- The paper introduces Neuroprobe, a standardized benchmark that transforms 40 hours of iEEG data from naturalistic movie-viewing into 15 distinct decoding tasks.
- The methodology leverages multimodal analyses with within-session, cross-session, and cross-subject splits to assess auditory, visual, and language processing in the brain.
- The results show that traditional linear baseline models, using spectrogram inputs and Laplacian re-referencing, often outperform complex models, highlighting practical implications for neural interface development.
Neuroprobe: Evaluating Intracranial Brain Responses to Naturalistic Stimuli
Introduction
The study introduces Neuroprobe, a benchmark designed for evaluating decoding tasks using intracranial EEG (iEEG) data aligned with naturalistic stimuli. This benchmark addresses the lack of standardized evaluation frameworks for iEEG recordings, facilitating the development and evaluation of neural foundation models for brain-computer interfaces (BCIs) and neurological research.
Neuroprobe is built upon the BrainTreebank dataset, which includes 40 hours of iEEG recordings from 10 subjects engaged in a naturalistic movie-viewing task. The high-resolution neural data enables insights into the temporal and spatial localization of language processing tasks, offering a mine of neuroscience insights. Neuroprobe serves to localize when and where computations for various language features occur in the brain and provides a framework for comparing neural decoding models systematically.
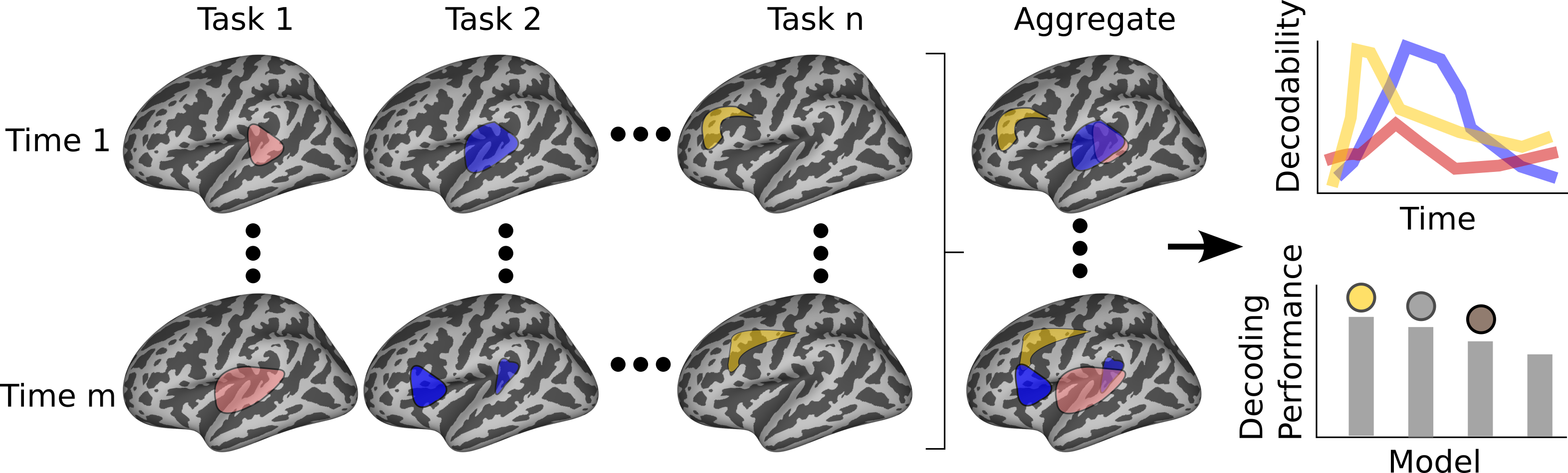
Figure 1: Overview of Neuroprobe's goals, highlighting its dual function in neural decoding task analysis and standardized model evaluation.
Methodology
Neuroprobe uses data from 26 movies, observed by subjects with stereoelectroencephalography electrodes implanted in different brain regions. The local field potentials recorded are used to define a suite of 15 visual, auditory, and language decoding tasks. This transform of raw neural data into decoding tasks offers a standardized evaluation benchmark.
