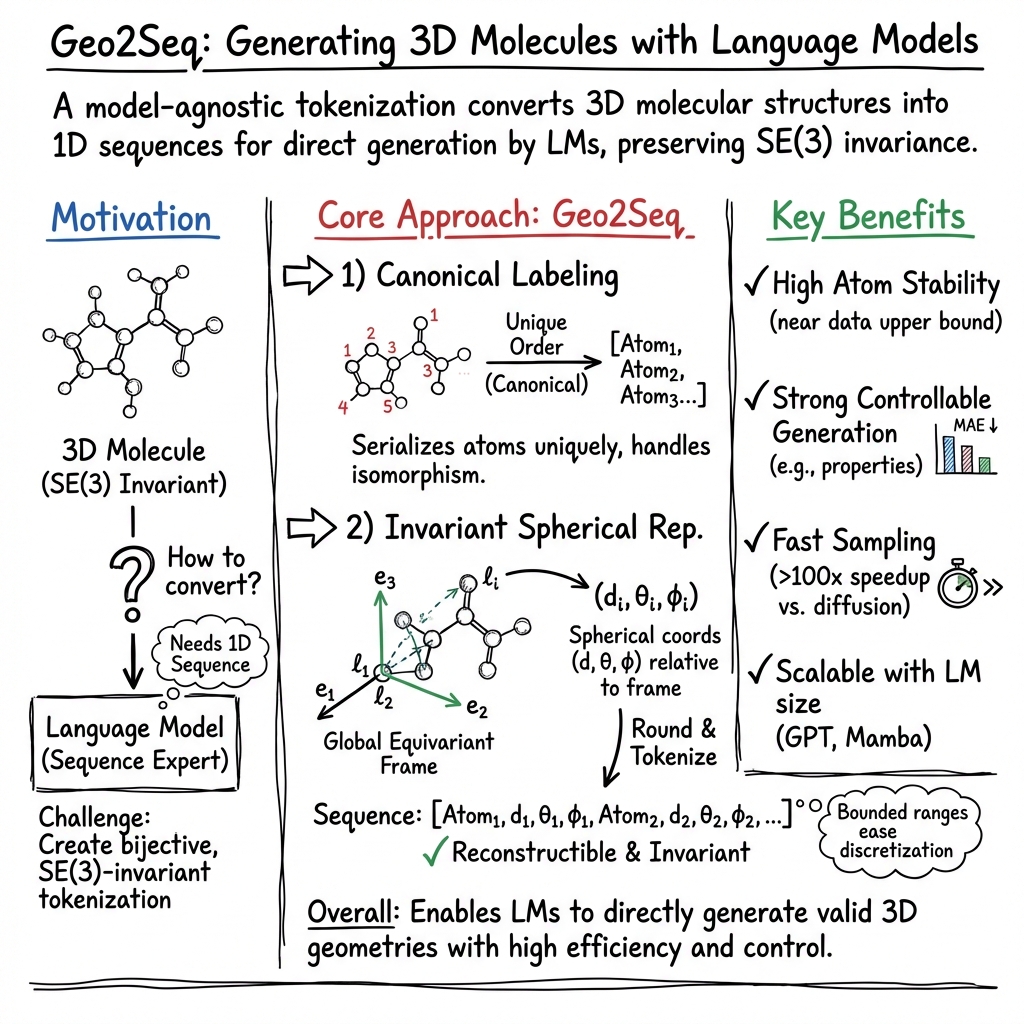
- The paper introduces Geo2Seq, a tokenization method that converts 3D molecular geometries into SE(3)-invariant 1D sequences for language model processing.
- It utilizes canonical labeling and invariant spherical representation to preserve atomic and structural fidelity, improving molecule stability.
- Geo2Seq outperforms diffusion-based methods on QM9 and GEOM-DRUGS datasets, enhancing controlled and valid molecular generation.
Introduction
The paper "Geometry Informed Tokenization of Molecules for LLM Generation" (2408.10120) introduces an innovative approach to molecular generation in three-dimensional (3D) space using LMs. It addresses the largely unexplored area of tokenizing 3D molecular geometries into discrete sequences, enabling the use of LMs for 3D molecule generation tasks. The authors propose Geo2Seq, a method that converts 3D molecular structures into SE(3)-invariant 1D discrete sequences, maintaining atomic and geometrical fidelity, which is conducive to processing by LMs. The study demonstrates the effectiveness of coupling various LMs with Geo2Seq, particularly in controlled generation tasks, yielding results with higher chemical validity compared to existing methods.
Geo2Seq Framework
Geo2Seq is comprised of two main components: canonical labeling and invariant spherical representation. The canonical labeling component ensures the uniqueness and distinction of the tokenized sequences by providing a standardized serialization of molecular graphs. This approach assigns a unique canonical form to each molecule, distinguishing between isomorphic and non-isomorphic structures effectively.
