- The paper introduces GraphSHAP-IQ, an algorithm that exactly computes any-order Shapley interactions in GNNs by exploiting message-passing locality to drastically reduce computational calls.
- It leverages the concept of receptive fields to restrict non-trivial interactions within local neighborhoods, enabling both exact and approximate solutions for sparse and dense graphs.
- Experimental evaluations demonstrate that this approach scales linearly with graph size for fixed neighborhood bounds, offering actionable insights for interpretable model predictions in domains like chemistry and infrastructure.
Exact Computation of Any-Order Shapley Interactions for Graph Neural Networks
Introduction and Motivation
The paper addresses the challenge of interpreting predictions made by Graph Neural Networks (GNNs) on graph-structured data, focusing on the limitations of classical Shapley Value (SV) explanations. While SVs provide axiomatic feature attributions, they fail to capture higher-order interactions among features (nodes), which are critical for understanding complex model decisions. Shapley Interactions (SIs) generalize SVs to quantify joint contributions of feature subsets, but their computation is exponentially complex in the number of features. The authors introduce GraphSHAP-IQ, a model-specific algorithm for exact and efficient computation of any-order SIs in GNNs, leveraging the architectural properties of message-passing GNNs and the locality induced by receptive fields.
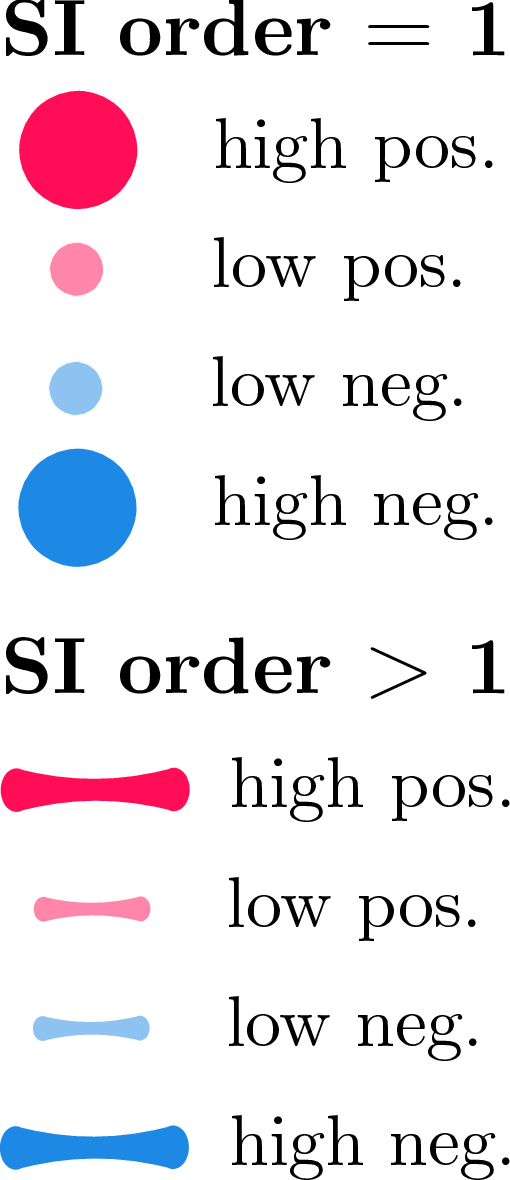
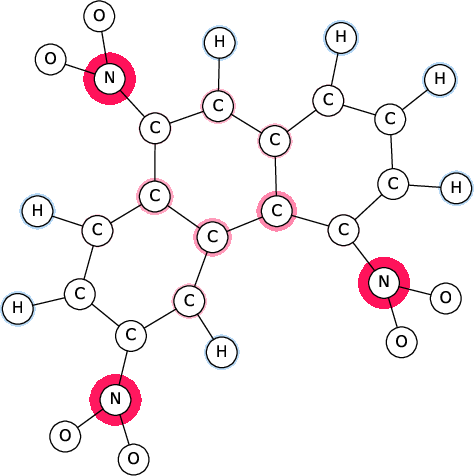
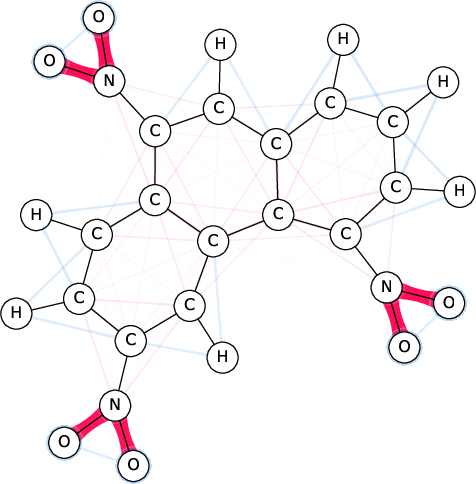
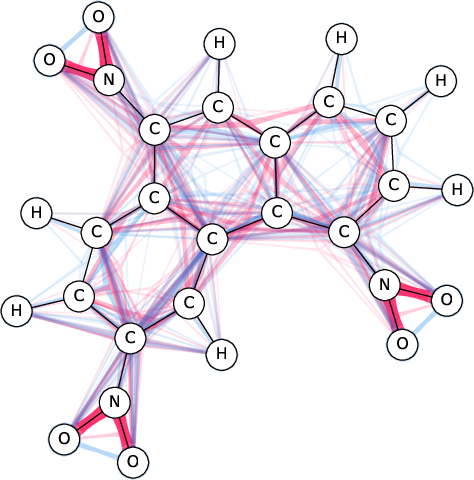
Figure 1: SI-Graphs overlayed on a molecule graph showing exact SIs for a mutagenicity prediction; GraphSHAP-IQ reduces the required model calls from 230 to $7,693$.
