- The paper presents innovative biomolecular controllers that integrate feedback and feedforward systems to ensure precise gene regulation in reprogramming applications.
- It shows that high-gain feedback on Oct4 dynamics can double reprogramming efficiency in hiPSCs by tightly controlling cellular heterogeneity.
- The study also addresses challenges like resource sharing and context-dependence, underscoring the need for adaptable, context-aware control designs.
Control Engineering in Synthetic Biology: Principles, Architectures, and Reprogramming Applications
Introduction: Motivation and Context
Control systems engineering underpins the quantitative manipulation of genetic circuits within cells for diverse applications, including cell therapy, biomanufacturing, and cell fate reprogramming. Central to these efforts is the accurate regulation of key regulatory factor concentrations despite environmental uncertainty and cellular perturbations. This essay reviews the system-level challenges in realizing such control, examines biomolecular controller architectures implementable in vivo, and discusses their relevance through the study of cell fate reprogramming—specifically, the induction of pluripotency in somatic cells.
The Controlled System: Synthetic Genetic Devices and Cellular Context
A synthetic genetic device constitutes an input/output module embedded in cellular genomes, orchestrating the expression of a target protein via transcription and translation.
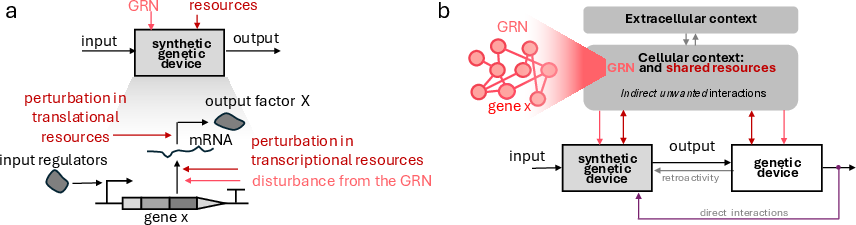
Figure 1: A synthetic genetic device as an input/output module for gene expression, highlighting typical environmental perturbations.
In mammalian and prokaryotic contexts, perturbations in transcriptional and translational resource pools, inherent gene regulatory network (GRN) crosstalk, endogenous expression, and retroactivity collectively degrade output predictability and stability. Resource-sharing effects and context-dependence now represent the predominant hurdles to modular circuit engineering.
Architectures for Biomolecular Disturbance Attenuation
Feedback and Feedforward Control: Framework and Rationale
Robust output regulation in biological circuits draws from control theory, with architectures classified into feedback and feedforward types:
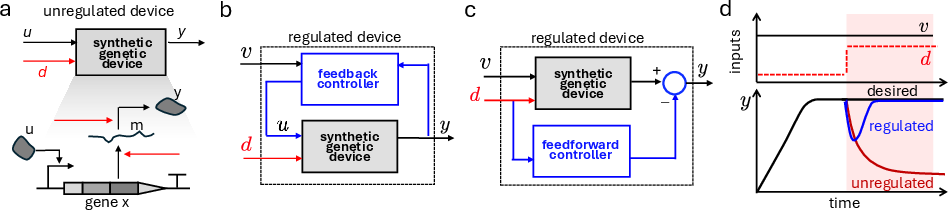
Figure 2: Biomolecular controller architectures—unregulated, feedback-regulated, and feedforward-regulated devices—and their disturbance rejection properties.
- Feedback controllers compute regulatory actions based on measurements of deviations of the output from setpoints, often using negative feedback to achieve homeostasis or adaptation.
- Feedforward controllers anticipate disturbances by acting directly on the pathways affected by a perturbation, designed to cancel disturbance effects via matched system structure.
Both classes aim for disturbance rejection and adaptation but differ in their implementation constraints and functional outcomes.
Biomolecular Feedback Control: Integral and Quasi-Integral Mechanisms
Biomolecular integral feedback, necessary for perfect adaptation, is theoretically demanding due to molecular decay (leaky integration). Architectural realizations employ fast covalent modification cycles (e.g., phosphorylation) to approximate timescale-separated integration.
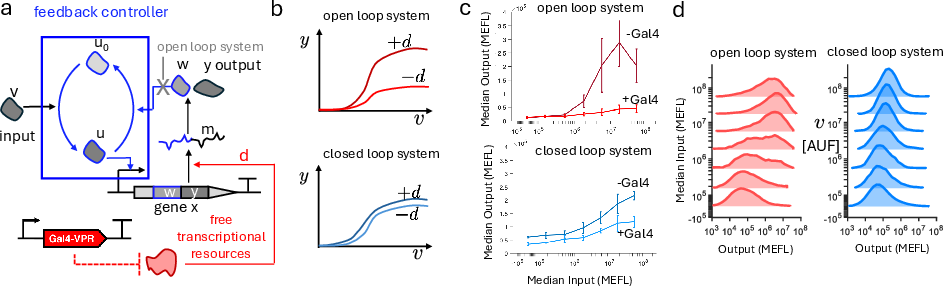
Figure 3: Biomolecular implementation of an integral feedback controller using a phosphorylation/dephosphorylation cycle and experimental validation.
