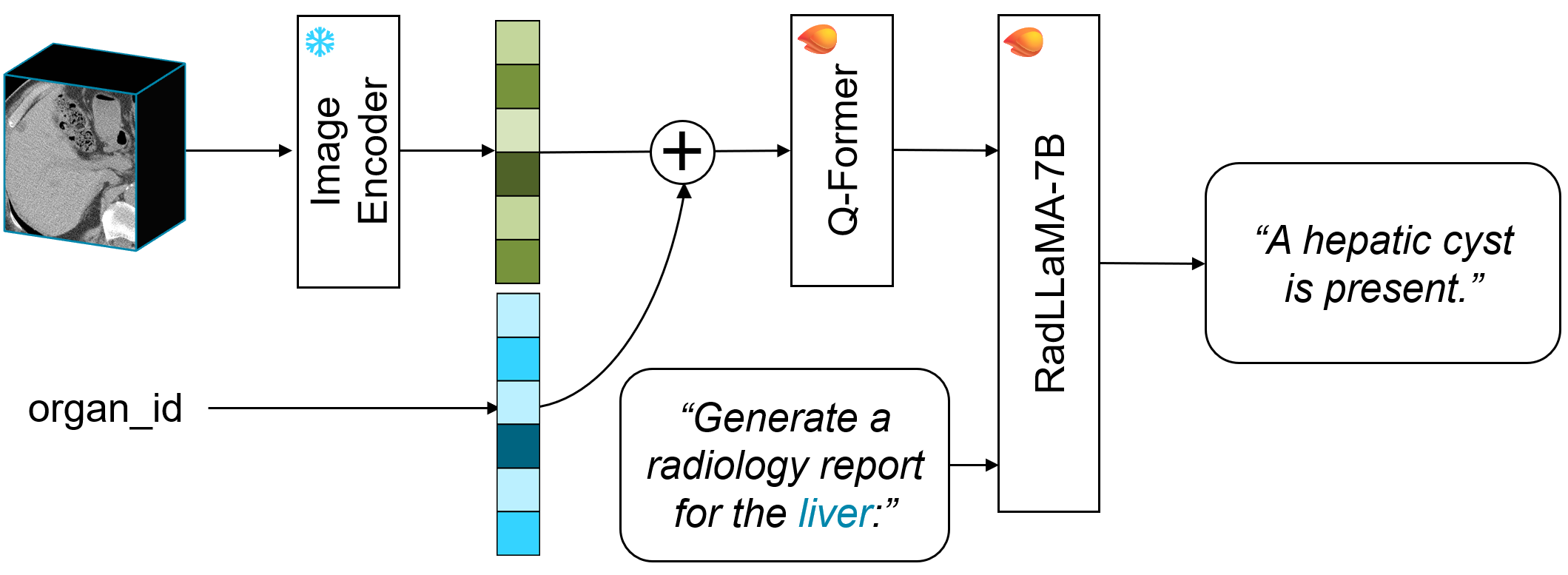
- The paper introduces an organ-separated approach that decomposes whole-body CT volumes into organ-specific units to overcome computational bottlenecks.
- The paper leverages a three-stage training process—self-supervised pre-training, organ-wise contrastive learning, and vision–language fine-tuning—to align CT images with radiology reports.
- The paper demonstrates improved organ-wise lesion classification and report generation, achieving higher F1 scores and robust zero-shot performance compared to prior models.
TotalFM: An Organ-Separated 3D-CT Vision Foundation Model Framework
Introduction and Rationale
TotalFM introduces an organ-separated methodology for 3D-CT vision foundation models, addressing the severe computational bottlenecks inherent to volumetric medical imaging contrastive learning. Prior work in 3D-CT modeling (e.g., CT-CLIP, RadFM, Merlin) consistently encounters a trade-off limiting batch size versus spatial resolution due to high GPU memory requirements. The TotalFM approach decomposes whole-body CT volumes into organ-specific units—emulating radiological workflows and harnessing recent advances in automatic segmentation (TotalSegmentator) and LLM-powered report processing—thus enabling organ-wise high-resolution processing with substantial computational efficiency gains.