Theoretical Foundations
Shapley Interactions and Complexity
The paper formalizes the computation of SIs using cooperative game theory, where the model's output for a subset of features (nodes) is treated as the value of a coalition. The Möbius Interaction (MI) provides a unique additive decomposition of the model output, and SIs of order k (k-SII) interpolate between SVs (k=1) and MIs (k=n). The exponential complexity of SIs (2n model calls) is prohibitive for practical use, especially in graphs with tens or hundreds of nodes.
GNN-Induced Graph Game and Receptive Fields
The authors define a GNN-induced graph game, where the value function is the GNN's output on a graph with a subset of nodes masked (using a baseline feature vector). Crucially, for message-passing GNNs with ℓ layers, the embedding of a node depends only on its ℓ-hop neighborhood. This locality property is formalized in Theorem 3.3, which states that the node game is invariant to maskings outside the receptive field. As a result, the non-trivial MIs are restricted to subsets contained within ℓ-hop neighborhoods, dramatically reducing the number of required model calls.
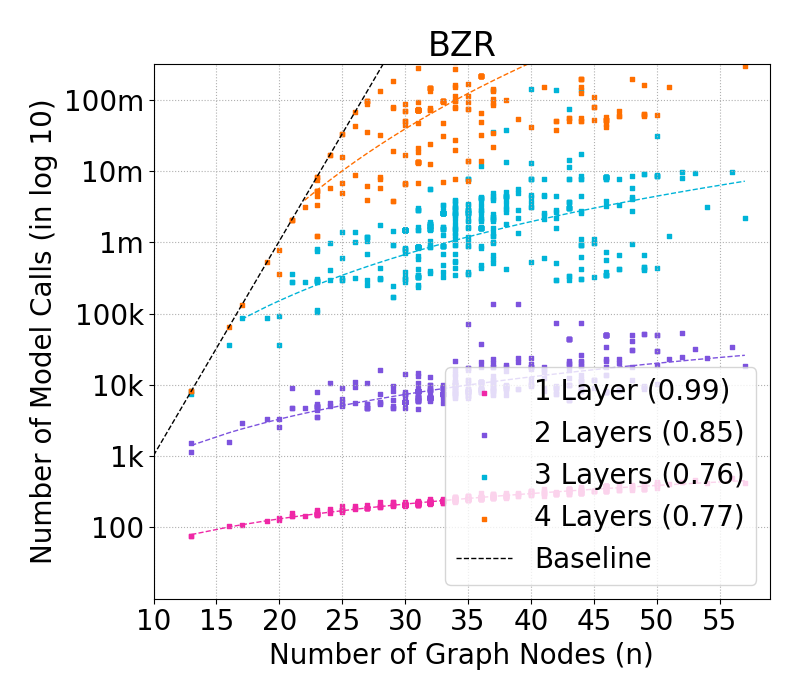
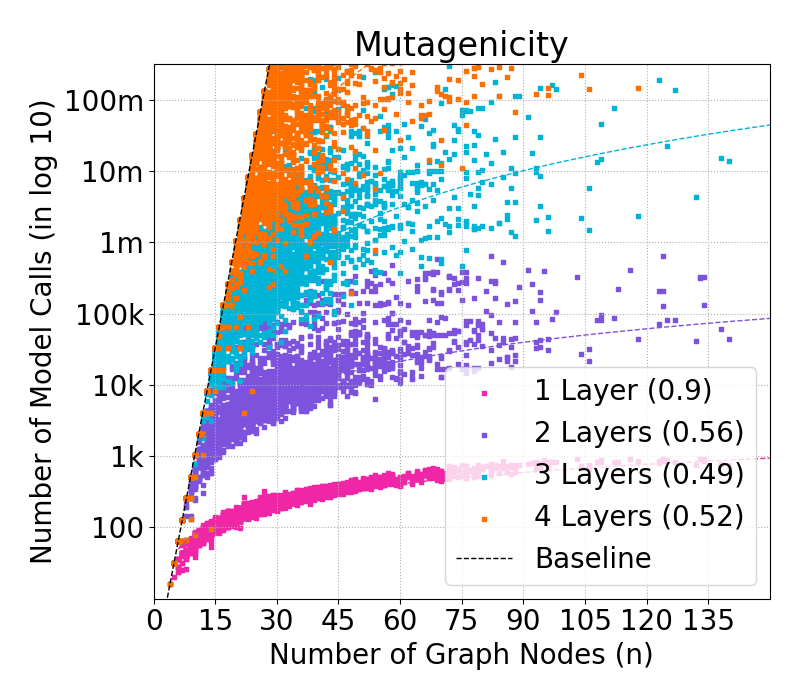
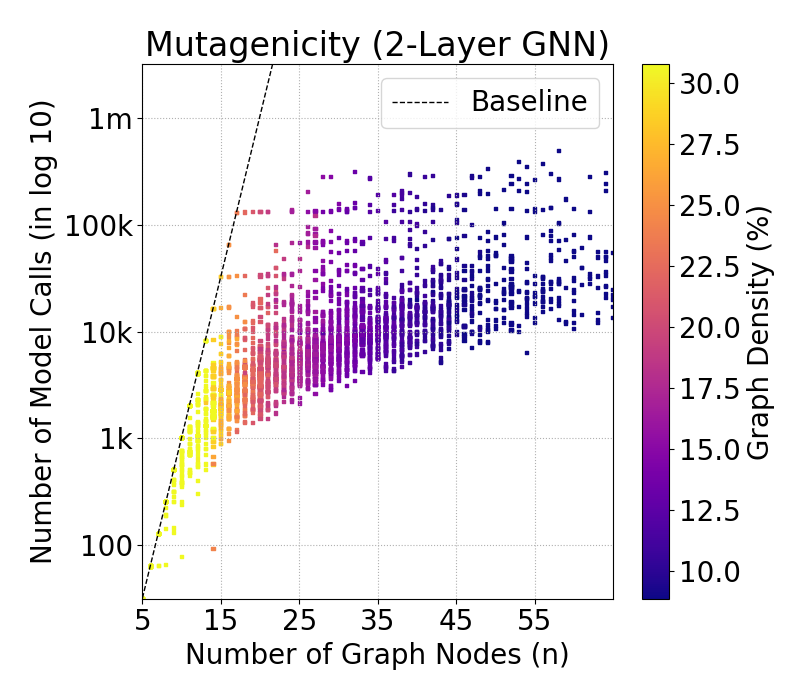
Figure 2: Complexity of GraphSHAP-IQ in model calls (log10) by number of nodes for BZR graphs; exponential baseline vs. linear scaling of GraphSHAP-IQ.
GraphSHAP-IQ: Algorithmic Implementation
GraphSHAP-IQ exploits the sparsity of non-trivial interactions induced by the GNN architecture. The algorithm proceeds as follows:
- Neighborhood Enumeration: For each node, enumerate all subsets of its ℓ-hop neighborhood.
- Model Evaluation: For each subset, mask the graph accordingly and evaluate the GNN.
- Möbius Interaction Computation: Compute MIs for all relevant subsets using inclusion-exclusion.
- SI Conversion: Convert MIs to SIs of desired order using established conversion formulas.
- Efficiency Guarantee: The sum of all SIs reconstructs the model's output due to the efficiency axiom.
The complexity is bounded by n⋅2nmax(ℓ), where nmax(ℓ) is the size of the largest ℓ-hop neighborhood. For sparse graphs and shallow GNNs, this is orders of magnitude smaller than 2n.