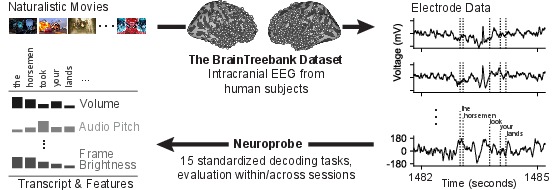
Figure 2: Transition from raw data to structured decoding tasks aligned with movie stimuli, encompassing auditory, language, and visual domains.
The evaluation is structured into three splits: within-session, cross-session, and cross-subject, each assessing a different aspect of model generalization. The cross-subject split poses the most significant challenge due to varying electrode placements among subjects.
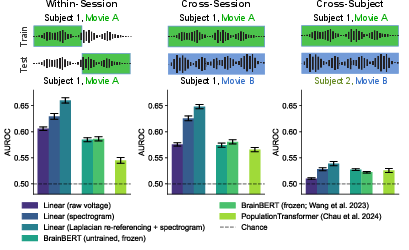
Figure 3: Neuroprobe evaluates model performance across session types, illustrating the challenges posed by cross-subject data variability.
Results
Spatial and temporal analyses reveal how multimodal stimuli are processed across the brain. Linear decodability shows that auditory and linguistic tasks, such as sentence onset, are decodable with high accuracy in the superior temporal gyrus. Visual features demonstrate distinct decodability patterns in the occipital lobe.
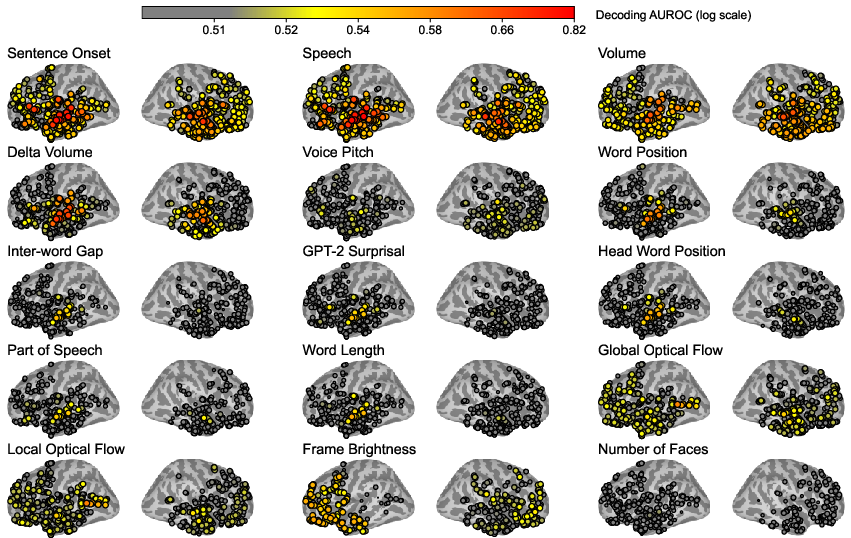
Figure 4: Visualization of multimodal stimulus processing trends across brain regions, highlighting task-specific decodability hotspots.
Temporal analysis tracks the evolution of sensory processing, with linguistic features showing peak decodability close to word onset and visual features lagging behind. This asynchronous processing is reflected in the distinct time courses of various tasks.
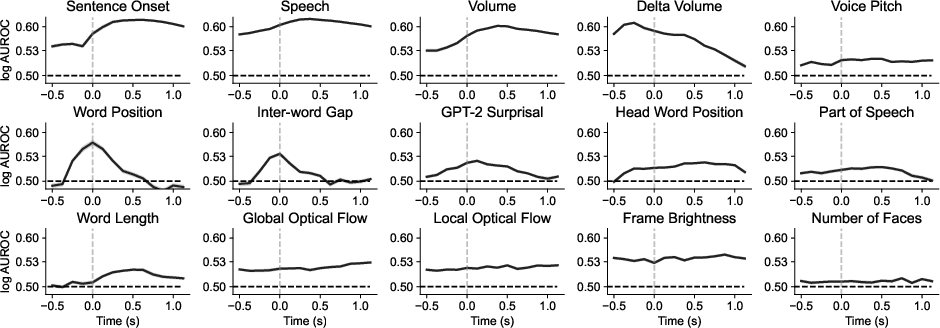
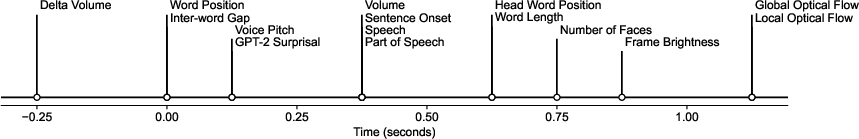
Figure 5: Time-tracking of sensory processing, revealing the staggered decodability of audio, language, and visual features in the brain.
