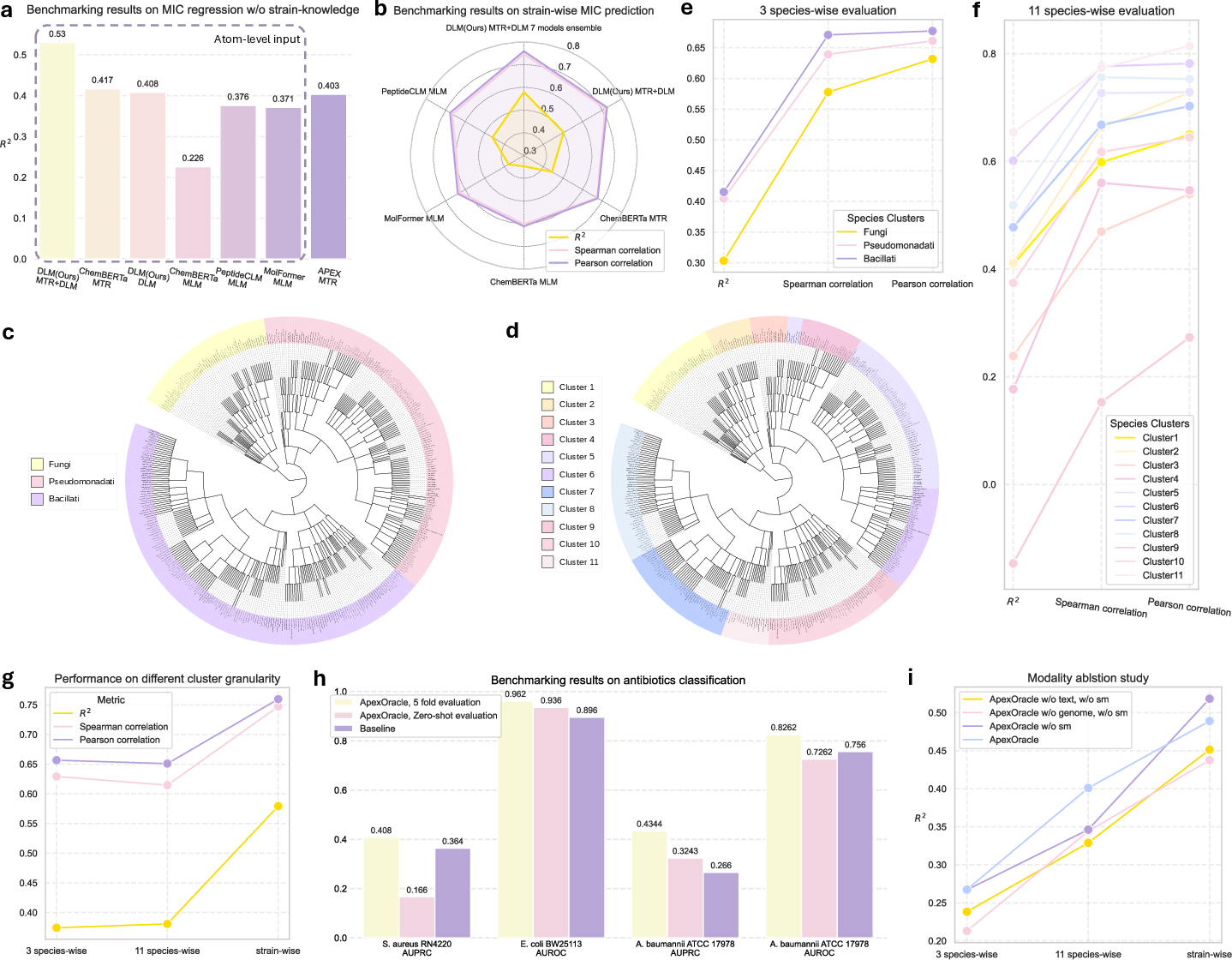
- The paper introduces ApexOracle, an AI-driven framework that integrates genomic, textual, and molecular data for antibiotic prediction and generation.
- It demonstrates a 27.1% increase in prediction accuracy for minimum inhibitory concentrations compared to state-of-the-art models.
- The system successfully generalizes to unseen bacterial strains and generates novel compounds with lower Tanimoto similarity to existing antibiotics.