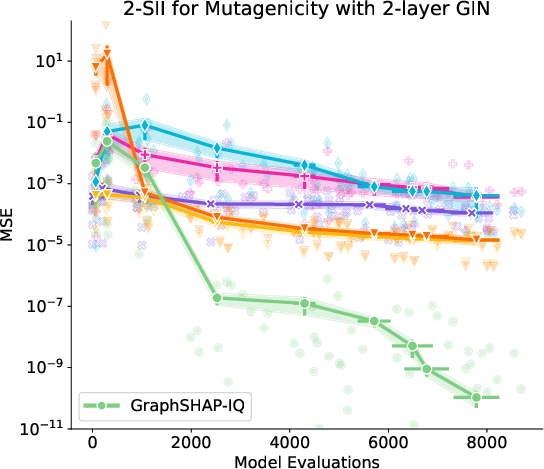
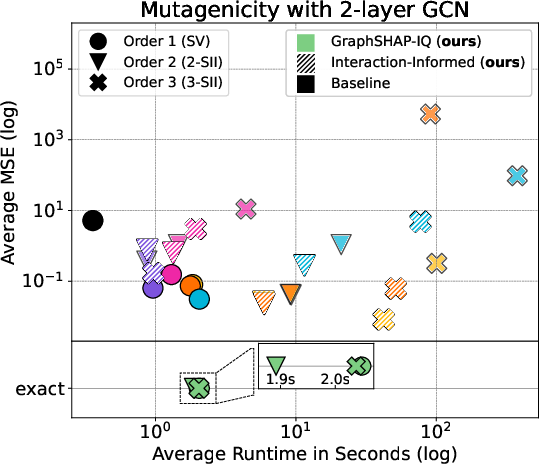
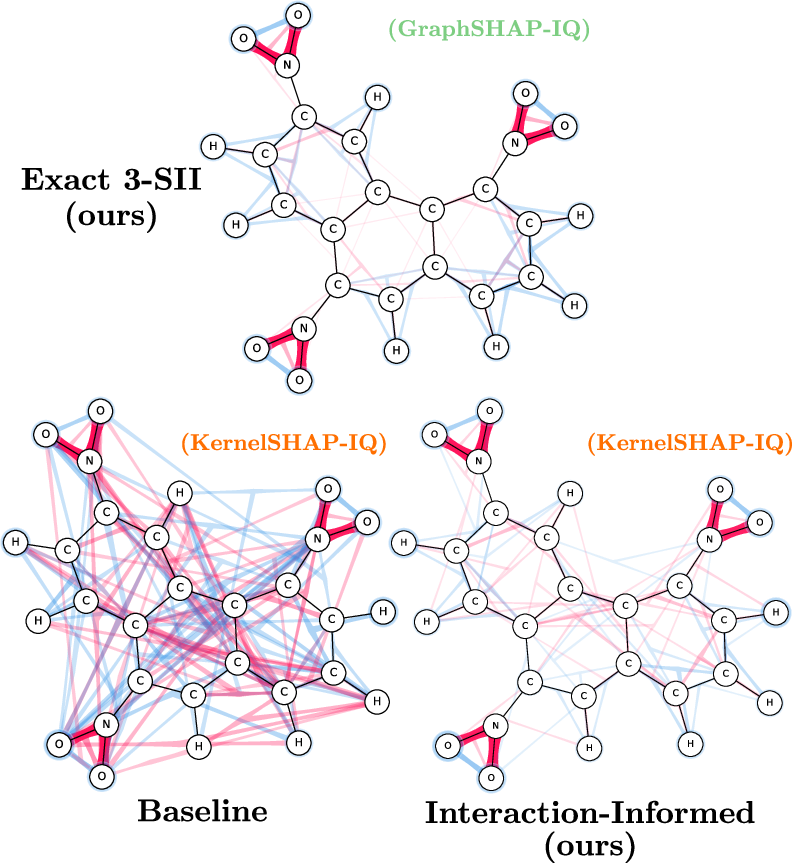
Figure 3: Approximation of SIs with GraphSHAP-IQ and model-agnostic baselines for Mutagenicity; GraphSHAP-IQ achieves exactness with far fewer model calls.
Empirical Evaluation
Complexity Reduction
Experiments on chemical and biological graph datasets (e.g., BZR, MTG) demonstrate that GraphSHAP-IQ enables exact computation of SIs for graphs