- The paper introduces LuMamba, a unified EEG model that uses topology-invariant latent encoding and bidirectional Mamba blocks for efficient long-sequence analysis.
- It demonstrates a hybrid pre-training strategy combining masked reconstruction and LeJEPA, achieving over 20% AUPR improvement in Alzheimer’s detection.
- The architecture scales linearly with sequence length, requiring 26× fewer FLOPS than LUNA and enabling robust cross-montage generalization with only 4.6M parameters.
LuMamba: Latent Unified Mamba for Electrode Topology-Invariant and Efficient EEG Modeling
Introduction
Electroencephalography (EEG) modeling faces persistent scalability and generalization challenges stemming from both the quadratic complexity of Transformer-based architectures and the pronounced variability in electrode topologies across datasets. "LuMamba: Latent Unified Mamba for Electrode Topology-Invariant and Efficient EEG Modeling" (2603.19100) addresses these with a unified self-supervised framework that (1) combines topology-invariant latent encodings, as pioneered by LUNA with cross-attention, and (2) leverages the temporal and computational efficiency of state-space models (SSMs) via FEMBA’s bidirectional Mamba blocks. LuMamba is further the first to systematically investigate the adaptation of Latent-Euclidean Joint-Embedding Predictive Architecture (LeJEPA) for biosignal time series, probing the interplay between isotropic latent regularization and masked signal reconstruction.
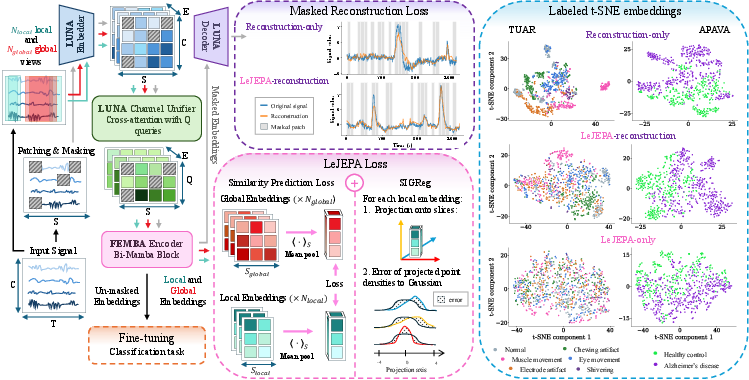
Figure 1: Overview of the LuMamba architecture, comprising topology-invariant cross-attention encoding and bidirectional Mamba temporal modeling. t-SNE projections reveal the latent structure under different pre-training objectives.
Methodology
Architecture
LuMamba’s core innovation lies in its fused architecture which achieves topology-invariant processing and efficient long-sequence modeling with only 4.6M parameters. The input EEG tensor x∈RB×C×T is first tokenized, projecting temporally patchified and spectrally encoded signals (augmented with 3D positional embeddings) into a token space. LUNA’s channel-unification module applies a learned-query cross-attention to yield montage-agnostic latent representations, effectively resolving the heterogeneity of varying electrode configurations. The output is processed through two bidirectional Mamba (bi-Mamba) blocks, yielding linear-time complexity for temporal modeling, essential for scaling to long EEG sequences where access patterns are non-stationary and context-rich. The architecture is completed by a cross-attention decoder (for reconstruction during pre-training) or a lightweight Mamba-based classifier for downstream supervised fine-tuning.
Pre-training Strategies
LuMamba is evaluated under three self-supervised objectives: (1) masked reconstruction, (2) LeJEPA-only, and (3) a hybrid LeJEPA+reconstruction loss. The masked reconstruction objective follows prior work—randomly masking 60% of input patches and reconstructing them from context. LeJEPA involves regularizing the latent embedding distributions toward an isotropic Gaussian by combining Sketched Isotropic Gaussian Regularization (SIGReg) with a joint-embedding alignment between global and local temporal views. The mixed objective leverages complementary properties: reconstruction enforces local structure and cluster compactness, whereas LeJEPA induces isotropy for enhanced out-of-distribution generalization.
