- The paper introduces Flowr, a generative model that employs flow matching for de novo 3D ligand design with structural conditioning and efficient interaction recovery.
- It demonstrates superior performance over existing models by achieving higher RDKit- and PoseBusters-validity scores and up to 70-fold faster inference speeds.
- Flowr.multi extends the model's capability by enabling conditional generation for scaffold elaboration and fragment-based design, significantly enhancing interaction recovery rates.
FLOWR: Flow Matching for Structure-Aware De Novo, Interaction- and Fragment-Based Ligand Generation
This essay provides a detailed examination of the paper "FLOWR: Flow Matching for Structure-Aware De Novo, Interaction- and Fragment-Based Ligand Generation" (2504.10564). The paper introduces Flowr, a generative model for the de novo generation and optimization of 3D ligands, offering advantages in structural conditioning, interaction recovery, and computational efficiency.
Introduction
Structure-Based Drug Discovery (SBDD) seeks to exploit the 3D structures of macromolecules to rationally design bioactive compounds, addressing challenges in accurately predicting binding poses and exploring chemical space. Traditional methods, such as molecular docking, often rely on simplified interaction approximations. Modern generative models, particularly diffusion models, have shown potential in capturing complex protein-ligand interactions for novel ligand design. However, iterative stochastic sampling methods, while efficient for exploration, often produce strained or non-drug-like molecules and can be computationally expensive. In this study, a novel model, Flowr, is introduced, which employs flow matching, integrating continuous and categorical data with equivariant optimal transport for the de novo generation of ligands. Further, Flowr.multi is proposed for conditional generation tasks, handling various interaction profiles and chemical substructures (Figure 1).
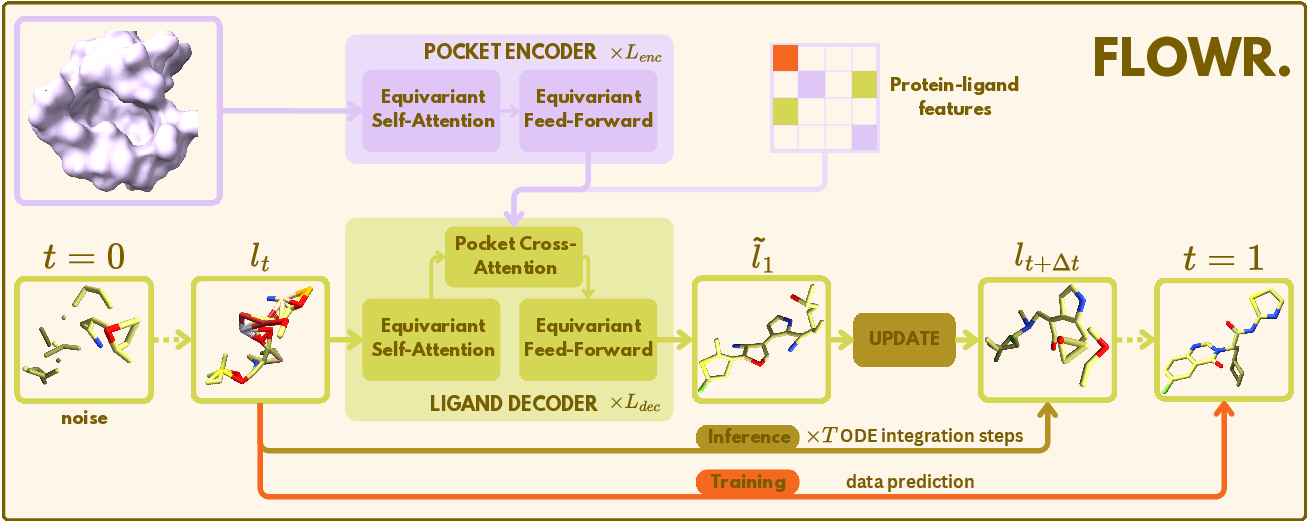
Figure 1: Overview of~Flowr. Schematical overview of the~Flowr model for 3D ligand generation. A protein pocket is encoded and passed, along with the noisy ligand lt, into the ligand decoder, which is trained to produce a denoised ligand l~t. A mixed continuous and categorical flow matching integration scheme pushes lt towards the data distribution to generate a sample l1~.
