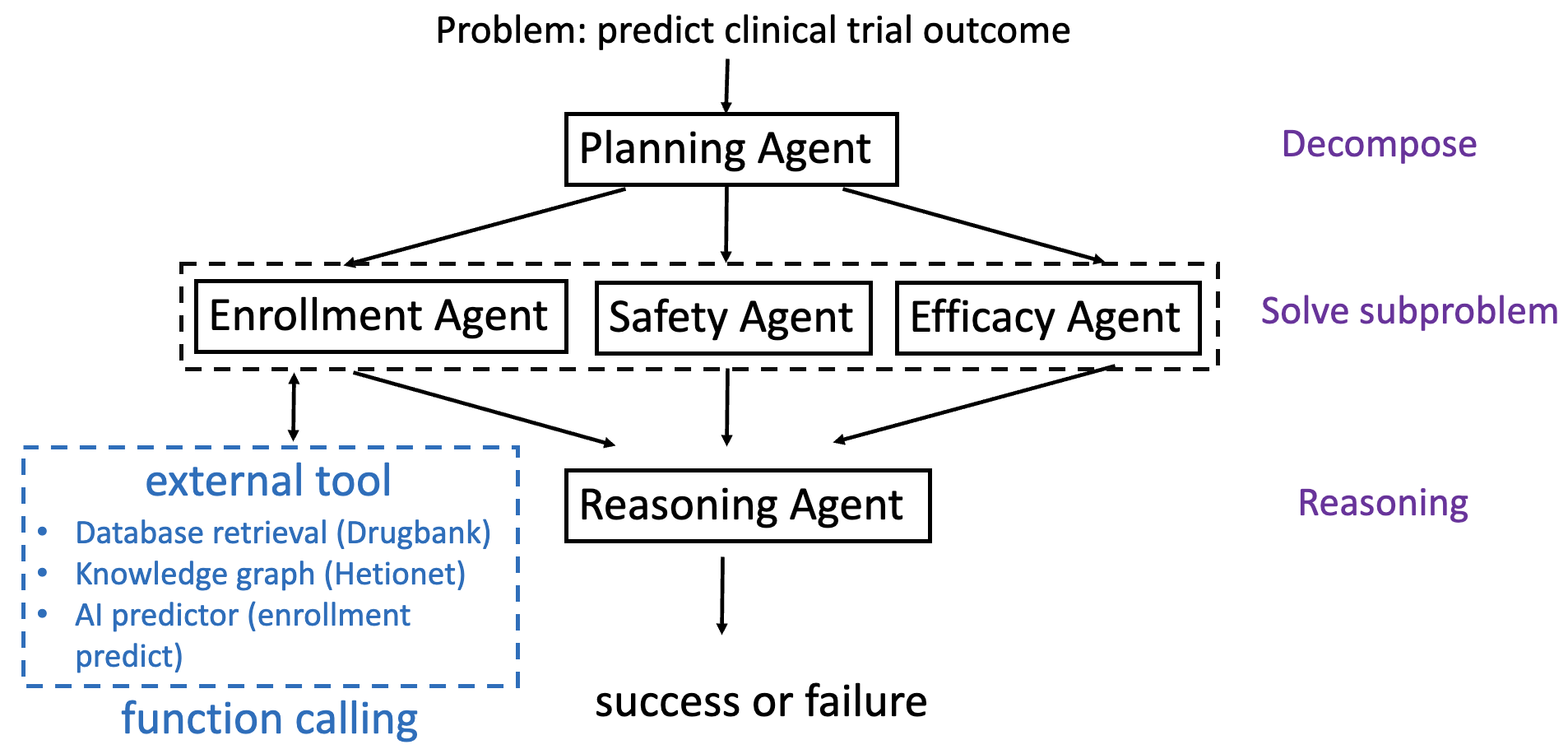
Figure 1: The CT-Agent framework highlights a Planning Agent that decomposes a clinical problem into subproblems (enrollment, safety, and efficacy) and routes these to respective specialist agents, each of which may invoke external tools and databases for task completion.
The agent roles are:
- Planning Agent: Receives the initial query (e.g., trial design, target drugs/diseases, eligibility criteria) and decomposes the high-level clinical problem using the Least-to-Most (LtM) strategy. This agent delegates subproblems, maintains workflow state, and aggregates sub-agent outputs.
- Enrollment Agent: Predicts enrollment feasibility based on trial eligibility criteria, applying a hierarchical transformer model (TrialDura) with contextual embeddings. Integrates external tools and outputs a binary or probabilistic enrollment success assessment.
- Safety Agent: Estimates risk (primarily drug toxicity and ADRs) based on historical DrugBank and trial registry evidence. Performs direct database lookups and applies statistical models to quantify safety failure probabilities.
- Efficacy Agent: Evaluates the potential therapeutic efficacy of the intervention, utilizing DrugBank, Hetionet, and pathway analysis. It leverages structured retrieval and knowledge graph reasoning on drug/disease associations, mechanisms, and prior outcomes.
Each agent operates as a GPT-4-driven instance receiving a role-specific system prompt and function definition suite. The Reasoning Agent then aggregates and synthesizes outputs, enabling explainable final predictions.
