- The paper presents a rigorous classification of generative AI models for enzyme design, distinguishing sequence-generating from backbone-generating approaches.
- It demonstrates significant improvements in catalytic activity, stability, and substrate specificity, with enhancements reaching up to 95-fold in specific enzyme designs.
- The study highlights current limitations and future directions, advocating for integrated experimental feedback and advanced steering techniques to overcome design challenges.
Generative AI Paradigms for Enzyme Design and Biocatalysis
Overview of Generative Model Classes for Enzyme Design
This work provides a rigorous classification of generative AI models leveraged for enzyme design, organizing them along two main axes: sequence-generating and backbone-generating modalities. Sequence-generating models encompass substitution models for improving existing enzyme scaffolds by point mutations, family expansion models producing divergent variants within enzyme families, and structure-conditioned models generating sequences for specified protein backbones. Backbone-generating models, based on denoising or diffusion frameworks, construct novel protein architectures from random noise, with varying levels of conditional granularity applied for precise active site scaffolding.
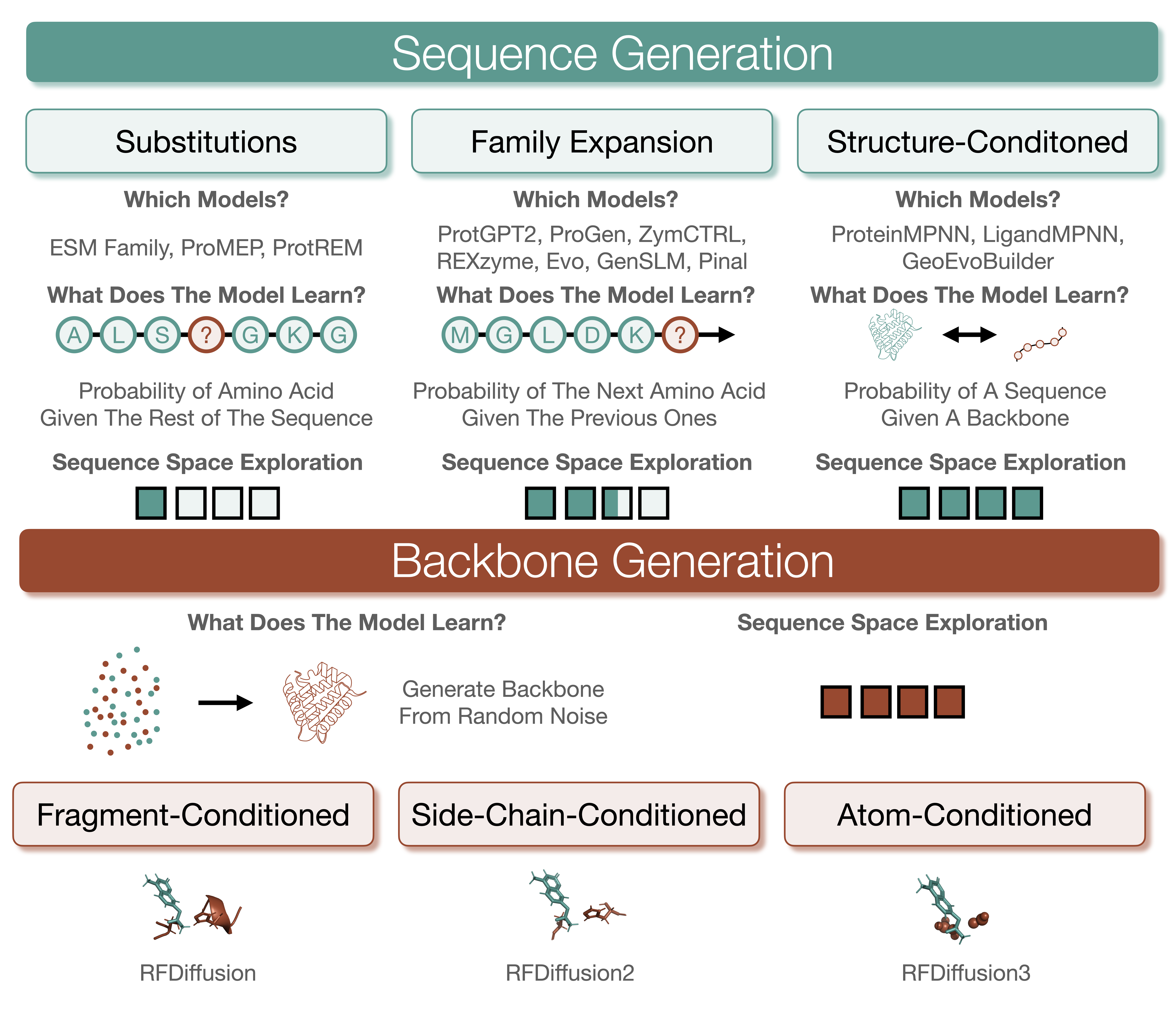
Figure 1: Classes of generative AI models for enzyme design, delineating models by sampled modality and granularity of conditioning.