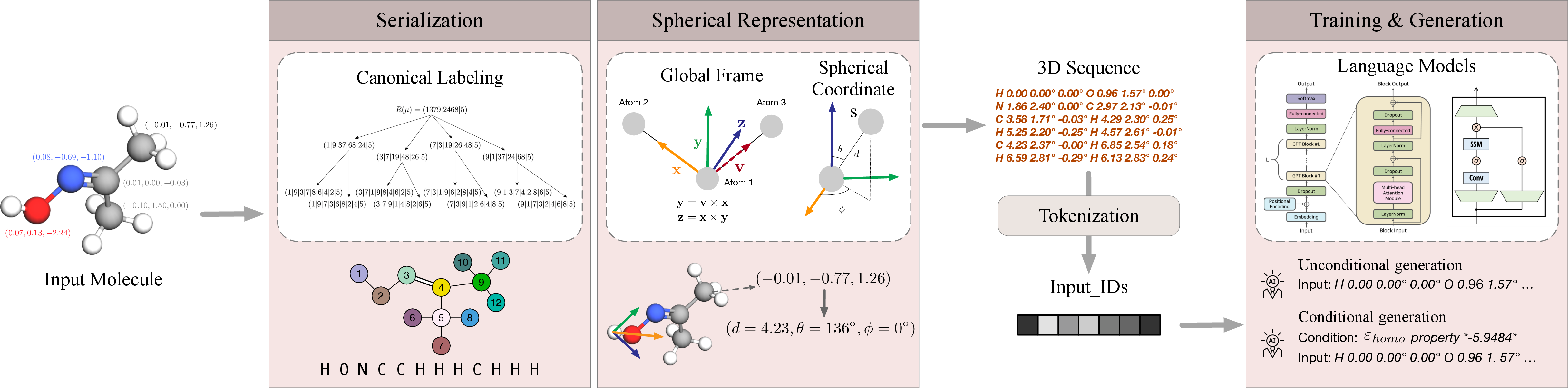
Figure 1: Overview of Geo2Seq. We use the canonical labeling order to arrange nodes in a row, fill each node's place with a vector [zi,di,θi,ϕi], and concatenate all elements into a sequence. Each node vector contains atom type and spherical coordinates, which are SE(3)-invariant.
In the invariant spherical representation component, Cartesian coordinates of atoms are transformed into spherical coordinates, providing a SE(3)-invariant representation that remains unchanged under rotation and translation transformations. This transformation minimizes information loss, ensuring that the sequences retain complete structural and geometrical information.
Experimental Insights
The experimental results highlight the advantages of using Geo2Seq with LMs for 3D molecule generation. On the QM9 dataset, Geo2Seq coupled with Mamba demonstrated marked improvements in atom and molecule stability, as well as in the generation of valid molecular structures. Additionally, when tested with the GEOM-DRUGS dataset, Geo2Seq with Mamba maintained high validity percentages and atom stability, outperforming several diffusion-based baselines.
For controllable generation tasks, Geo2Seq enabled significant advancements in generating molecules with specified quantum properties. The method exhibited superior performance across multiple property types compared to diffusion models, affirming the potential of LMs in goal-directed molecular design.
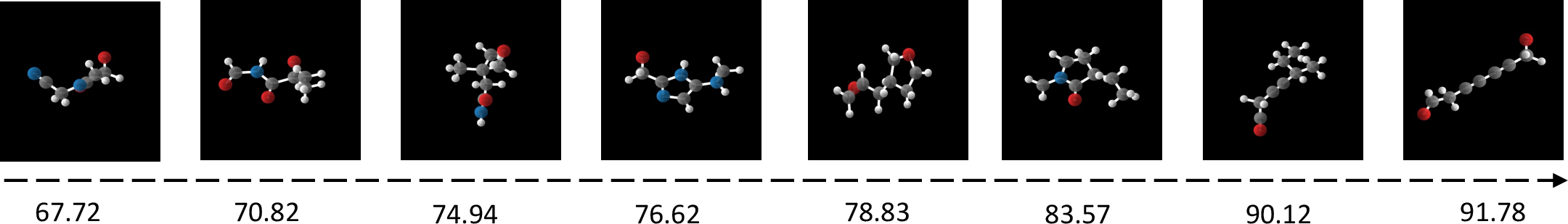
Figure 2: Visualization of generated molecules conditioned on the property of Polarizability α.
Token Embedding and Structure Learning
The paper also explores the learned token embeddings through UMAP visualizations, which reveal that Mamba models trained with Geo2Seq capture significant structural information. These insights suggest that LMs can effectively understand and generate 3D molecular structures by capitalizing on geometrical data transformed into sequence representations.
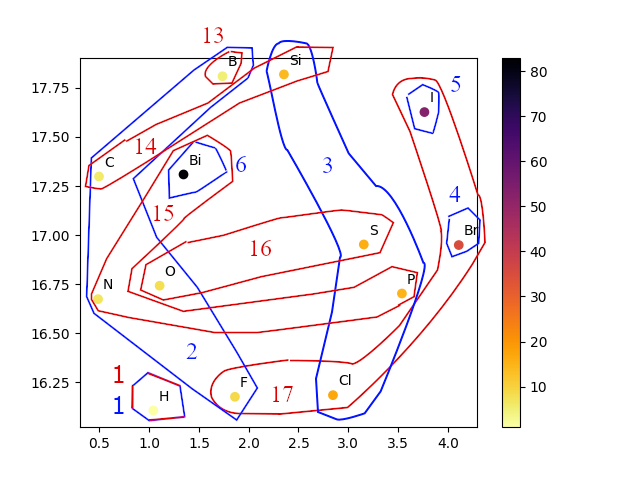
Figure 3: UMAP visualization of element token embeddings learned by a Mamba model trained on GEOM-DRUGS, illustrating the model's capacity to capture the periodic table's structure.
Conclusion
The research outlined in "Geometry Informed Tokenization of Molecules for LLM Generation" (2408.10120) underscores the transformative potential of Geo2Seq in adapting LLMs for 3D molecule generation tasks. By enabling the effective tokenization of 3D molecular geometries, Geo2Seq paves the way for using advanced sequence-processing capabilities of LMs in molecular design and drug discovery. The framework addresses existing limitations of diffusion models by providing efficient, scalable solutions with enhanced structural fidelity in generated molecules. Future work could further explore its application in broader chemical and pharmaceutical contexts, leveraging LMs' emergent abilities and adaptability to diverse molecular datasets and tasks.