Empirical implementation in mammalian cells utilizes a bacterial phosphorylation cycle orthogonal to mammalian signaling, with co-expression of output protein and a feedback phosphatase. This architecture achieves:
- Significant attenuation of output disturbance due to resource variability and targeted mRNA decay
- Reduced cell-to-cell variability in output due to copy number variation
Theoretical analysis confirms that steady-state error can be made arbitrarily small as the timescale separation parameter ε is minimized, though global convergence in nonlinear scenarios remains unresolved.
Feedforward Control Motifs: Incoherent Loops and Adaptation
Feedforward designs leverage incoherent motifs in which the disturbance simultaneously activates and inhibits the output via different paths.
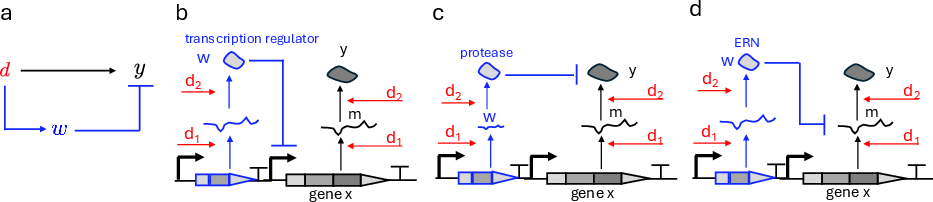
Figure 4: Incoherent feedforward controllers realized via transcriptional repression, proteolysis, or endoribonuclease-mediated mRNA degradation.
Their key properties include:
- Output adaptation to perturbations in transcription rate (DNA copy number, resource sharing) and, when implemented with protein/endonuclease effectors, to translation-level perturbations.
- The potential for perfect adaptation in the limit of high controller gain and matched disturbance pathways
- For post-transcriptional feedforward controllers using microRNAs, adaptation is limited to transcriptional disturbances if promoter contexts are matched.
Experimental validation demonstrates that feedforward-regulated modules maintain constant output under resource depletion, suppress output variability across cell lines, and attenuate sensitivity to DNA copy number changes.
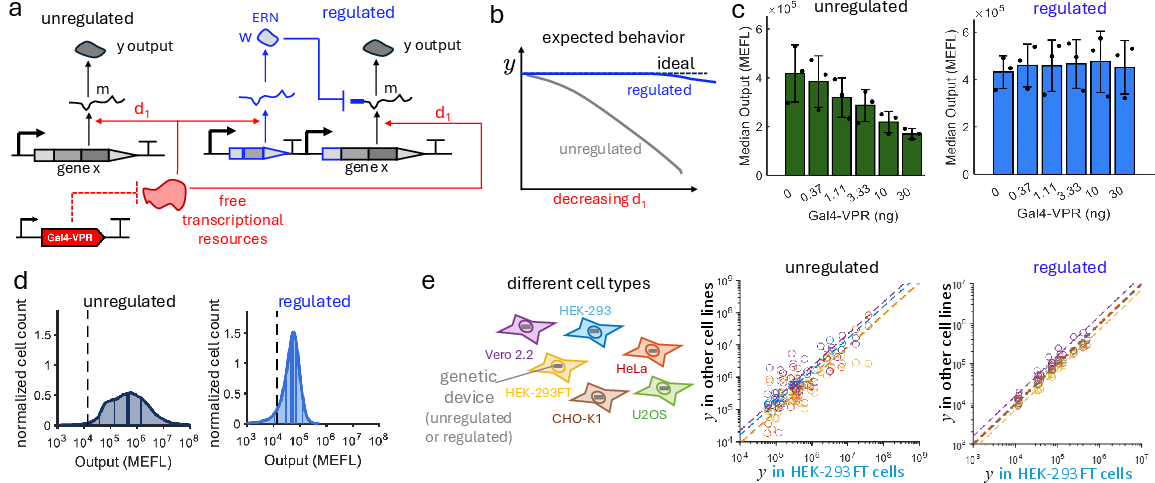
Figure 5: Implementation and performance of a feedforward controller in mammalian cells, demonstrating robust output despite resource perturbation.
Case Study: High-Precision Control of Cell Fate via Reprogramming
Human induced pluripotent stem cells (hiPSCs) are generated by enforcing expression of pluripotency master regulators (Oct4, Sox2, Nanog), but efficiency and fidelity depend critically on the controlled level of these factors due to the multistable nature of the underlying GRNs.
