- The paper introduces a diffusion-based class prompt encoder that eliminates manual prompt generation for efficient, autonomous segmentation.
- It employs an uncertainty-aware joint optimization with multiple loss functions to enhance generalization across various medical modalities.
- Experimental results on datasets like AbdomenCT-1K, BraTS, and Kvasir-SEG reveal superior DSC and NSD scores compared to traditional SAM-based methods.
Diffusion-empowered AutoPrompt MedSAM
AutoMedSAM demonstrates a novel approach to overcoming the challenges in medical image segmentation posed by traditional models, particularly those based on the Segment Anything Model (SAM) framework. The proposed method effectively addresses the need for labor-intensive manual prompts and enhances usability by integrating semantic labels, thus widening its applicability to clinical environments.
Introduction and Challenges
Traditional SAM-based models rely heavily on manual prompt generation, demanding significant effort from medical experts and often limiting their deployment in real-world clinical settings. Furthermore, the lack of semantic labeling in output segmentation masks restricts their utility for non-experts, who may struggle to interpret raw binary masks without clear contextual labels (Figure 1).
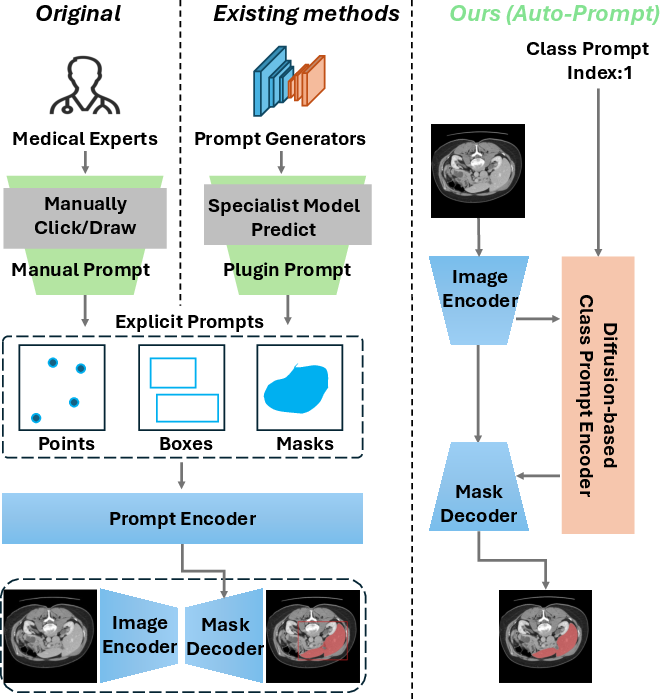
Figure 1: Comparison with SAM-based models.(Left) The original SAM model relies on manual prompts from medical experts, restricting its usability and scenarios. (Middle) Current SAM-based methods employ specialist models for prompt generation, but these models are organ- or lesion-specific, limiting SAM's generalizability. (Right) Our method introduces an automatic diffusion-based class prompt encoder, removing the need for explicit prompts, adding semantic labels to masks, and enabling accurate, end-to-end segmentation for non-experts in diverse medical contexts.
Technical Approach
AutoMedSAM introduces a diffusion-based class prompt encoder, which replaces manual interactions with automated, end-to-end segmentation capabilities (Figure 2). The encoder generates prompt embeddings guided by class indices, integrating semantic structure and enhancing the prediction pipeline with meaningful knowledge. This approach eliminates the need for manual prompts and facilitates a streamlined, fully autonomous segmentation process.