with up to 100 nodes, where model-agnostic baselines are infeasible. The complexity scales linearly with graph size for fixed ℓ, and graph density is a strong proxy for computational cost.
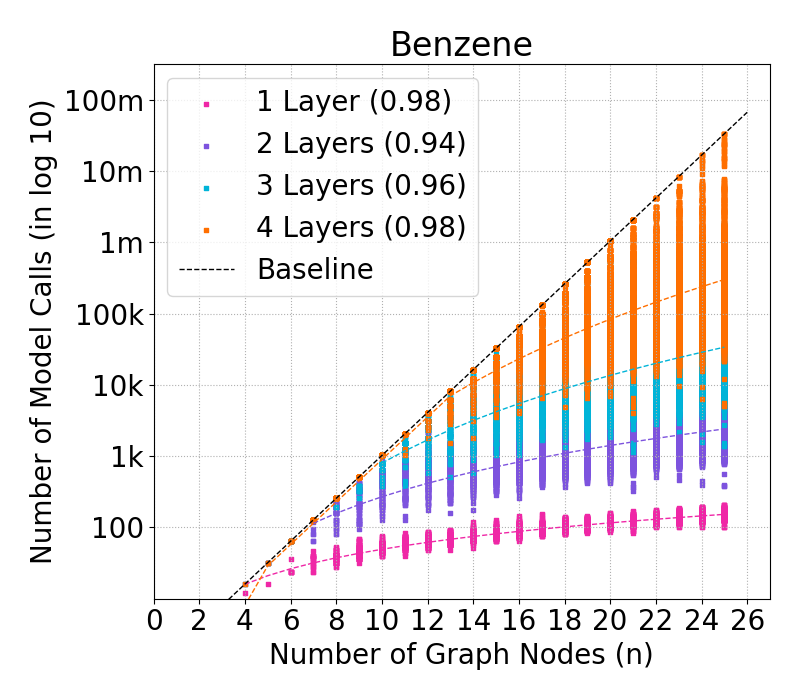
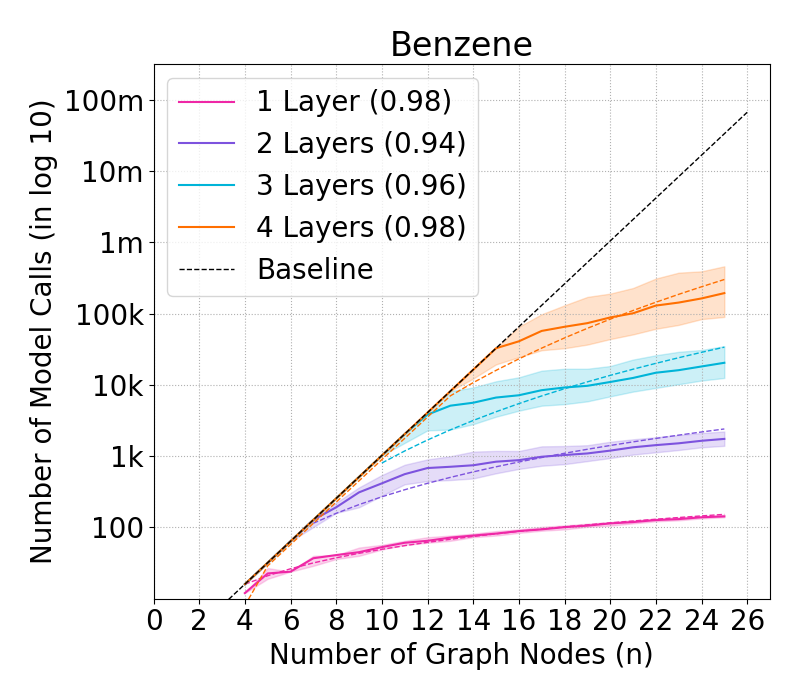
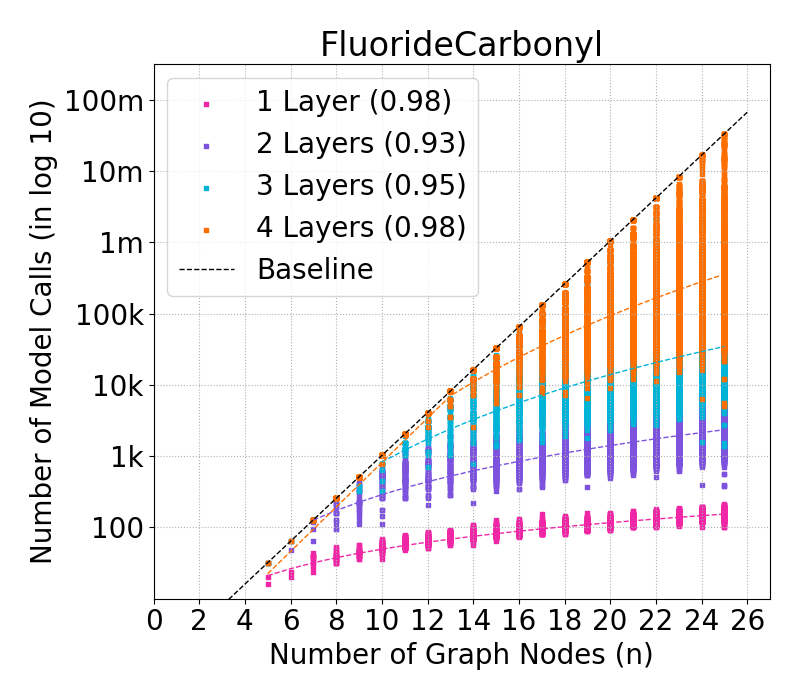
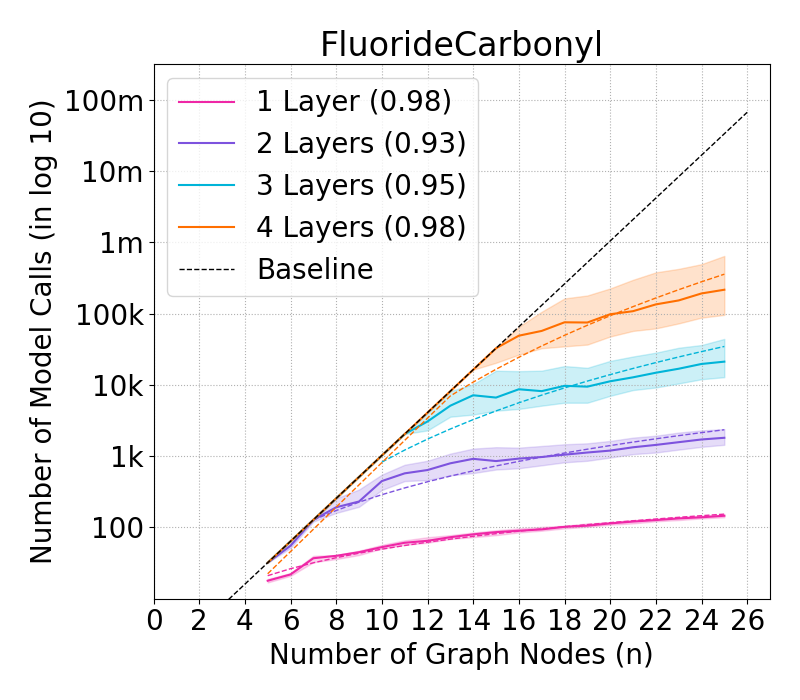


Figure 4: Complexity of GraphSHAP-IQ vs. baseline in model calls (log10) for BNZ; median, Q1, Q3 per graph size.
