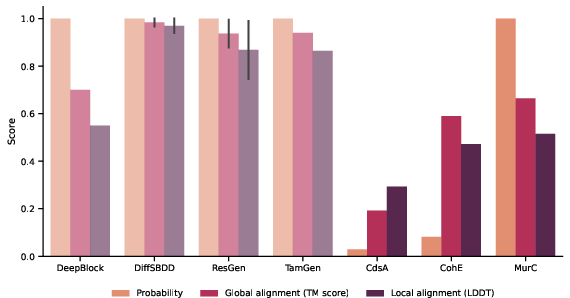- The paper introduces a comprehensive AI pipeline for antibiotic discovery, spanning target identification to compound validation.
- The study evaluates six state-of-the-art 3D structure-aware models, with DeepBlock and TamGen leading in compound validity and scaffold diversity.
- It leverages structure-based clustering and rigorous post-processing to filter candidates, establishing a replicable approach for overcoming antibiotic resistance.
AI-guided Antibiotic Discovery Pipeline from Target Selection to Compound Identification
This essay provides a comprehensive analysis and summary of the AI-guided pipeline designed for antibiotic discovery, highlighting the methodologies employed, evaluation of generative models, and steps toward practical drug development. The study integrates advanced methods in structure-based drug design (SBDD), leveraging deep learning techniques for potential therapeutic discovery.
Introduction
Antibiotic resistance is a critical challenge in global health, necessitating innovative approaches in drug discovery. This study introduces an end-to-end AI-driven pipeline for antibiotic discovery, beginning with target identification and concluding with compound realization. Using structure-based clustering, it identifies potential antibacterial targets devoid of human homologs. The paper evaluates six state-of-the-art 3D structure-aware generative models (spanning diffusion, autoregressive, GNN, and LLM architectures) on usability, chemical validity, and biological relevance.
Target Identification
Structure-based clustering serves as a foundational step in identifying novel antibacterial targets. Utilizing Foldseek, the clustering of predicted proteomes from various pathogenic bacteria permits the identification of conserved, essential proteins without human analogs (Figure 1). This approach identifies promising targets such as AccD, FtsW, and LolE, which hold potential for pan-strain antibiotic development.
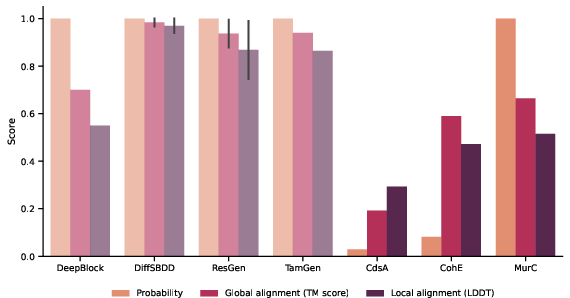
Figure 1: Foldseek search similarity of target proteins to the CrossDocked2020 training set, highlighting structural conservation.
The structural similarity of targets to CrossDocked2020 training sets suggests that generative models trained on similar datasets could yield better prediction performance. Protein structures resembling the training set structures, such as those of MurC, serve as positive controls in validating target selection.
Molecular Generation and Model Evaluation
The study employs six models (DeepBlock, DiffSBDD, Pocket2Mol, ResGen, TamGen, and TargetDiff), each evaluated for ease of implementation, usability, output quality, and structural diversity (Figure 2). The performance of these models in generating valid compounds varied significantly, with DeepBlock showing the highest validity at approximately 90%.
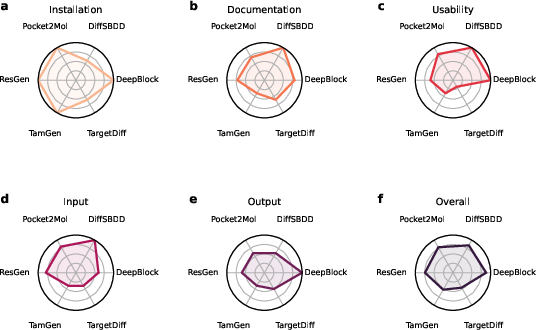
Figure 2: Model implementation ranking reveals differences in ease of use, documentation quality, and output usability.