Spindr: A New Benchmark Dataset
The evaluation of novel SBDD models relies heavily on data quality and appropriate benchmarks. Existing datasets like PDBBind and CrossDocked2020 suffer from data quality issues and structural defects, hindering their utility in realistic drug discovery applications. The Spindr dataset was created to address these problems, offering 35,666 high-quality, refined protein-ligand complexes derived from the Plinder dataset. The new benchmark eliminates structural defects, infers accurate interaction profiles, adjusts protonation states, and mitigates data leakage, thus enabling a more robust assessment of generative models. This dataset sets new standards for 3D generative modeling benchmarks, particularly with respect to ligand binding poses and data integrity.
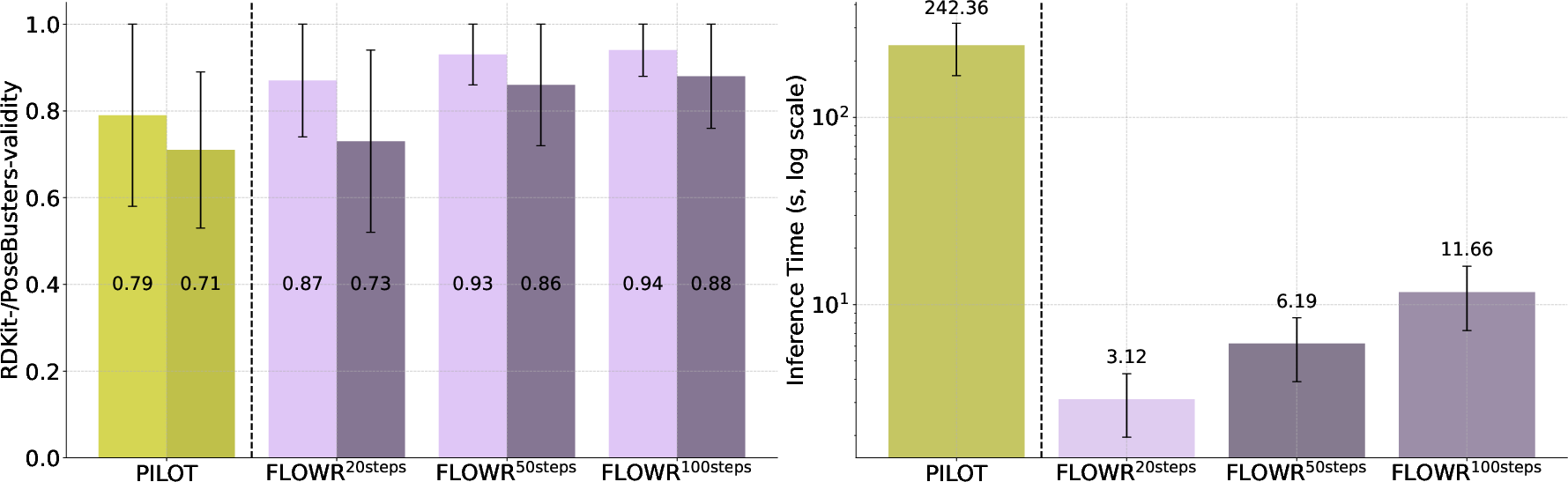
Figure 2: Comparison of~Pilot and~Flowr on validity and inference speed. We compare~Pilot and~Flowr in terms of RDKit- and PoseBusters-validity and inference speed. Results for~Flowr are reported using three different inference step settings: 20, 50, and 100 steps.