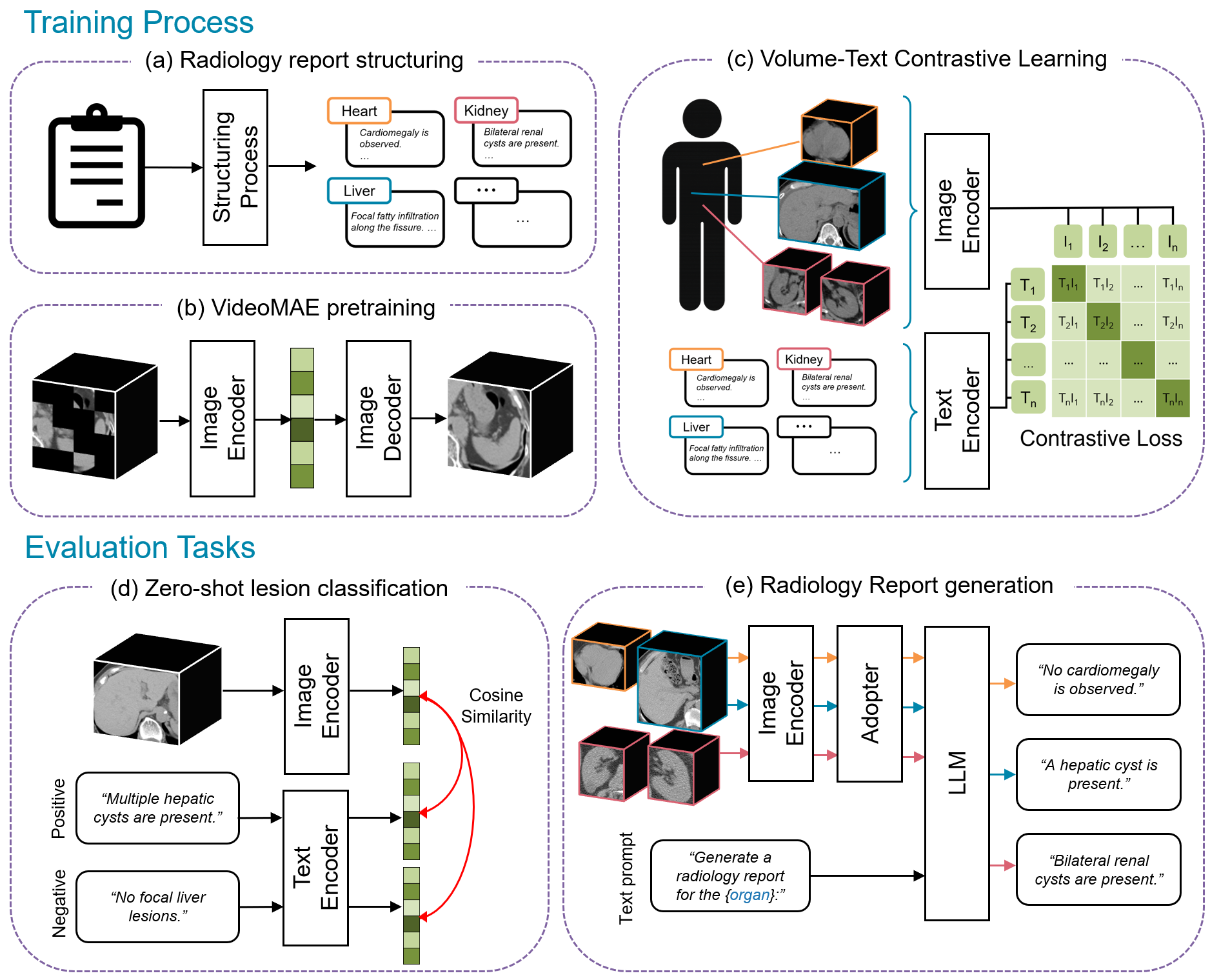
Figure 1: Framework overview—CT volumes and reports are structured to organ-level units, pre-trained with VideoMAE, subjected to organ-specific contrastive learning, and finally configured into a vision–LLM using a Q-Former and LoRA.
Data Pipeline and Automated Volume–Text Pair Construction
Central to TotalFM’s scalability is a fully automated pipeline that generates organ-specific CT volume and report sentence pairs. Original DICOM data are segmented by TotalSegmentator to derive clean organ masks. Simultaneously, associated radiology reports are parsed and split into atomic findings; each finding is classified to its imaging region and contrast phase by a prompt-based LLM, and matched by rule-based logic to the best-fitting CT series. The resulting pairs are pruned—findings without high-confidence anatomical correspondence are discarded—and negative examples are produced by templating for unmentioned-but-present organs, ensuring robust negative sampling for contrastive learning.
