- The paper introduces ChemTEB, a new benchmark to evaluate chemical text embeddings using domain-specific tasks.
- It assesses 34 models, including transformer-based and proprietary variants, highlighting trade-offs between performance and efficiency.
- Results underscore that domain adaptation and specialized datasets lead to improved embedding quality in chemical NLP applications.
ChemTEB: Evaluating Chemical Text Embeddings
This paper introduces the Chemical Text Embedding Benchmark (ChemTEB), a new framework for evaluating embedding models in the domain of chemical text data. It builds upon existing benchmarks by offering domain-specific tasks aimed at capturing the linguistic and semantic nuances inherent in chemical literature. This initiative underscores the increasing need for specialized approaches to cater to the unique challenges presented by chemical data.
Introduction to ChemTEB
Recent advances in NLP have spurred sophisticated techniques for text embedding, facilitating better information retrieval and understanding across domains. Embedding models have evolved from basic representations such as GloVe and Word2vec to context-aware transformer-based models like BERT. ChemTEB zeroes in on embedding models pertinent to chemical sciences, proposing tasks that evaluate model performance in this specialized domain.
ChemTEB's significance arises from the limitations of general frameworks like MTEB when applied to chemistry-specific texts, where context and precision are vital. The benchmark facilitates the development and evaluation of models tailored for chemistry applications, ensuring that both efficiency and performance are optimized for domain-specific tasks.
Dataset and Tasks
ChemTEB uses a collection of datasets derived from PubChem, Wikipedia, BeIR, CoconutDB, and Safety Data Sheets. It encompasses tasks such as Classification, Bitext Mining, Retrieval, Clustering, and Pair Classification, each capturing distinct aspects of chemical data processing. These diversified tasks allow models to be rigorously tested on text classification, semantic similarity, retrieval, and more complex tasks like SMILES representation matching.
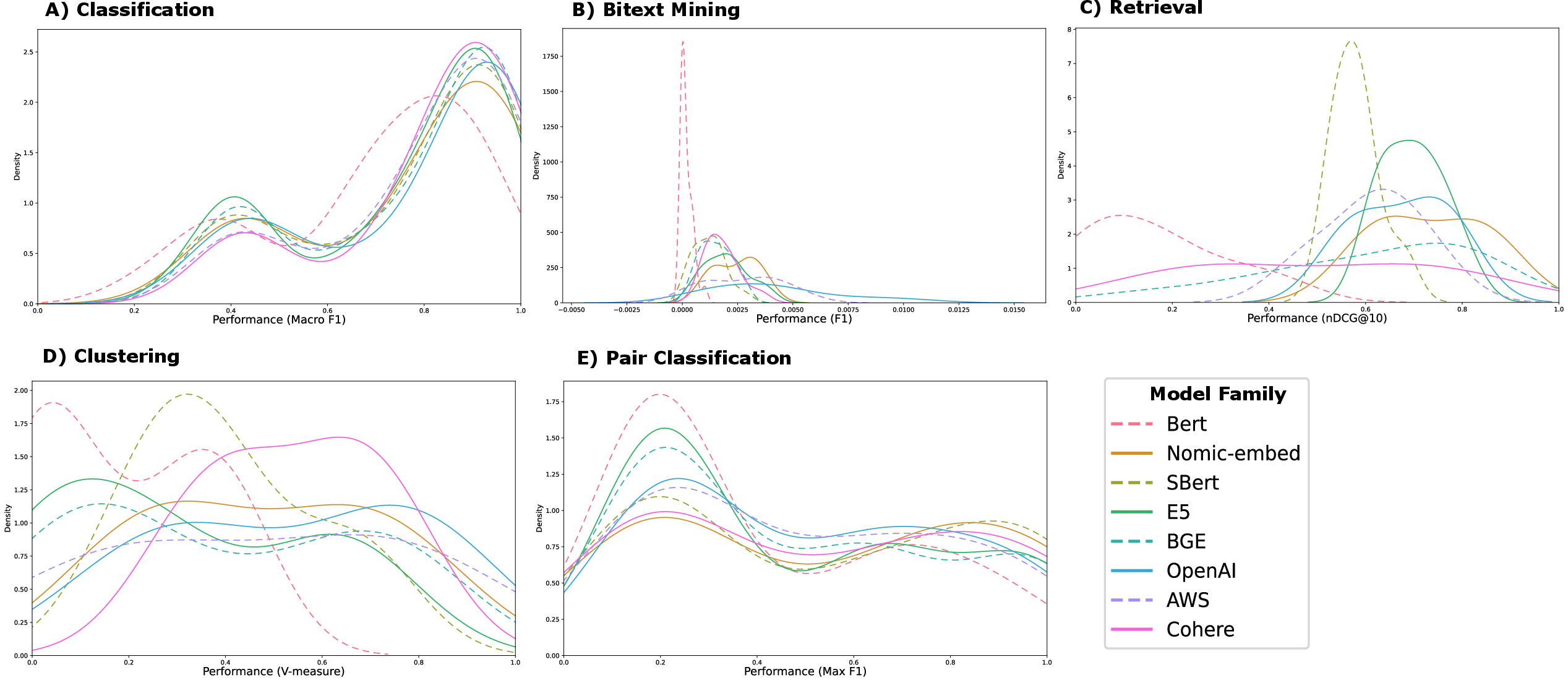
Figure 1: Distribution plots for five categories of tasks. The KDE plots show the probability density functions, where the x-axis represents the range of predicted values (performance distribution over tasks of each category and models of each family) and the y-axis represents the estimated density. Each colored line corresponds to a unique model family, enabling a clear visual comparison of their value distributions.