Predicting and Generating Antibiotics Against Future Pathogens with ApexOracle
The paper "Predicting and generating antibiotics against future pathogens with ApexOracle" presents a novel AI-driven framework, ApexOracle, for predicting and generating effective antibiotics against both known and emerging pathogens. It leverages a unique integration of pathogen genomic, textual, and molecular data to create a unified system that not only forecasts antibiotic efficacy but also generates novel antibiotic candidates. The following sections provide a detailed overview of ApexOracle's architecture, evaluation, and implications.
ApexOracle Architecture
ApexOracle is designed to address the limitations of existing antimicrobial models, which often focus on isolated strain-specific datasets. Its core architecture consists of three key representation modules:
- Genomic Encoder: Utilizes Evo2, a DNA LLM, to encode pathogen genomic data into numerical representations capturing genetic hallmarks.
- Textual Trait Encoder: Employs a fine-tuned Me-LLaMA model to process descriptions of pathogen traits, providing phenotypic context that complements genomic data.
- Molecular Representation Learning and Generation Module: Based on a Diffusion LLM (DLM), this module transforms molecular structures into latent spaces and generates new molecules.
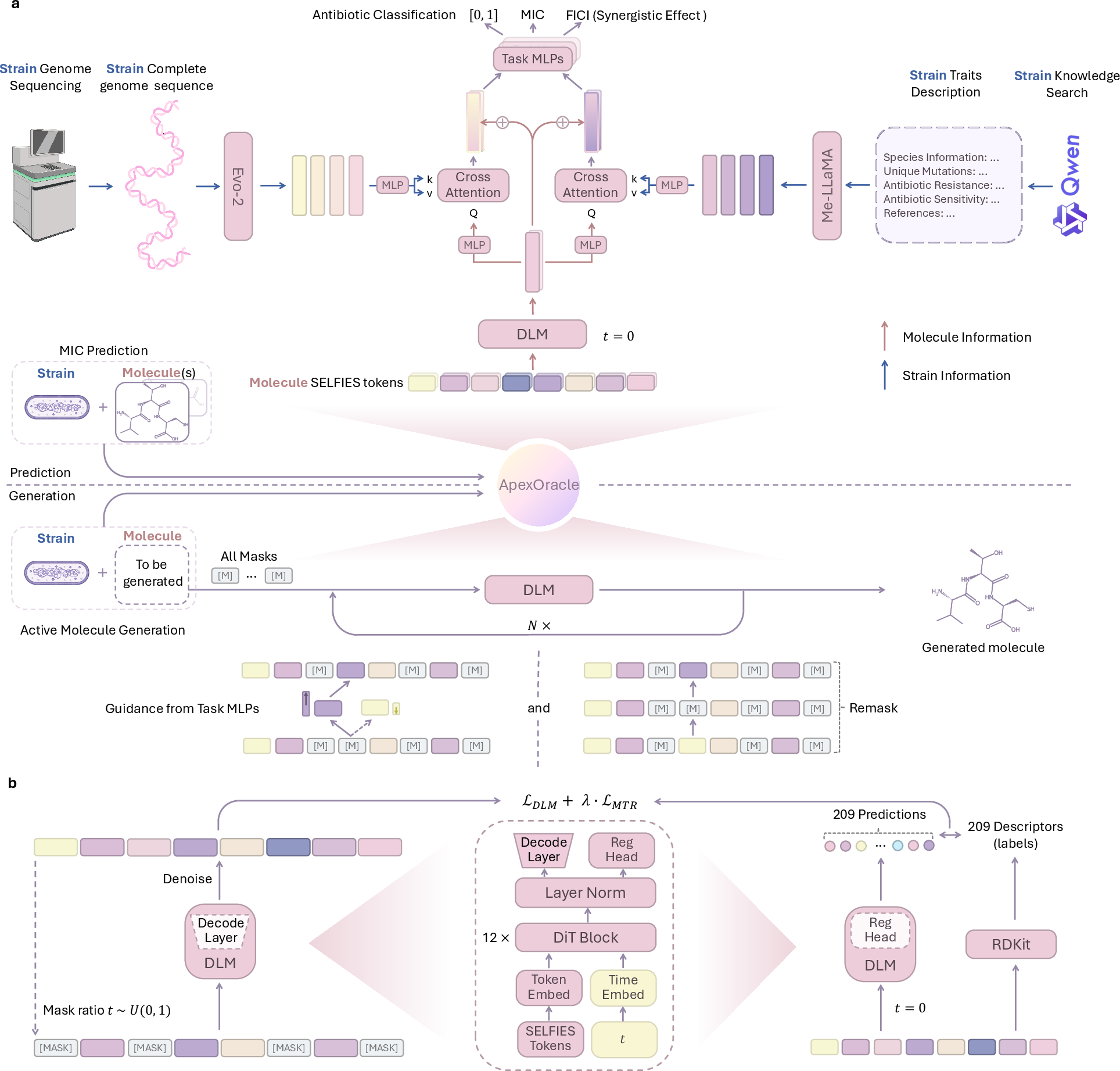
Figure 1: The architecture of ApexOracle and DLM training tasks, illustrating the integration of pathogen strain knowledge for antimicrobial prediction and molecular generation.