Approximation and Baseline Comparison
For dense graphs or deep GNNs, exact computation may still be infeasible. The authors propose an approximation variant of GraphSHAP-IQ, limiting the order of computed MIs. Interaction-informed baseline methods (e.g., KernelSHAP-IQ, SVARM-IQ) are adapted to exploit the sparsity structure, yielding substantial improvements in estimation quality and runtime.
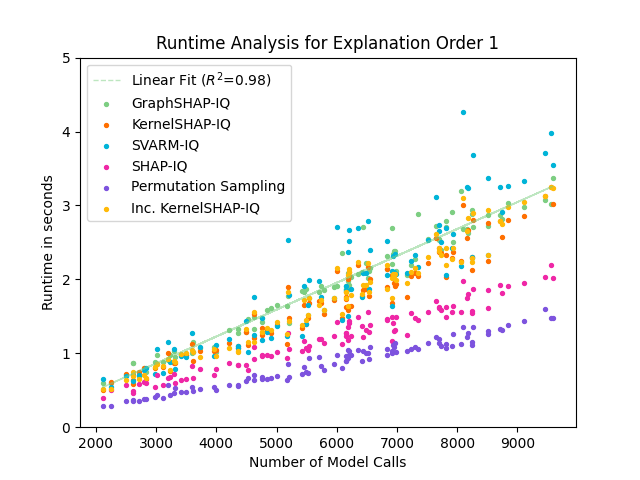

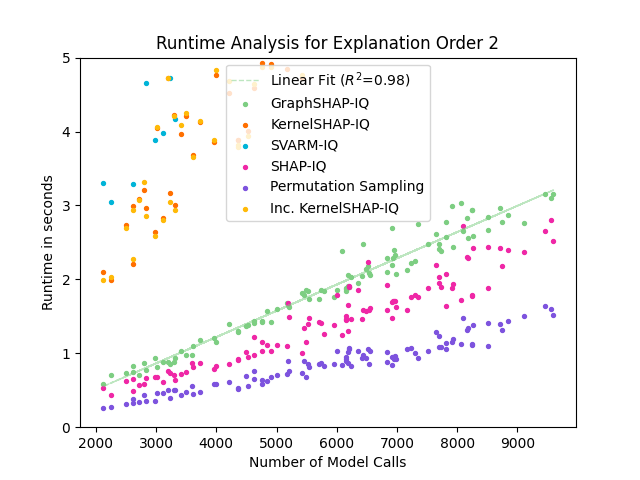

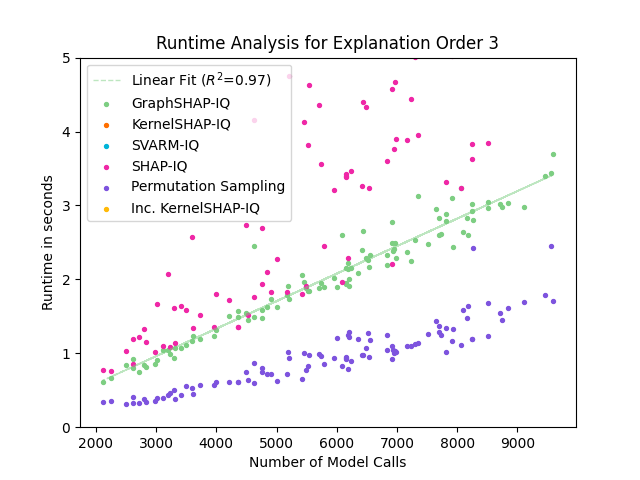

Figure 5: Runtime comparison of GraphSHAP-IQ and baseline methods for SVs; GraphSHAP-IQ runtime scales linearly with model calls and is independent of graph size.