Models Evaluated
ChemTEB evaluates 34 models, including both open-source and proprietary models. Open-source variants encompass transformer models with self-supervised training methods such as MLM and contrastive learning, extending from generic models like BERT to domain-specific variants like MatSciBERT and ChemicalBERT. Proprietary models from OpenAI, Amazon, and Cohere provide insights into industry-oriented embedding solutions.
Several model families were created for comparative analysis: BERT Family, Nomic embedding family, SBERT family, E5 family, and others. Interestingly, domain adaptation of models like MatSciBERT proved advantageous in tasks requiring chemical domain expertise, suggesting that specialized corpora significantly impact performance.
Results and Analysis
Performance and efficiency metrics were key areas of investigation. Models generally performed best in classification tasks, while bitext mining proved challenging due to SMILES representation complexities. Proprietary models from OpenAI outperformed many open-source models, likely due to architectural optimizations and extensive training data.
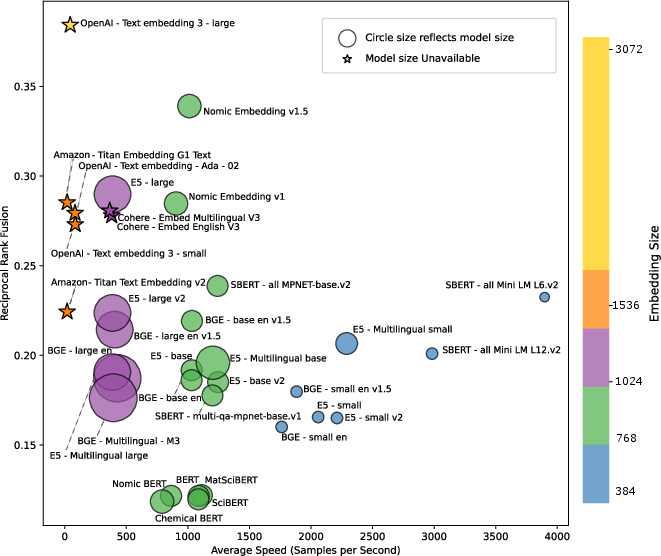
Figure 2: Summary of evaluated models in terms of efficiency. All evaluated models are depicted in the form of (i) circles (with circle size being proportional to the number of parameters) for open-source models, and (ii) stars for proprietary models. The color of the depicted models reflects their embedding dimension. The x-axis denotes the averaged inference speed (embedded samples/sec) calculated over seven pair classification tasks.
The efficiency analysis highlighted a trade-off between model size, embedding dimension, and inference speed, with larger models offering better performance at the cost of computational efficiency. Open-source models like Nomic Embedding v1.5 were notable for a balance of high speed and competitive performance.
Implications for Domain-Specific Models
ChemTEB emphasizes the need for advancing domain-adapted models, particularly at harnessing novel architectures that go beyond traditional methods. Performance metrics suggest that improvements in embedding quality could substantially benefit applications in chemical sciences, from literature analysis to molecular prediction tasks.