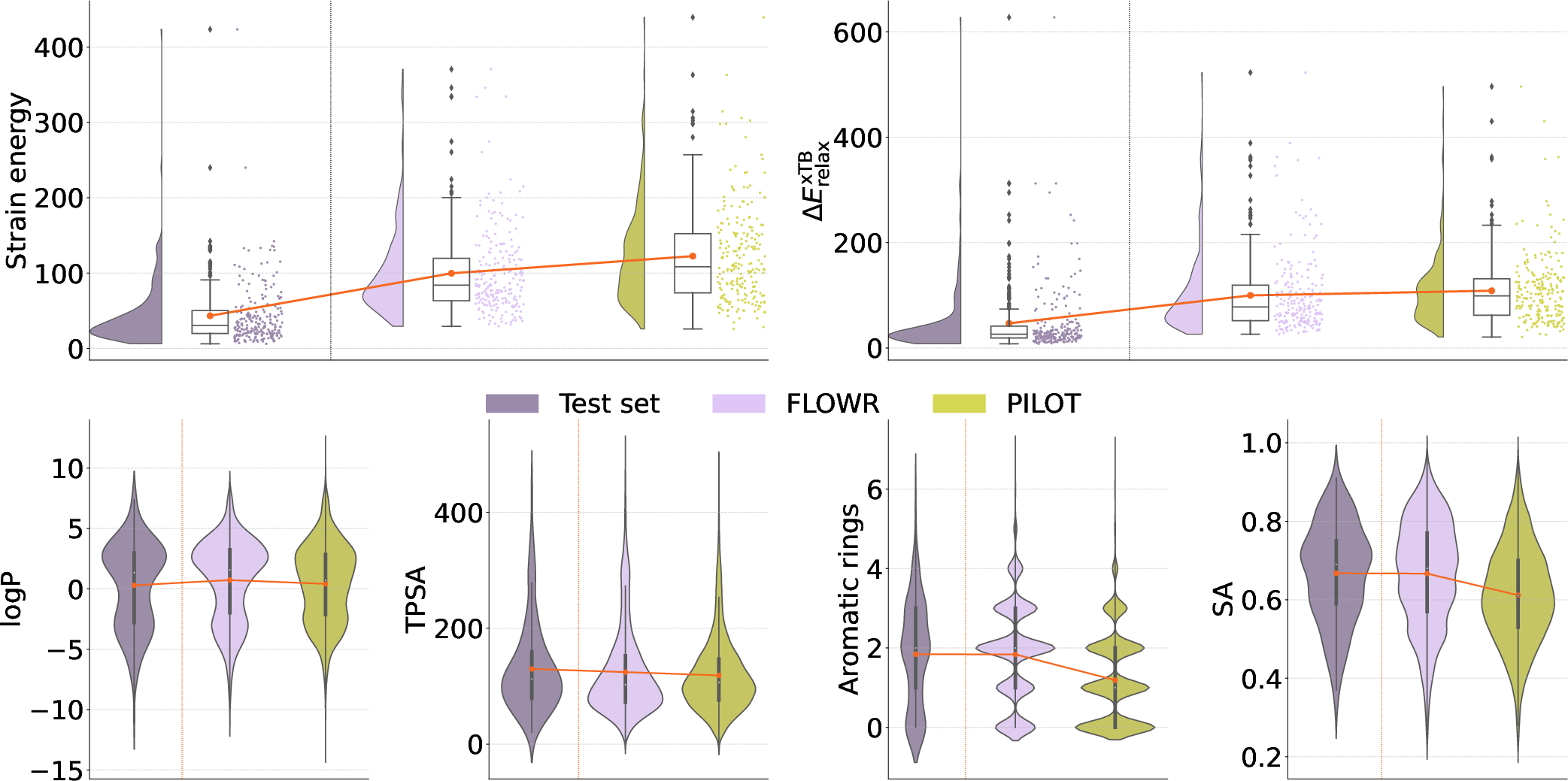
Figure 3: Comparison of~Pilot and~Flowr on molecular properties.
FLOWR Model Architecture
The Flowr model employs flow matching techniques for efficient, de novo ligand design, focused on structural constraints. Flowr utilizes the SemlaFlow architecture, augmented with a pocket encoder and a cross-attention module within the ligand decoder, enabling structural conditioning on protein pockets and optional desired interaction profiles or chemical substructures. Flowr simultaneously generates continuous (3D coordinates) and discrete features (atom types, bond orders) using flow matching for continuous and discrete data (Figure 1).
Flowr.multi offers an additional layer of versatility by allowing multi-purpose conditional generation without retraining or re-sampling. This model uses predefined interaction profiles and chemical substructure conditions for scaffold elaboration, scaffold hopping, and fragment-based design tasks. These autoencoders adhere to the learned data distribution while maintaining computational efficiency.