Real-World Applications
GraphSHAP-IQ is applied to water distribution networks (WDNs) and molecular graphs. In WDNs, SIs reveal the trajectory of chlorination, aligning with physical flow patterns. In molecular graphs, SIs highlight functional groups (e.g., NO2 in mutagenicity, benzene rings), providing interpretable substructure attributions.
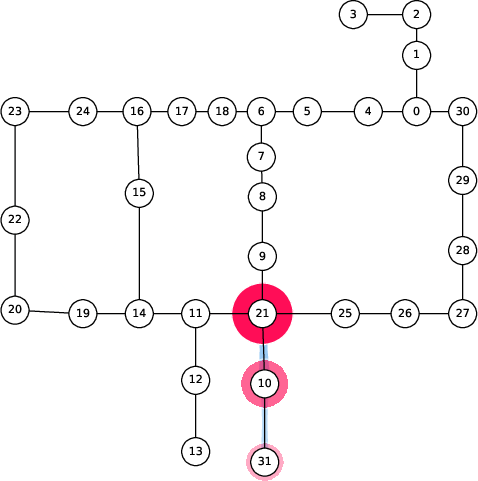
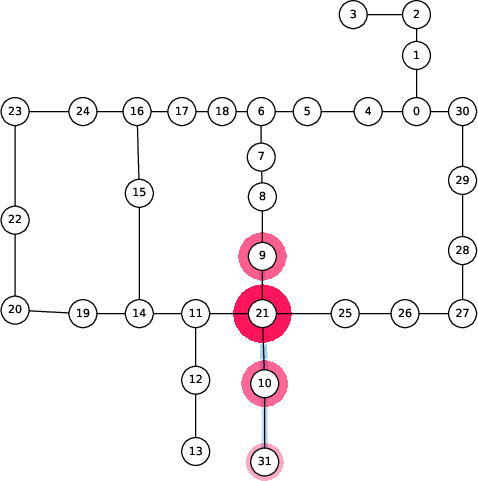
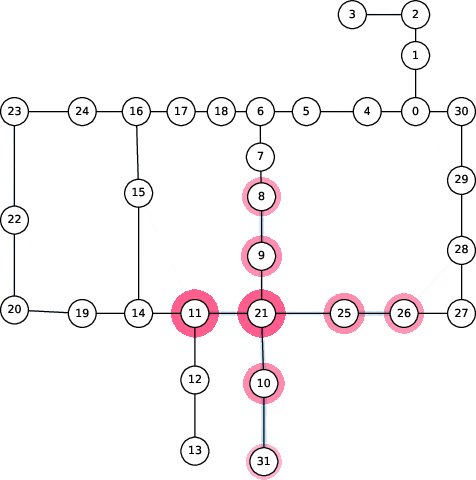
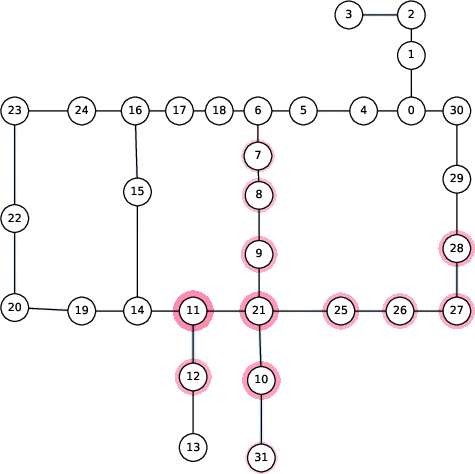
Figure 6: Spread of chlorination through a WDN over time as explained by 2-SII.
Architectural Trade-offs and Limitations
The method requires linear global pooling and output layers, which is standard in many GNN benchmarks but excludes architectures with deep readouts. Empirical comparison shows that non-linear readouts can induce interactions outside the receptive field, violating the locality assumption and leading to fundamentally different SI structures.