The paper identifies notable advances in self-supervised learning, such as the efficacy of masked LLMs (MLMs) and causal LLMs (CLMs), paired with Transformer architectures, in modeling high-dimensional distributions of protein sequences and structures. Each model type demonstrates different strengths regarding the scope of design, from local mutational optimization to the generation of structurally novel enzymes. The authors highlight that generalist models—trained across diverse protein families—are particularly valuable for non-expert users due to broad applicability without family-specific retraining.
Experimental Validation and Quantitative Outcomes
A major emphasis is placed on reporting experimentally validated enzyme designs produced via generative AI, including documentation of strong numerical gains in catalytic efficiencies, stability, substrate scope, and other relevant biocatalytic metrics. The work summarizes benchmarked examples across numerous enzyme classes, with activities spanning from moderate improvements to several orders of magnitude enhancement relative to natural or wild-type sequences.

Figure 2: Representative structures of best-performing enzymes designed using generative AI models, illustrating diversity across classes and folds.
- Notable, validated improvements include:
- TEV protease redesigned using ProteinMPNN exhibiting a 26-fold increase in catalytic activity and a 40°C increase in melting temperature.
- Stabilization and subsequent activity optimization of α-ketoglutarate-dependent oxygenase, achieving an 80-fold increase via combinatorial application of ProteinMPNN and directed evolution.
- De novo serine hydrolase and retro-aldolase designs with backbone-generating models (RFDiffusion family), reaching median activities of natural enzymes.
- LigandMPNN-mediated modification of substrate specificity and enantioselectivity in de novo bundle scaffolds, with up to 95-fold improvements.
The authors also provide a comprehensive tabulation of engineered enzymes, their associated generative pipelines, sequence identity statistics, quantitative improvements, and referenced validation studies. These results robustly demonstrate the current state of generative AI for empirical enzyme engineering, transcending prior low-success paradigms.
Architectural Innovations and Design Workflows
The paper details operational workflow distinctions among model classes:
- Masked LLMs (MLMs): Used for scoring and prioritizing substitutions in natural scaffolds. Integration of experimental feedback greatly improves their design utility, as seen in iterative rounds of mutagenesis guided by measured activities.
- Family Expansion Models (CLMs): Capable of generating novel, low-identity sequences within enzyme families, recapitulating properties present in their training data, but limited in exploring properties that are not evolutionarily represented.
- Structure-Conditioned Models (ProteinMPNN, LigandMPNN): Direct generation of entire sequences for user-defined backbones or ligand-binding contexts. These have shown marked success in stability optimization, facilitating the subsequent introduction of activity-enhancing but destabilizing mutations.
- Backbone-Generating Models (RFDiffusion Family, Proteus2): Denoising/diffusion networks that construct backbones with tailored active sites. The latest iterations have circumvented manual fragment libraries and enable direct conditioning on catalytic residue coordinates.
Model selection is advised to be property- and protein-family specific, necessitating empirical benchmarking (e.g., ProteinGym) and consideration of available context (structural data, MSAs).
Technical Limitations and Research Challenges
Several conceptual and practical limitations are critically analyzed:
- Training data implicitly encode evolutionary fitness rather than industrially desirable properties, constraining sequence-function mapping and chemical diversity.
- Most current models neglect second-shell interactions and overall fold dynamics, which dominate real-world catalytic performance.
- No genuinely end-to-end generative solution exists capable of autonomously deducing chemical requirements for new-to-nature reactions—a significant bottleneck is manual chemical hypothesis construction during backbone design and active site placement.
- Scaling of model parameters has shown diminishing returns in real design tasks, underscoring the necessity of integrated experimental feedback and explainable AI for future development.
Advancement in steering techniques (e.g., reinforcement learning, DPO, hybrid physicochemical models) and incorporation of explicit catalytic priors are highlighted as future technical imperatives.
Implications and Prospects for AI-Driven Biocatalysis
The maturity of generative AI models for practical enzyme engineering now supports their adoption in industrial biocatalysis, especially in scenarios where conventional directed evolution and rational design methods are inefficient. The integration of automated experimental pipelines and active learning paradigms can accelerate the design-build-test-learn cycle, enabling the rapid creation and optimization of biocatalysts for diverse chemical transformations.
From a theoretical perspective, the observed performance ceilings invite deeper investigation into sequence-function causality, diversification of model architectures beyond Transformer scaling, and novel training objectives that transcend evolutionary datasets. Future models will likely blend generative distributions with explicit physicochemical constraints, potentially enabling fully autonomous and interpretable enzyme design for bespoke reactions.
Conclusion
This work provides an authoritative overview of the landscape and capabilities of generative AI models in enzyme design and biocatalysis. Through extensive classification, empirical analysis, and critical assessment of model performance, it evidences both the transformative potential and current bottlenecks facing AI-driven protein engineering. The field’s advancement will hinge on continued methodological hybridization, rigorous benchmarking in task-relevant contexts, and integration with experimental data streams to fulfill the promise of programmable catalysis (2602.03779).