Empirical Evaluations
When benchmarked on the CrossDocked2020 dataset, Flowr outperformed existing models including Pocket2Mol, DiffSBDD, TargetDiff, DrugFlow, and Pilot in all major metrics, such as PoseBusters-validity (0.92 ± 0.22) and significantly improved inference speed (12.05 ± 8.01 seconds, achieving up to 70-fold faster performance than its competitors) (Figure 2). Flowr also demonstrated a reduction in strain energy values, indicating a superior ability to generate stable conformations with a better geometric alignment with actual structures.
Subsequent comparisons on the Spindr dataset also illustrate Flowr's superior RDKit-validity and PoseBusters-validity when compared to Pilot, with average RDKit-validity scores of 0.94 ± 0.24 versus 0.79 ± 0.39 for Pilot. Furthermore, Flowr's mean Vina scores suggested improved pose accuracy in docking assessments.
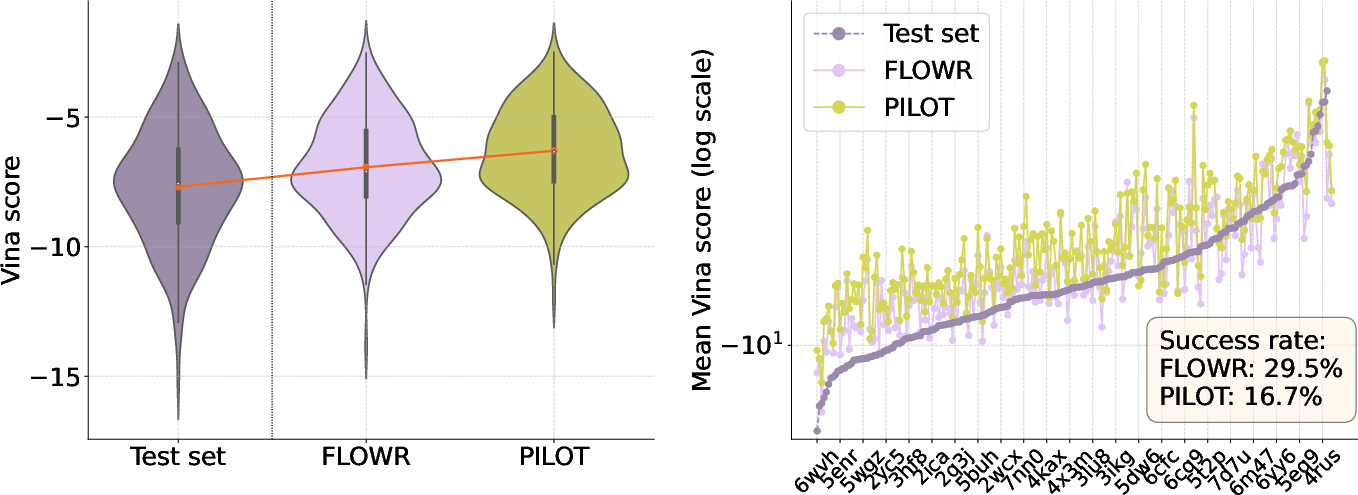
Figure 4: Comparison of~Pilot and~Flowr on AutoDock-Vina scores.
Flowr.multi's interaction-conditional generation notably enhanced interaction recovery rates, achieving higher accuracy than Pilot, particularly in scenarios involving explicit hydrogen modeling (Figure 5). On the Spindr test set, Flowr.multi achieved an average interaction recovery rate of 76.1%, highlighting its efficacy for fragment-based design tasks.