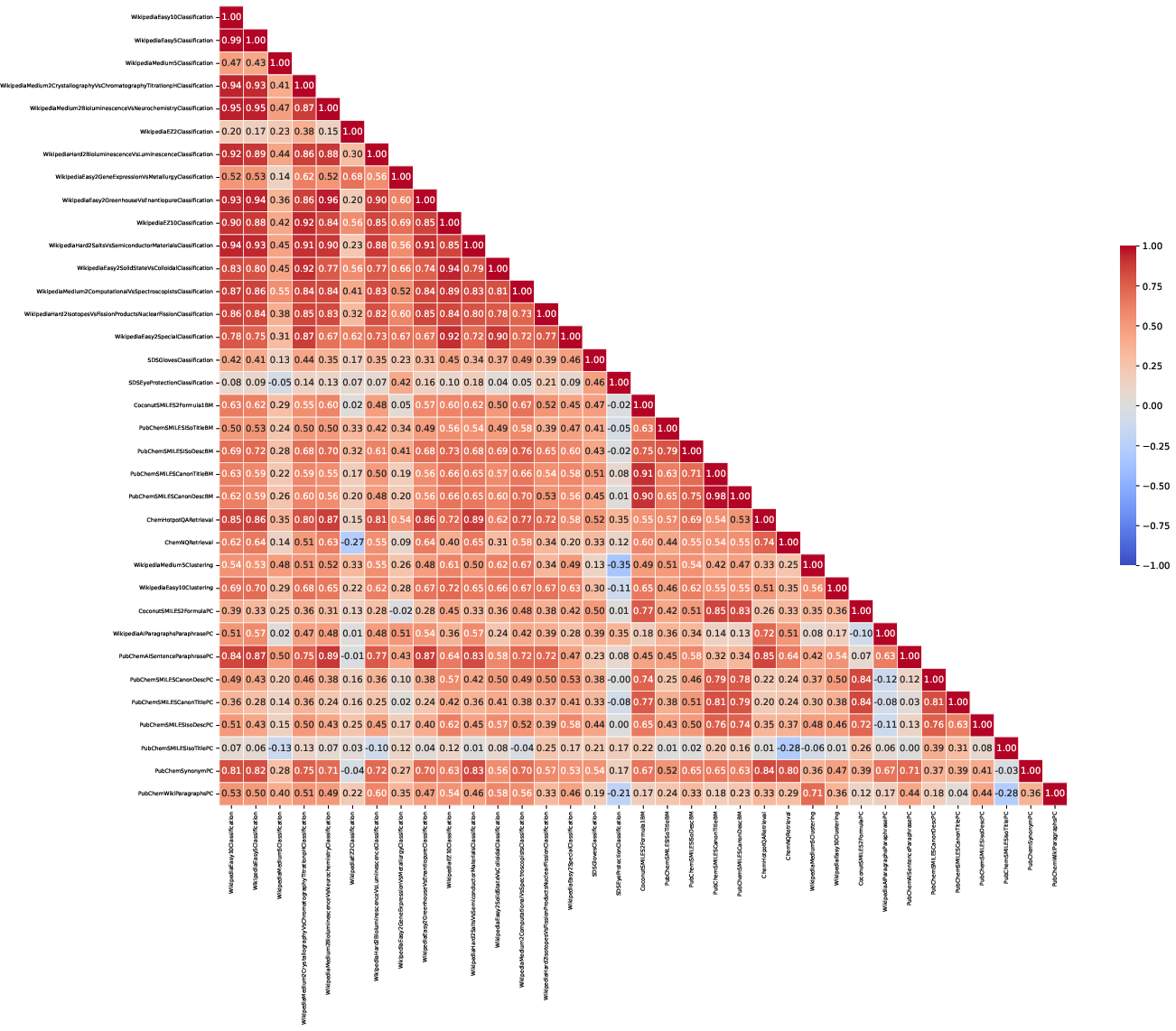
Figure 3: Correlation Matrix over datasets. Each row/column represents a separate dataset tested in the ChemTEB benchmark. The values and associated color reflect the correlation between the performance of different models on each pair of these datasets.
Conclusion
ChemTEB provides a structured, comprehensive framework to evaluate text embeddings in the chemical domain, facilitating the innovation in NLP models tailored to this field. It reveals the critical need for stronger domain-adapted models and underscores areas of potential development for embedding models in specialized scientific domains.
By focusing on domain-specific benchmarks, ChemTEB encourages the research community to refine and develop innovative solutions tailored to the demanding requirements of chemical data representation. Its contribution lies in driving the dialogue towards more precise and efficient models for chemical NLP applications, reinforcing the importance of specialized benchmarks in advancing scientific discovery.