Empirical Results
Latent Space Analysis
t-SNE projections of LuMamba’s embeddings (Figure 1) reveal that reconstruction-only objectives yield notably structured and clustered latent spaces, while LeJEPA introduces isotropy and dispersion. The hybrid (LeJEPA+reconstruction) objective achieves a trade-off—preserving sufficient structure for robust classification but with increased distributional smoothness, empirically associated with superior cross-montage generalization performance.
LuMamba is pretrained on 21,600 hours of unlabeled EEG from the TUEG corpus and evaluated across five benchmarks: TUAB (normal/abnormal detection), TUAR (artifact recognition), TUSL (event classification), and two disease-specific datasets with distinct montages—APAVA (Alzheimer’s, 16 channels) and TDBrain (Parkinson's, 26 channels). Noteworthy observations:
- On TUAB, LuMamba attains 80.99% balanced accuracy and an AUPR of 0.892—on par with the best reported self-supervised approaches and trailing only transformer-based LaBraM/LUNA models, despite a substantially lower parameter count.
- For APAVA (Alzheimer’s detection), the hybrid objective achieves 0.970 AUPR— an improvement of over 20% compared to reconstruction-only pre-training, indicating strong benefits for distributional regularization under unseen electrode topologies.
- On TDBrain (Parkinson’s), LuMamba maintains competitive performance (0.96 AUROC), generalizing well even with 10 more channels than in pre-training.
Quantitative results also demonstrate that for in-distribution tasks (same-channel), reconstruction-only achieves marginally higher scores, underscoring that strong cluster structure is helpful for i.i.d. settings, but detrimental for out-of-distribution (electrode shift) scenarios.
Efficiency and Scaling
LuMamba’s computational efficiency is underscored by its linear scaling with sequence length due to the Mamba backbone. Comparative FLOPS analysis shows:
(Figure 2)
Figure 2: FLOPS vs. sequence length for state-of-the-art EEG foundation models; LuMamba’s linear scaling enables substantially longer sequence processing before out-of-memory barriers.
- LuMamba requires 26× fewer FLOPS than LUNA and 377× fewer than LaBraM at matched sequence lengths, and can process 12× longer sequences within standard GPU memory than leading transformer-based baselines.
- This represents a critical improvement for real-world EEG applications, which often operate with long, continuous sequences unsuitable for attention-based models.
Implications and Future Perspectives
LuMamba represents a significant advance for EEG foundation models in heterogeneous real-world settings. Practically, its architecture unlocks:
- Montage-invariant deployment: Direct transfer across datasets/institutions without retraining or data discard due to montage mismatch.
- Low-cost long-horizon modeling: Efficient analysis of clinical and experimental EEG without severe window length restriction.
- Improved generalization: The demonstrated robustness under cross-montage transfer suggests utility for large-scale federated or multi-site studies.
The introduced adaptation of LeJEPA for biosignal time series suggests that self-supervised objectives which promote isotropy and regularization can systematically enhance model robustness to hardware and protocol differences, a primary barrier for EEG model translation. Theoretically, the findings motivate further study of the trade-off between local latent structure and global smoothness in SSL objectives for temporal and biomedical signals.
Future directions should include scaling up pre-training corpora and electrode diversity, extending the range of downstream evaluations (including regression and multi-label tasks), and investigating modularity for simultaneous multi-montage inference.
Conclusion
LuMamba demonstrates that high-performance, computationally efficient EEG modeling across diverse electrode configurations is attainable through a principled fusion of topology-invariant latent encoding and SSM-based temporal blocks. The hybridization of masked reconstruction and LeJEPA objectives yields state-of-the-art cross-montage generalization, especially notable under severe latent topology shifts, without reliance on the computational cost of full attention. LuMamba thus establishes a robust new baseline for foundation models in large-scale and heterogeneous EEG analysis.