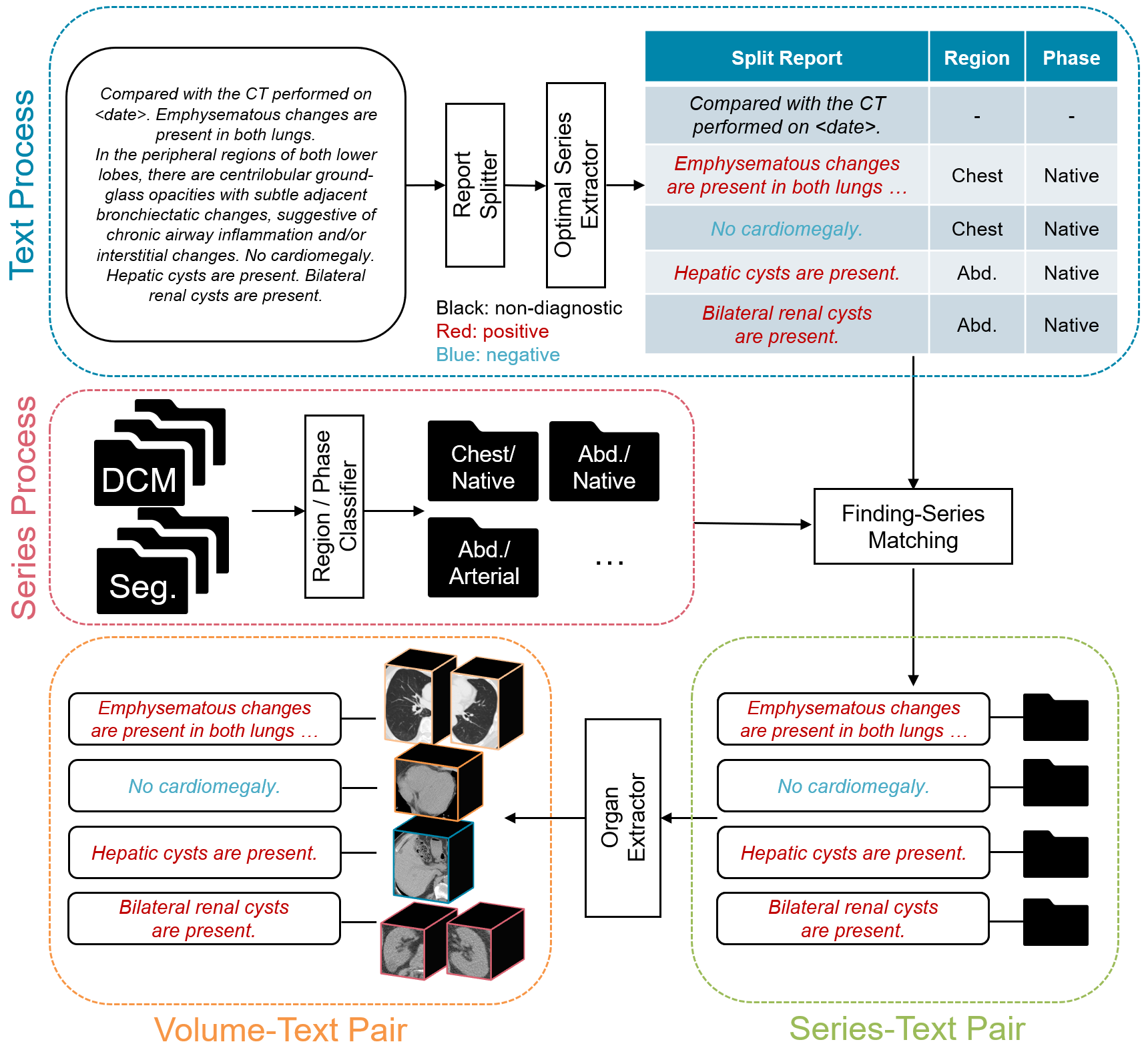
Figure 2: Automated pipeline—DICOM images are segmented, report sentences classified and mapped, yielding organ-level volume–text pairs.