ApexOracle's unified framework allows it to predict antimicrobial efficacy and design new compounds within the same architecture, ensuring that generated antibiotics are contextually tailored to specific pathogens.
Evaluation of ApexOracle
The evaluation of ApexOracle demonstrates its superior performance across multiple dimensions of antimicrobial prediction and generation. Key findings include:
Molecule Generation and Novelty
A significant capability of ApexOracle is its ability to generate novel antibiotic candidates. Using predictor-guided generation, the model proposes molecular structures with high predicted potency and structural novelty, as evidenced by lower Tanimoto similarities to existing compounds (Figures 3a-d).
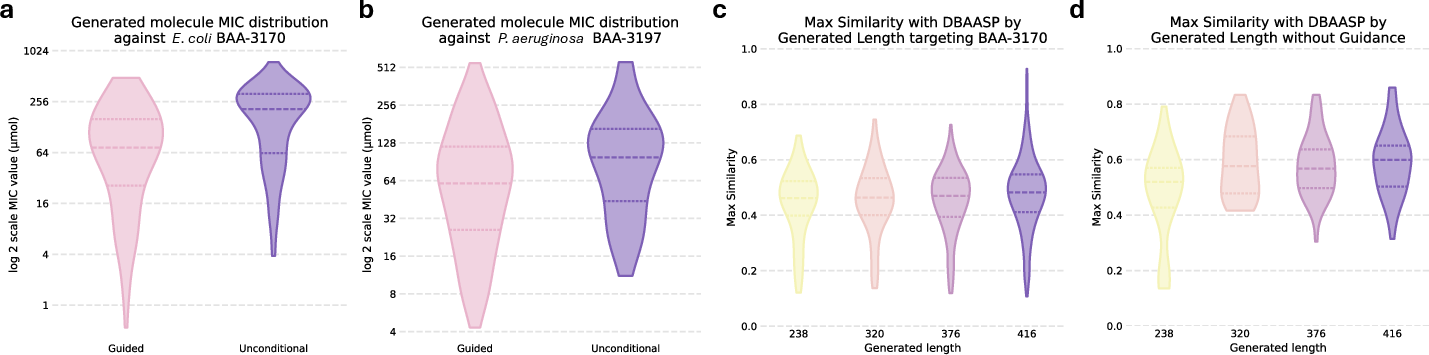
Figure 3: Predicted MIC distributions and structural novelty of generated molecules, highlighting the efficacy of pathogen-guided generation.
Implications and Future Directions
ApexOracle represents a substantial advancement in antimicrobial discovery by integrating multimodal data streams to predict and generate candidate antibiotics. Its ability to generalize to unseen pathogens and explore unconventional chemical spaces offers promising strategies for preemptively combating antimicrobial resistance (AMR).
Future developments could focus on addressing current limitations, such as incorporating toxicity and synthetic feasibility into the generation process and expanding the model to other pathogen types such as viruses. With further refinement, ApexOracle could become a critical component in rapid antibiotic discovery, capable of immediately responding to new infectious threats through AI-driven design.
Conclusion
ApexOracle establishes a new paradigm in antibiotic development, combining pathogen knowledge with advanced AI techniques to predict and design therapeutics against both current and future bacterial threats. It bridges critical gaps in existing methods, offering a scalable and proactive approach to combat AMR and infectious diseases. Continued efforts to enhance its capabilities and integrate real-time data will further align AI-driven methodologies with urgent clinical needs.