ClinicalAgent systematically overcomes knowledge limitations inherent in closed LLMs by incorporating tool use via function-calling infrastructure. These function definitions (in a strict JSON schema) allow agents to execute:
- Fast database retrieval (e.g., drug properties, historical outcomes, contraindications)
- Knowledge graph path extraction (e.g., Hetionet compound–gene–disease relationships)
- Predictive model inference (enrollment models, risk calculators)
The Reasoning Agent synthesizes subproblem responses, performing ReAct-style context-aware reasoning and, where appropriate, few-shot prompted deduction, to arrive at a comprehensive recommendation or prediction.
Workflow and Execution Details
The agent workflow is:
- Planning Agent ingests the user/submitted clinical trial description and decomposes it via LtM reasoning into atomic subproblems along enrollment, safety, and efficacy axes.
- Delegation to expert agents, which independently invoke external tools, retrieve current evidence, and produce specialized sub-responses.
- Aggregation of subproblem outputs. The system dynamically determines which function/tool to call using function selection in context.
- Contextual reasoning by the Reasoning Agent, often employing few-shot paradigms (by embedding prior cases or response templates), integrating the three axes into a structured, explainable prediction.
- Explicit, interpretable output to the user, including calculated metrics (e.g., predicted trial success/failure, supporting factors, risk rationales).
The micro-architecture ensures both transparency (exposing reasoning chains) and extensibility (support of new tools or agents).
Experimental Results
CT-Agent is evaluated on clinical trial outcome prediction benchmarks with rigorous, reproducible methodology. Major points:
- Data: Clinical trials benchmark dataset (including historical, synthetic, and generated data) is utilized for model calibration and test evaluation; 40 samples each for train/test are employed, ensuring controlled balance and reproducibility.
- Comparisons: Baseline methods include Gradient Boosted Decision Trees (with BioBERT feature encoding), Hierarchical Attention Transformer (deep feature attention), and LLM baselines (standard GPT-4/3.5 prompts).
- Metrics: Outcome prediction assessed via ROC-AUC, PR-AUC, accuracy, precision, recall, and F1. Notably:
- CT-Agent: ROC-AUC = 0.8347 (highest among all evaluated baselines), PR-AUC = 0.7908
- Improvement over standard GPT-4 prompting method by 0.3326 in PR-AUC
- Consistent superiority in multi-metric evaluation, particularly in recall and AUC, indicating improved sensitivity to positive outcome classification.
Ablation studies explicitly compare agent variants with/without few-shot learning and varying LLM versions, revealing:
- Few-shot adaptation provides robust gains in recall and ranking-based metrics (AUC), at the expense of minor reductions in precision.
- Upgrading from GPT-3.5 to GPT-4 as the agent LLM backbone yields nontrivial performance lifts across all metrics.
Implementation Considerations
- System Prompts and Role Conditioning: Each agent is instantiated with a bespoke prompt, encoding its task (role-playing specialist paradigms), thereby sharpening LLM focus and reducing hallucination.
- Function Definition and Execution: JSON-based schemas define input arguments and outputs for all tools/databases, enabling tightly coupled, deterministic interaction between agents and external resources.
- Few-shot Prompting: For decomposition and reasoning agents, explicit inclusion of diverse solved example cases in the system prompt scaffolds robust context-sensitive inference.
- External Knowledge Integration: Direct lookup and semantic matching (e.g., with string normalization in drug/disease names) ensure retrieval is robust to naming variation and missingness in knowledge base coverage.
- Infrastructure: Implementation utilizes Python 3.8, PyTorch, and standard LLM APIs (e.g., OpenAI, Azure), executed on high-performance commodity GPU compute nodes with sufficient memory to support multi-agent concurrent tasks and large prompt objects.
Limitations and Future Outlook
While ClinicalAgent achieves significant improvement over conventional and baseline LLM-based methods, its scalability is bounded by the manual configuration of agent roles and toolsets. Full autonomy in agent specialization and adaptive tool integration will require meta-learning or modular automatic reasoning algorithm advances. Moreover, further expansion of external knowledge integration (including real-time EHR, registries, and multi-omics) is necessary for deployment in heterogeneous or dynamically evolving trial contexts. Additional work is needed to mitigate over-reliance on historical data bias, assure generalization across therapeutic areas or trial phases, and safeguard against adversarial/erroneous tool outputs.
Theoretically, this system foregrounds a productive pathway for LLM-driven symbolic and retrieval-augmented reasoning in structured medical decision support, opening the door for task generalization beyond clinical trials—e.g., multidisciplinary panel decision emulation, guideline synthesis, or regulatory impact assessment.
Conclusion
The ClinicalAgent (CT-Agent) system advances LLM-based clinical trial reasoning by embedding task decomposition, external knowledge/tool augmentation, and explicit, explainable multi-agent architecture. Empirical benchmarks confirm substantial gains over strong baselines in outcome prediction, especially in challenging multi-factorial contexts where single-agent LLMs stagnate. Limitations centering on manual agent/tool configuration highlight key opportunities for meta-agent learning and autonomous orchestration. The integration demonstrated herein exemplifies the next stage in moving from conversational to actionable, autonomous clinical informatics, and its modularity enables adaptation to a wide spectrum of biomedical reasoning domains.