This pipeline produced, from 140k CT series, 340k volume–text pairs for training and evaluation. The data curation system is generalizable and automates a previously labor-intensive step, with the added benefit of aligning with clinical granularity.
Model Architecture and Training Strategy
The image branch is a 3D ViT (ViT-B scale, input 192×192×32), which receives organ-cropped CT subvolumes with three channel windowings (lung, soft tissue, bone), patch size 16×16×4. The text branch is a lightweight Japanese BERT variant. Training proceeds in three sequential stages:
TotalFM’s architecture thus supports efficient high-resolution contrastive training, and can be flexibly adapted for downstream zero-shot tasks or full report generation.
Experimental Protocols
Organ-wise lesion classification is posed as a binary VQA task per organ, directly aligning volume embeddings with positive and negative report sentences. Prior models could not exploit this granularity; comparison models (CT-CLIP, Merlin) receive whole-volumes and generic prompts, with non-trivial translation required for Japanese report sentences.
Finding-wise lesion classification uses zero-shot AUROC against manually defined disease label queries, across standard datasets (J-MID, Merlin Test), with a focus on generalization to findings across diverse anatomical sites.
Report generation is measured by BLEU, ROUGE-2, BERTScore, and RadGraph-F1, with qualitative clinical appraisal by a board-certified radiologist.
Results
Organ-wise Lesion Classification
TotalFM achieved higher F1 in 83% (5/6) of organs versus CT-CLIP, and in 64% (9/14) of organs compared to Merlin. Average F1 improved by 37.5% (vs CT-CLIP) and 6.5% (vs Merlin), with all organ F1 above 0.6, except the aorta—where the performance lagged, likely owing to longitudinal shape and multi-label mapping complexities.
