- The paper presents a novel 3D self-supervised learning framework employing superpatch partitioning and dual-masking to preserve vital volumetric context.
- It achieves robust reconstruction on CT datasets with PSNR of 30.28, SSIM of 0.98, and LPIPS of 0.26, outperforming AE, VAE, and VQ-VAE baselines.
- The study demonstrates practical implications for segmentation and lesion detection while highlighting opportunities for reducing fixed-pattern artifacts.
MAESIL: Masked Autoencoder for Enhanced Self-supervised Medical Image Learning
Problem Context and Motivation
Self-supervised learning (SSL) is increasingly critical in 3D medical image analysis due to the scarcity and expense of labeled data. CT imaging, with its inherently volumetric structural information, presents unique challenges: most prior SSL approaches simplify 3D data into independent 2D slices, causing a loss of axial coherence and critical spatial relationships necessary for robust downstream tasks. Existing frameworks such as MedMAE address cross-domain pretraining issues but do not adequately preserve the structural context of 3D inputs. The limitations of 2D-based approaches and the need for effective, scalable 3D SSL forms the main motivation for the MAESIL framework proposed by the authors.
Methodological Framework of MAESIL
MAESIL (Masked Autoencoder for Enhanced Self-supervised Medical Image Learning) directly tackles the problem of 3D context loss via a tailored encoder-decoder Transformer architecture operating on a custom-designed "superpatch" input unit. The processing pipeline is designed for a balance between spatial context preservation and computational efficiency:
After taking a volumetric CT scan, MAESIL partitions the input into non-overlapping superpatches (128×128×128 voxels). Each superpatch is densely subdivided into standardized patches (8×8×8), which are embedded via a 3D convolution and positional encoding to populate the Transformer input sequence.
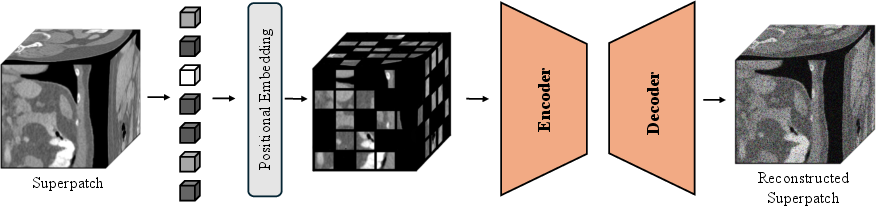
Figure 1: Overview of the MAESIL framework, illustrating masked 3D superpatch encoding and Transformer-based reconstruction.
The masking paradigm extends 2D MAE concepts to the volumetric case with a dual-masking strategy. Specifically, each superpatch undergoes:
- Plane-wise masking: randomly masking 75% of the patches within each 2D plane.
- Axis-wise masking: dropping a contiguous block of 50% of slices along the S-I axis.
This combination imposes high information loss, ensuring that the model must capture long-range 3D dependencies rather than overfitting to local patterns.
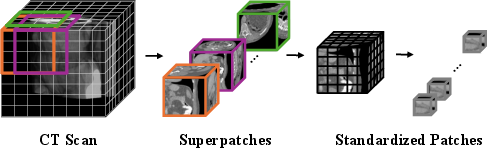
Figure 2: The 3D input processing pipeline—CT scans are partitioned into superpatches then densely tokenized, forming the representation for the Transformer.
The encoder processes the unmasked tokens into latent embeddings, which are then reassembled in the decoder with the addition of learnable mask tokens inserted at missing locations. The sequence is restored to its canonical ordering and passed through Transformer blocks for reconstructive generation.
Quantitative and Qualitative Results
The efficacy of MAESIL is empirically validated on three heterogenous, large-scale medical CT datasets: BTCV (abdominal), LIDC-IDRI (thoracic), and TotalSegmentatorV2 (whole-body, multi-structure). All models, including strong baselines (AE, VAE, VQ-VAE), are trained uniformly on the combined dataset for robustness to anatomical and technical variability.
MAESIL demonstrates strong numerical gains in all key reconstruction quality metrics:
- PSNR: 30.28
- SSIM: 0.98
- LPIPS: 0.26
compared with AE (25.23, 0.97, 0.39), VQ-VAE (20.04, 0.88, 0.49), and VAE (11.91, 0.25, 0.91). Notably, the VAE baseline exhibits a catastrophic drop in SSIM, underscoring the critical importance of 3D context modeling.
Within-dataset ablations reveal significant performance variance (e.g., PSNR 32.09 on BTCV vs. 18.72 on TotalSegmentatorV2), reflecting the challenge of multi-structure, full-body scans. Importantly, all ablations utilize a 75% mask ratio with 768-dimensional embeddings, indicating robust representation learning under aggressive information suppression.
The accompanying qualitative results present high-fidelity reconstructions in axial, coronal, and sagittal views, with clear preservation of core anatomic features and boundaries.
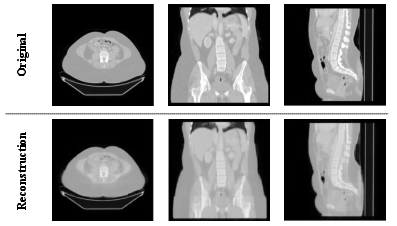
Figure 3: Qualitative reconstruction results across anatomical views; the outputs retain fine-grained structure and 3D relationships.
However, fixed-pattern artifacts, a known challenge in MAE-based decoders, are evident and constitute a target for subsequent architectural refinement.
Implications and Future Directions
MAESIL substantiates the importance of explicitly leveraging volumetric structure in medical image SSL. The superpatch design improves context retention across scales, while dual-masking prevents shortcutting by enforcing information synergy. These findings have immediate practical implications for generalizable pretraining backbones in segmentation, lesion detection, and disease classification tasks—domains where high efficiency per label is vital.
On the theoretical side, MAESIL demonstrates that masking paradigms can be extended to multi-axis, highly-structured medical domains without degrading model capacity or scalability. The observed artifact patterns and datasets' inherent noise suggest room for work in robust context restoration and dynamic masking schedules.
Future research will likely focus on:
- Masking schedule optimization and curriculum.
- Decoder redesigns for artifact suppression.
- Downstream task transferability, such as cross-modality and cross-dataset generalization.
- Application to other high-dimensional bioimaging modalities.
Conclusion
MAESIL presents a principled approach to SSL in 3D medical imaging, with a novel superpatch masking strategy yielding state-of-the-art quantitative and qualitative results relative to established generative baselines. The framework's data efficiency and robustness highlight its potential as a foundation for transferable, annotation-efficient representation learning in clinical imaging pipelines. Further research is warranted to refine artifact handling and broaden downstream applicability (2604.00514).