Figure 6: Medical relevance of hiPSCs for regenerative medicine and therapy.
Mechanistic modeling indicates that constant overexpression is inadequate for targeting desired attractors of the pluripotency GRN, while high-gain feedback on Oct4 trajectories ensures convergence to appropriate levels.
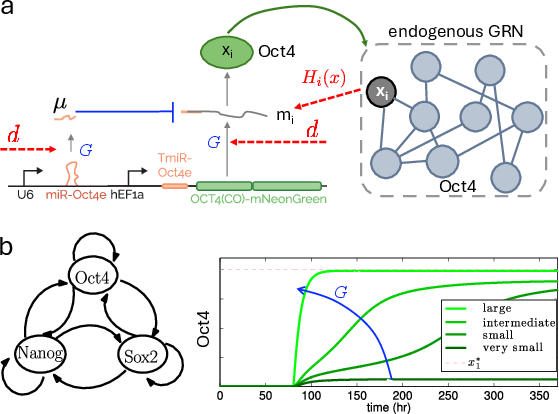
Figure 7: Synthetic genetic implementation of high-gain feedback and feedforward control for Oct4; ODE simulations validate disturbance attenuation properties.
Empirical studies combining high-gain feedback with feedforward architecture demonstrate:
- A narrow distribution of Oct4 levels among successfully reprogrammed cell populations, aligning with desired reprogramming trajectories
- Superior reprogramming efficiency—variant v4 of the controller achieves a two-fold improvement over traditional overexpression protocols
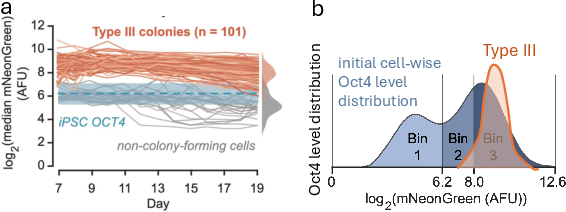
Figure 8: Single-cell Oct4 trajectories reveal the narrow desired range required for successful iPSC reprogramming.
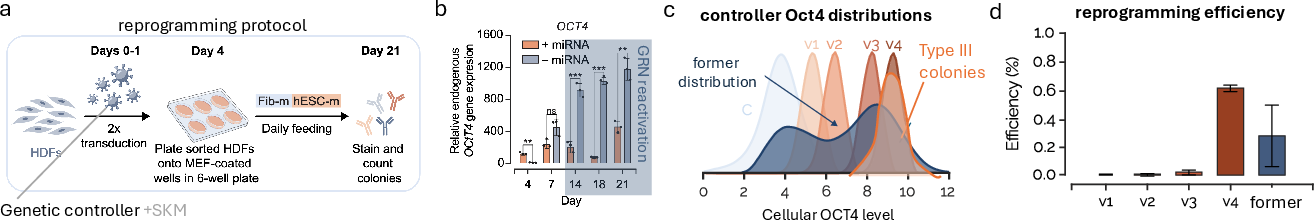
Figure 9: Experimental outcomes: controller variants yield sharply defined Oct4 distributions and robust, enhanced reprogramming efficiency.
Nevertheless, proliferation-dependent dilution leads to selection against high Oct4-expressing cells over time, limiting theoretical controller performance. This indicates the necessity for linking cell survival/growth modulation to target expression levels to maintain the desired population distribution.
Implications and Future Directions
This research demonstrates that theoretical principles of control theory are realizable within the constraints of biochemistry, providing robust solutions to stochasticity, environmental variability, and network crosstalk in synthetic and endogenous circuits. Key implications include:
- Standardization and Modular Design: Next-generation architectures should anticipate context effects not only from endogenous GRN coupling but also dynamic resource allocations and growth feedback.
- Field Translation: Most controllers have not been validated outside laboratory strains; operational robustness under in vivo and field conditions remains a significant challenge.
- Integration with Learning-Based Design: Hybrid approaches that combine mechanistic modeling with data-driven learning can enhance context awareness and inform controller tuning in poorly characterized environments.
Conclusion
Biomolecular control system engineering is essential for robust synthetic biology, especially when tight quantitative regulation is required for applications such as cellular reprogramming, production, and therapy. This work establishes a rigorous foundation for the implementation and analysis of both feedback and feedforward control in genetic circuits, with direct translation to notably improved outcomes in applied systems such as hiPSC reprogramming. Key open challenges include controlling coupled growth rate effects, extending validation to complex environments, and developing controllers with integrated context-awareness—potentially via machine learning—ushering in a new era of modular, predictable, and adaptive design in synthetic biology (2601.20135).