Finding-wise Lesion Classification
For zero-shot AUROC, TotalFM outperformed Merlin in 83% (25/30) of findings in J-MID, and generally matched or exceeded Merlin in the curated Merlin Test findings—except in cases where anatomical extension (e.g., aorta, lungs) induced information dilution via sliding window inference.
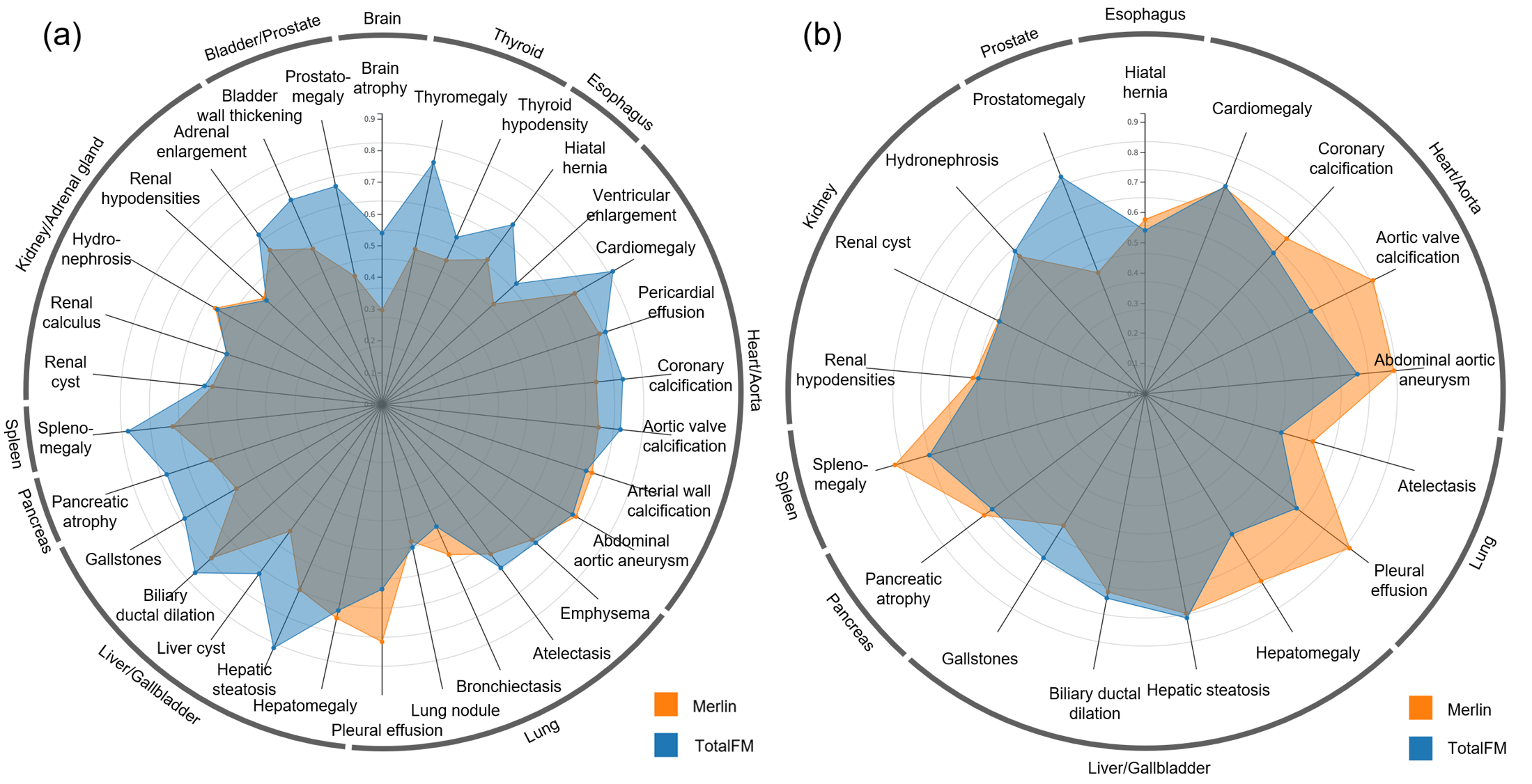
Figure 4: AUROC comparison by finding—TotalFM matches or exceeds Merlin in the majority of evaluated findings.
Vision–LLM Report Generation
TotalFM matched or marginally exceeded Merlin in aggregate BLEU, ROUGE-2, BERTScore, and RadGraph-F1 on nine organ categories. Strongest numerical improvements were observed for gallbladder and pelvic organ cohorts.
Human evaluation confirmed that auto-generated reports approached expert standards, though identified areas for further precision, especially in nuanced anatomic delineation and less common findings.
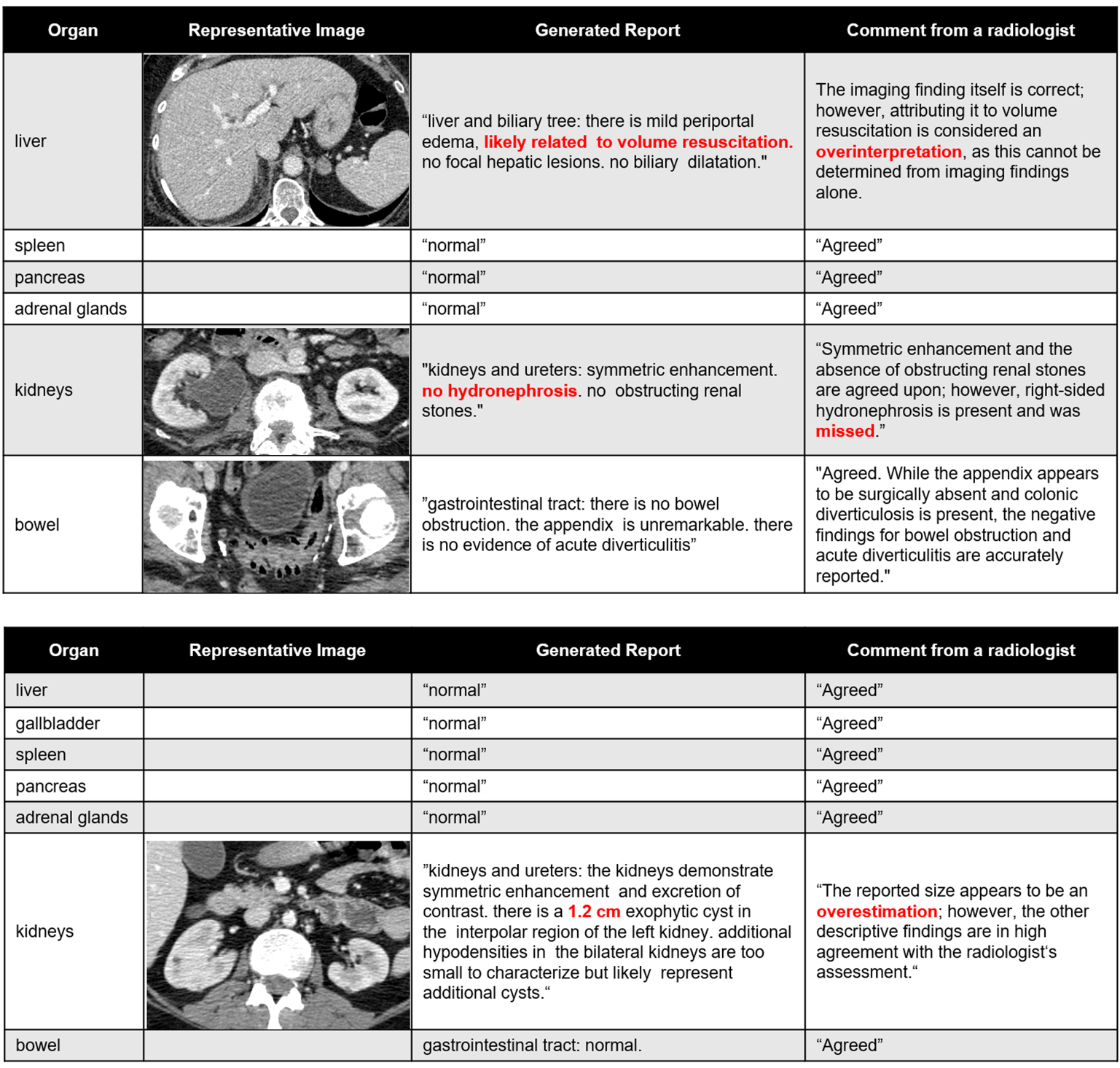
Figure 5: Examples of auto-generated organ-specific reports and expert clinician evaluation, highlighting content accuracy and areas for refinement.
Practical and Theoretical Implications
The introduction of organ-level decomposition reshapes the batching–resolution trade-off in 3D foundation models. By permitting higher batch sizes (up to 32 batch/GPU on H100 with bf16) while preserving fine anatomical detail, TotalFM enables contrastive learning at a scale and fidelity not previously attainable without excessive resource requirements. The organ-separated approach aligns with clinical reasoning granularity, potentially enabling both retrieval-based systems for structured reporting and fine-tuning for organ-specific CAD.
Organ-wise contrastive tasks, as used in TotalFM, offer a rigorous and clinically meaningful evaluation method, more robust to prompt variations than finding-wise tasks and more reflective of true report complexity. This evaluation modality could inform future benchmarking protocols.