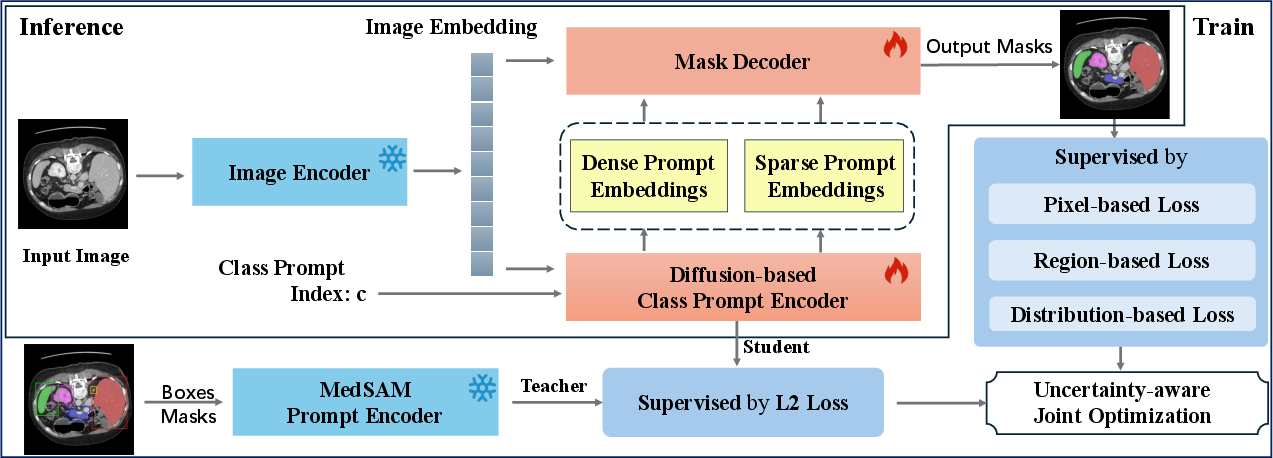
Figure 2: An overview of the AutoMedSAM. AutoMedSAM generates dense and sparse prompt embeddings through a diffusion-based class prompt encoder, eliminating the need for explicit prompts. During training, we employ an uncertainty-aware joint optimization strategy with multiple loss functions for supervision, while transferring MedSAM's pre-trained knowledge to AutoMedSAM. This approach improves training efficiency and generalization. With end-to-end inference, AutoMedSAM overcomes SAM's limitations, enhancing usability and expanding its application scope and user base.
Diffusion-Based Class Prompt Encoder
The diffusion mechanism is central to AutoMedSAM's design. By utilizing a forward conditional generation process, class prompts are projected and interwoven with noise generation steps, enabling the encoder to focus more effectively on class-specific features (Figure 3). The encoder-decoder framework, with two independent branches, extracts both local and global features, contributing to precise prompt embedding generation necessary for accurate mask outputs.
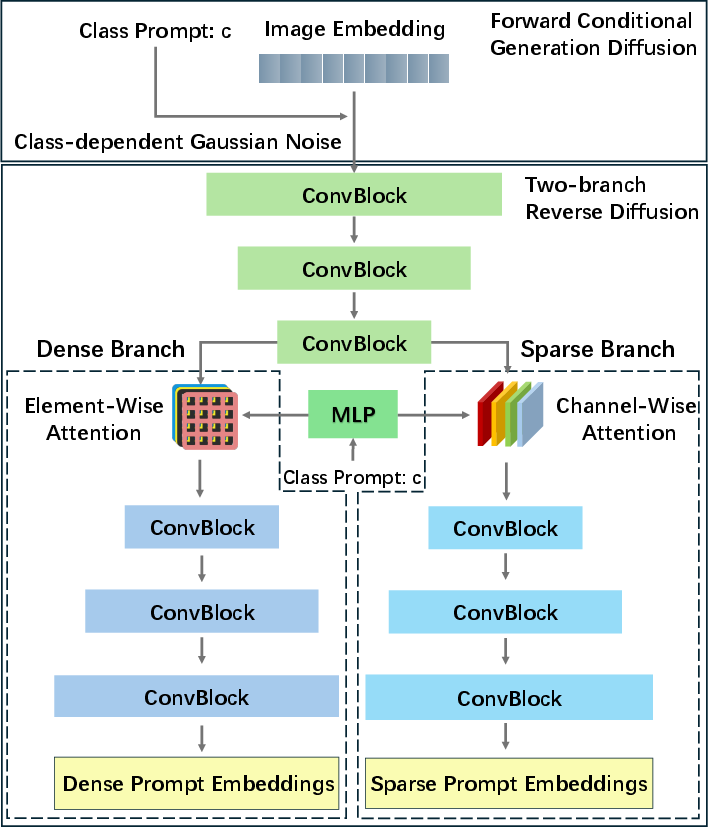
Figure 3: Structure of the diffusion-based class prompt encoder. It is designed with an encoder and two independent decoder branches to extract local and global features, based on the practical significance of sparse and dense prompts.
Uncertainty-Aware Joint Optimization
Adding to its robust feature extraction capabilities, AutoMedSAM employs an uncertainty-aware joint optimization strategy. This approach integrates multiple loss functions, including pixel-based, region-based, and distribution-based losses, to maximize the model's generalization ability. By leveraging MedSAM's pre-trained knowledge alongside these losses, AutoMedSAM shows remarkable adaptability across various medical image modalities.