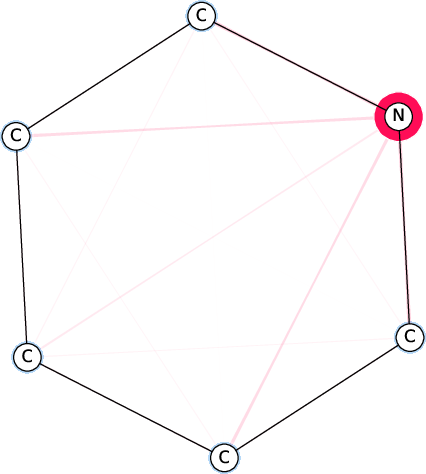
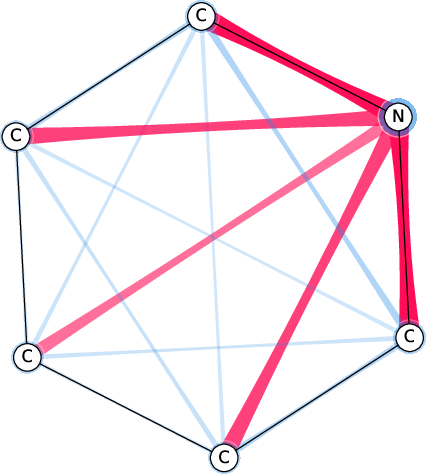
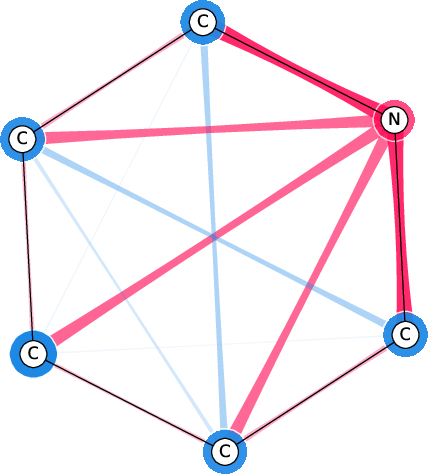
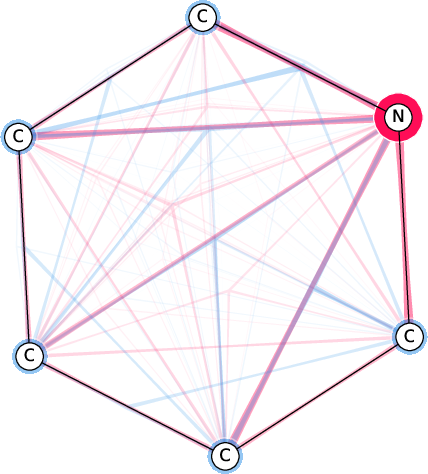
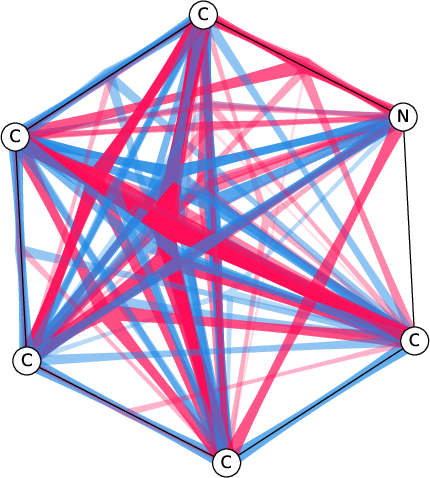
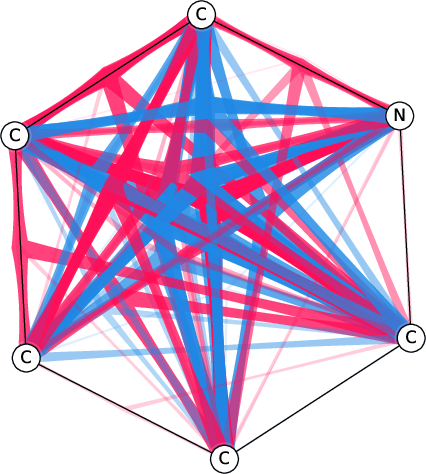
Figure 7: Comparison of the SI-Graphs for pyridine molecule across GCN, GIN, and GAT architectures; all models accurately predict non-benzene.
Implementation Considerations
- Computational Requirements: For sparse graphs and shallow GNNs, exact SIs are tractable; for dense graphs, approximation is necessary.
- Deployment: The method is implemented in the shapiq package and is compatible with PyTorch Geometric; parallelization is recommended for large-scale evaluation.
- Masking Strategy: Baseline feature masking (BSHAP) is used; alternative strategies (e.g., induced subgraphs, edge removal) may emphasize different aspects and are suggested for future work.
- Scalability: The approach is extendable to other models with spatially restricted features, such as CNNs.
Implications and Future Directions
GraphSHAP-IQ provides a principled, efficient framework for computing exact and any-order SIs in GNNs, enabling interpretable decomposition of graph-level predictions. The locality-induced sparsity is a key insight, with practical impact on explainability in domains such as chemistry, biology, and infrastructure networks. The findings challenge the implicit assumption in many GNN XAI methods that interactions outside receptive fields are negligible, especially for non-linear readouts. Future work should explore alternative masking strategies, extend the approach to other architectures, and develop novel approximation methods tailored to the sparsity structure.
Conclusion
The paper establishes that, under linearity assumptions, the complexity of computing SIs in GNNs is determined solely by the receptive fields, enabling exact explanations for large graphs. GraphSHAP-IQ and its interaction-informed variants substantially reduce computational cost and improve estimation quality compared to model-agnostic baselines. Visualization of SIs as SI-Graphs provides actionable insights into model reasoning, with demonstrated utility in scientific and engineering applications. The approach sets a new standard for interpretable GNN explanations, with broad implications for XAI research and practice.