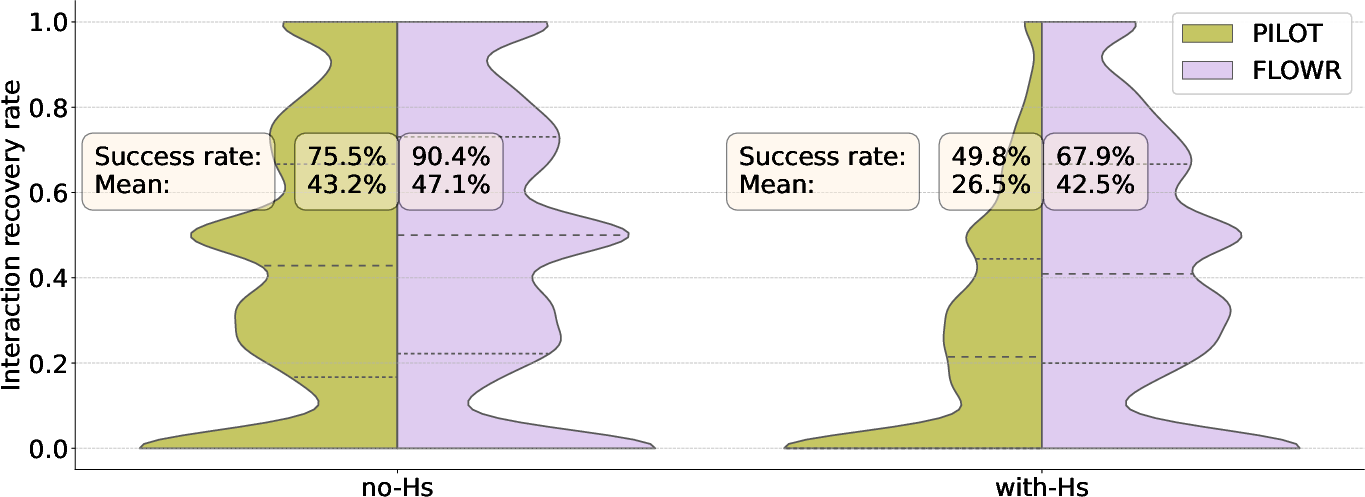
Figure 5: Comparison of~Pilot and~Flowr on interaction recovery.
Flowr.multi: Multi-Purpose Conditional Generation
Flowr.multi demonstrates proficiency in generating ligands conforming to interaction profiles or chemical substructures, evidenced by empirical results on randomly selected targets (5YEA and 4MPE) (Figures 7 and 8). The model maintained high PB-validity and demonstrated improved interaction recovery, indicating that it effectively generated ligands with desired characteristics, suitable for scaffold elaboration, fragmentation, and structure-guided optimization, thus advancing the practical applications of generative models in drug discovery.
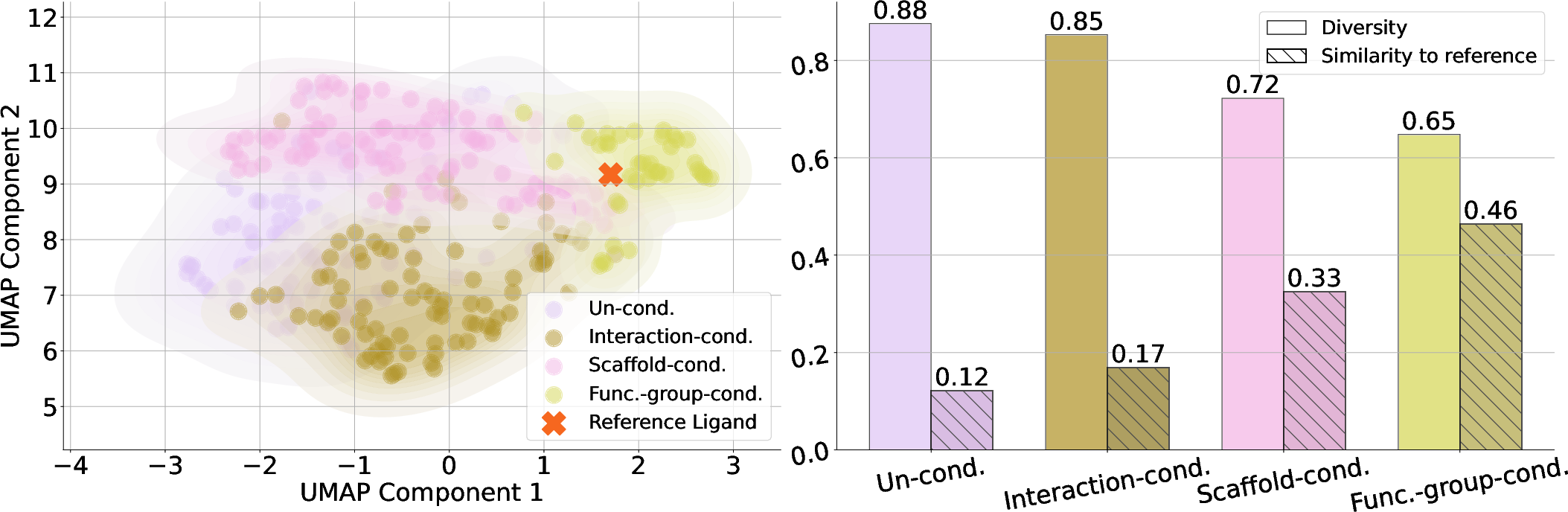
Figure 6: Evaluation of chemical space coverage on 5YEA with~Flowr.multi.