Experimental Evaluations
AutoMedSAM significantly outperforms existing models on datasets like AbdomenCT-1K, BraTS, and Kvasir-SEG, achieving superior Dice Similarity Coefficient (DSC) and Normalized Surface Distance (NSD) scores. The model's efficacy is particularly evident in settings where traditional models struggle due to the need for explicit prompts or poor generalization across different medical contexts (Figure 4 and Figure 5).
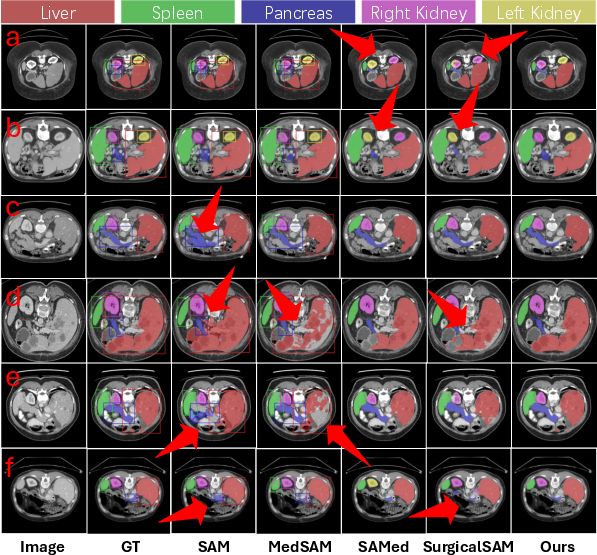
Figure 4: The qualitative results of AutoMedSAM and other comparison models on AbdomenCT-1K. The bounding box represents the input prompt.
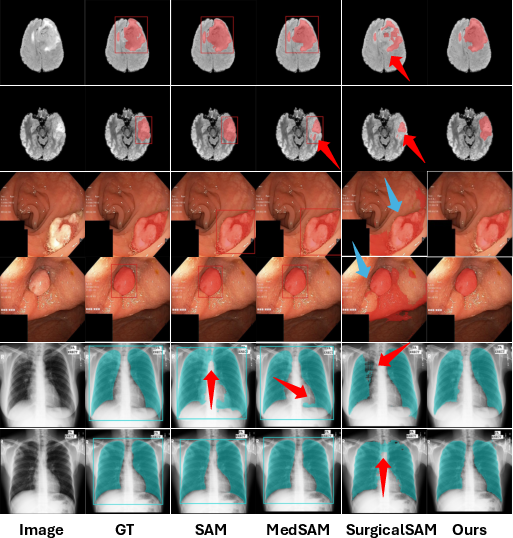
Figure 5: The qualitative analysis results of AutoMedSAM and other comparison models on BraTS, Kvasir-SEG, and Chest-XML.
The study reveals that segmentation accuracy correlates with prompt box precision, highlighting how explicit prompt errors can hinder model performance. AutoMedSAM's approach circumvents these limitations by embedding semantic class information directly, thus reducing dependency on exact prompt sizing (Figure 6).
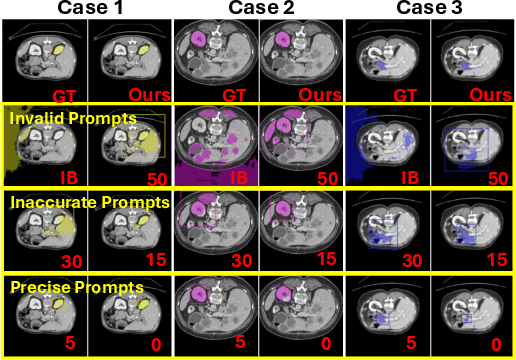
Figure 6: The effect of different sized prompt boxes on segmentation masks. The marked numbers indicate the offset pixel size of the prompt box. IB represents the image boundary.
Conclusion
AutoMedSAM represents a significant advance in medical image segmentation technology, offering a more efficient and adaptable framework than traditional SAM-based models. By automating prompt generation and integrating semantic knowledge, this model extends its utility to non-expert users and enhances its applicability in diverse clinical settings. Future work could focus on scaling this approach across larger datasets to further improve its robustness and practical adoption in healthcare systems.