A key limitation is that not all findings can be associated with existing segmentation vocabulary (notably, vascular syndromes and lymphatic findings are underrepresented). Further, false-negative sampling in InfoNCE at this scale can introduce noise, which may be mitigated via adaptive hard negative mining or improved supervision for ambiguous pairs. Finally, while TotalFM provides a mechanism to integrate partial organ findings, clinical practice requires comprehensive integration—multi-series and multi-organ reasoning—which may be further supported with hierarchical or graph-based reasoning modules.
Future Directions
Enhancements in text–region grounding (e.g., open-vocabulary segmentation, visual grounding [Zhao2025-tl, Ichinose2023-qe]), adaptive hard-negative construction [Kim2025-js], and multi-series integration represent immediate axes for research exploration. Interfacing TotalFM with large-vocabulary segmentation models or composite prompt-based CAD tools can extend both its accuracy and clinical utility.
Conclusion
TotalFM demonstrates that organ-separated learning is a practical and effective framework for constructing high-performance, high-efficiency 3D-CT foundation models. By leveraging modern segmentation and LLM techniques, this pipeline achieves strong zero-shot generalization and near state-of-the-art report generation, while substantially reducing computational demands. Organ-wise contrastive tasks should be considered for future clinical evaluation of 3D medical foundation models, and the TotalFM architecture constitutes a compelling template for subsequent systems in this domain.
Reference: "TotalFM: An Organ-Separated Framework for 3D-CT Vision Foundation Models" (2601.00260)