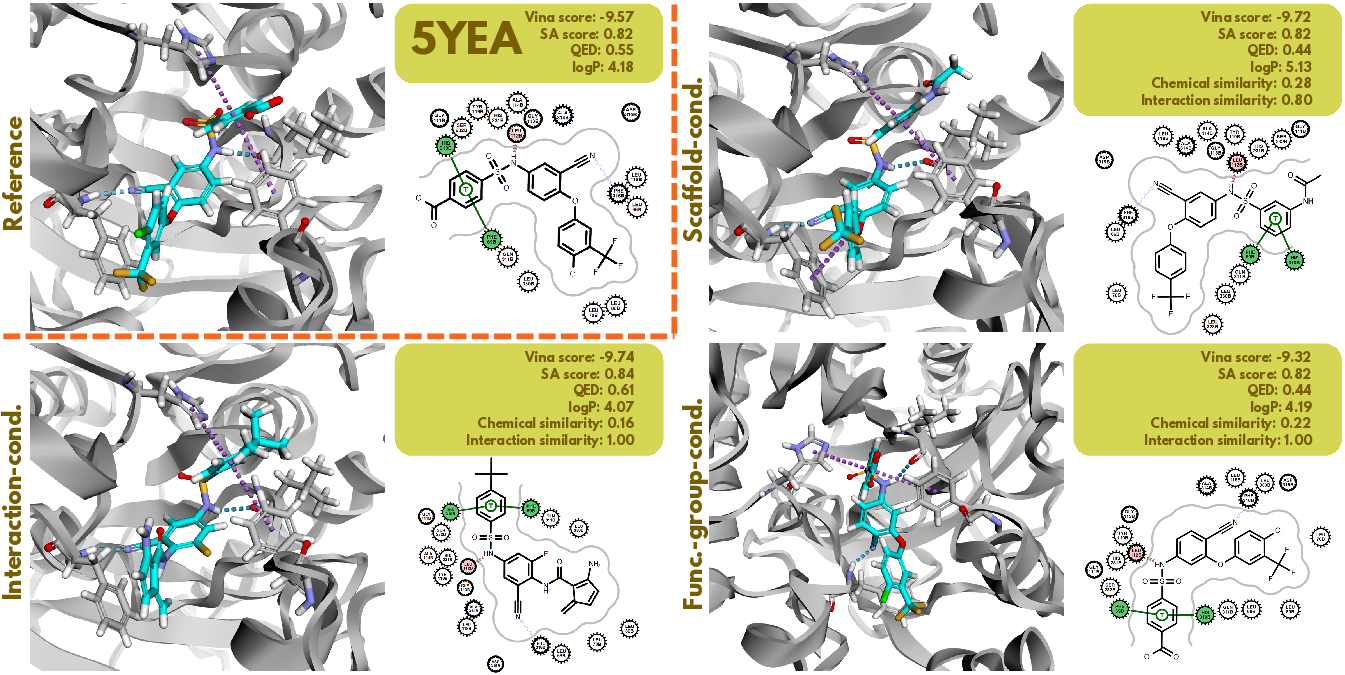
Figure 7: Visualization of conditional generation on 5YEA with~Flowr.multi.
Conclusion
Flowr and its multi-purpose variant, Flowr.multi, advance the landscape of AI-driven ligand generation for structure-based drug design, directly addressing limitations found in diffusion-based generative models with improved validity, pose accuracy, and computational efficiency. The introduction of the Spindr dataset provides a critical resource for evaluating generative models, emphasizing data quality and minimizing leakage. Flowr.multi's performance across diverse modalities in ligand generation substantiates its potential in facilitating targeted therapeutic development through AI, suggesting robust applications in hit exploration and lead optimization while setting new benchmarks for future research.