(Figure 3)
Figure 3: Overview of generated molecules based on model and target protein, illustrating generation variability.
Despite challenges in achieving uniform molecule generation, DeepBlock and TamGen demonstrate a commendable balance between scaffold diversity and structural validity. In comparison, models like Pocket2Mol and ResGen exhibited limitations in chemical space exploration.
Post-Processing and Structural Validation
The study emphasizes extensive post-processing to filter generated compounds through REOS/Dundee alerts, ensuring molecular validity and alignment with the desired chemical spaces for antibiotics (Figures 4 and 5).
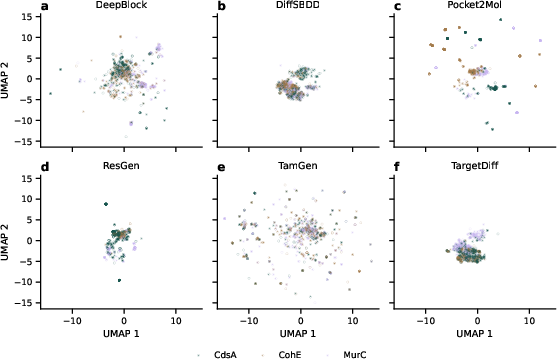
Figure 4: Structural alerts for generated molecules, showcasing Dundee alert analysis based on model.
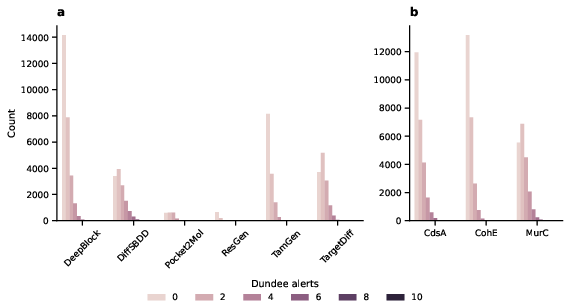
Figure 5: Top-ranking structural alerts highlighting common pitfalls in molecular structures.
The curated pipeline refines compounds based on physicochemical properties, bringing the focus onto the commercially accessible candidates that exhibit favorable binding pocket predictions via AlphaFold 3 (Figure 6).
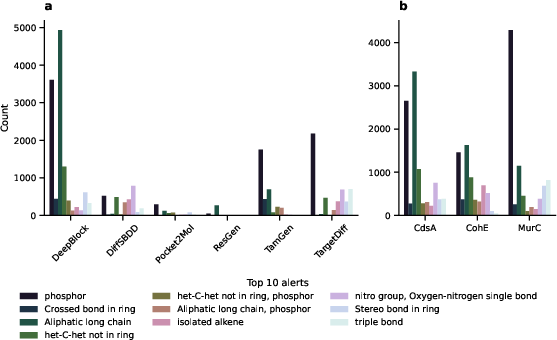
Figure 6: Processing of de novo structures, evaluating molecules through curation and filtering.
Discussion and Conclusion
This study delineates a framework for integrating AI in antibiotic discovery, elucidating the strengths and limitations of contemporary SBDD models. The analysis highlights how AI-driven pipelines can streamline early-stage antibiotic design by focusing on structurally conserved proteins and employing rigorous filtering to enhance compound validity.
Empirical findings suggest that while DeepBlock and TamGen set the benchmark in model performance by generating the highest number of valid candidates, the study underscores the potential for future model improvements, especially concerning structural diversity and usability.
In summary, this work marks a substantive step in leveraging AI for antibiotic discovery, providing a replicable prototype for future research endeavors aiming to overcome antibiotic resistance challenges. The detailed comparison of model outputs and comprehensive post-processing adoption set a robust foundation for continued development in AI-assisted pharmaceutical research.