The benchmark results indicate strong performance for traditional linear baseline models, particularly when using spectrogram inputs and Laplacian re-referencing, often outperforming more complex foundation models like BrainBERT and PopulationTransformer.
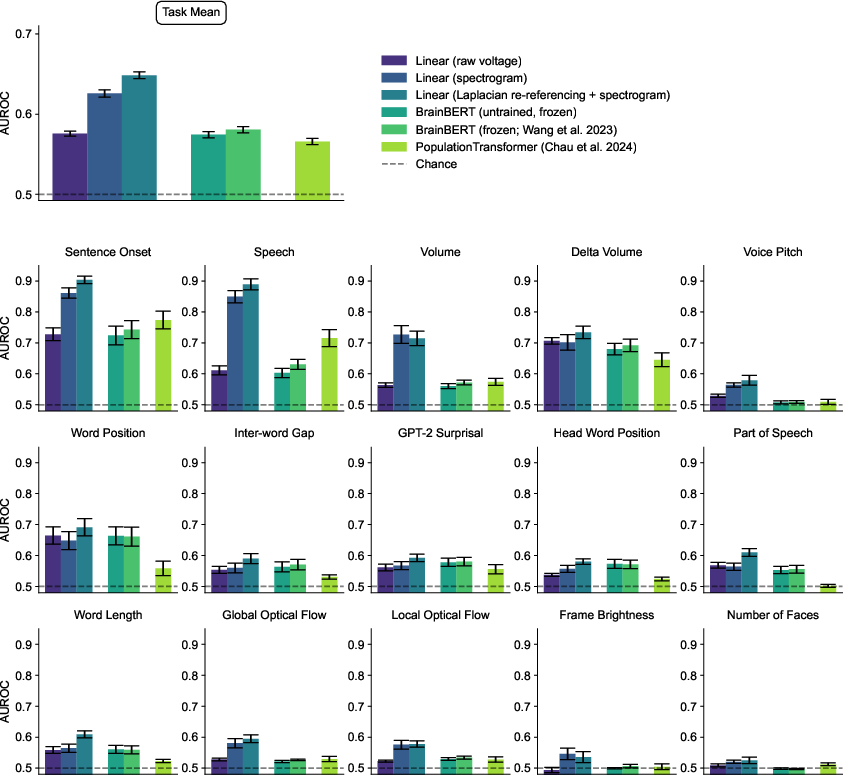
Figure 6: Performance of baseline and frontier models on Neuroprobe's cross-session tasks, showcasing comparative effectiveness.
Discussion
Neuroprobe offers a robust framework for advancing iEEG foundation models, promoting research into brain-computer interfaces and multimodal sensory processing. It provides standardization benefits akin to popular benchmarks in other domains, lowering the entry barrier for new researchers and fostering community growth.
Despite its innovation, Neuroprobe is limited by the clinical setting of iEEG data collection and a small number of subjects. Future work aims to expand the library of tasks and datasets, enhancing the neurobiological insights derived from this benchmark.
Conclusion
Neuroprobe represents a significant step forward in standardizing neural data analysis, enabling rigorous comparisons across iEEG models. Its implementation opens new avenues for understanding the intricacies of brain computation and promotes the development of advanced neural interfaces. This benchmark is positioned to drive measurable progress in the evaluation and development of iEEG-based foundation models, potentially leading to significant innovations in neuroscience and clinical applications.








