- The paper introduces chaos-controlled reservoir computing to stabilize live neuron dynamics and significantly boost pattern recognition accuracy.
- It employs pre-flight dynamical diagnostics and optical modulation to mitigate biological variability and extend substrate longevity.
- Knowledge Transplant geometrically aligns latent state spaces across cultures, reducing training time and enhancing cross-sample model performance.
Chaos-Controlled Computing with Living Neurons: A Detailed Synthesis
Introduction
The manuscript "Computing with Living Neurons: Chaos-Controlled Reservoir Computing with Knowledge Transplant" (2604.02552) delineates a systematic approach to exploiting in vitro neural cultures as dynamic computational substrates under the reservoir computing (RC) paradigm. The work addresses key obstacles—biological variability, fragility, and limited lifespan—by integrating pre-experimental dynamical diagnostics, optical chaos control for stabilization, and a geometric procedure for cross-sample model transfer (Knowledge Transplant). The empirical investigation leverages optogenetically actuated mouse embryonic stem cell-derived motoneuron cultures interfaced with high-density microelectrode arrays (MEAs) and custom electrophysiological platforms. The multi-pronged methodology achieves robust real-time pattern classification with significant enhancements in accuracy and operational longevity, while Knowledge Transplant (KT) generalizes trained models across substrate instances, paving the way for cumulative and cross-generational knowledge in biological compute systems.
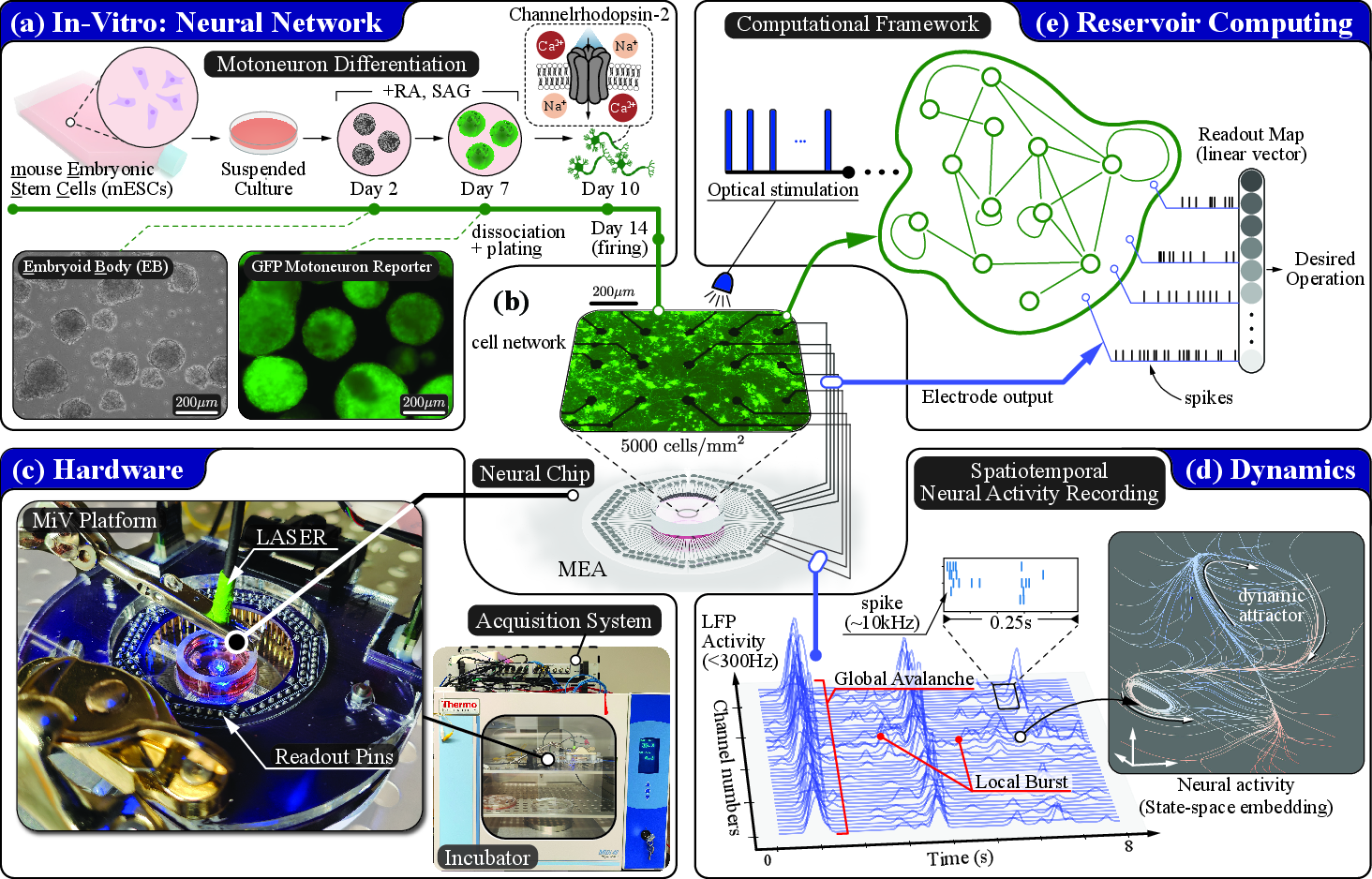
Figure 1: Overview of cc-RC experimental architecture, highlighting cell preparation, MEA integration, multimodal interfacing, and statespace embedding for dynamical diagnostics and computational readout.
Pre-flight Substrate Characterization: Metrics and Impact
Neural cultures present substantial heterogeneity even under identical protocols. The authors systematically classified over 500 cultures using dynamical system metrics: branching ratio (criticality), kernel rank (representation diversity), spectral radius (short-term memory), and transfer entropy (functional connectivity). Cultures were grouped (Types A–D) by synchronization and bursting profiles, with Types C and D exhibiting attractive balances of criticality and temporal alignment with input timing, thus offering maximal computational substrate fitness. This pre-selection predicts downstream RC performance, with Type C cultures achieving peak classification accuracies (∼80%) and best-in-class samples nearing 95% over 1–2 hours.
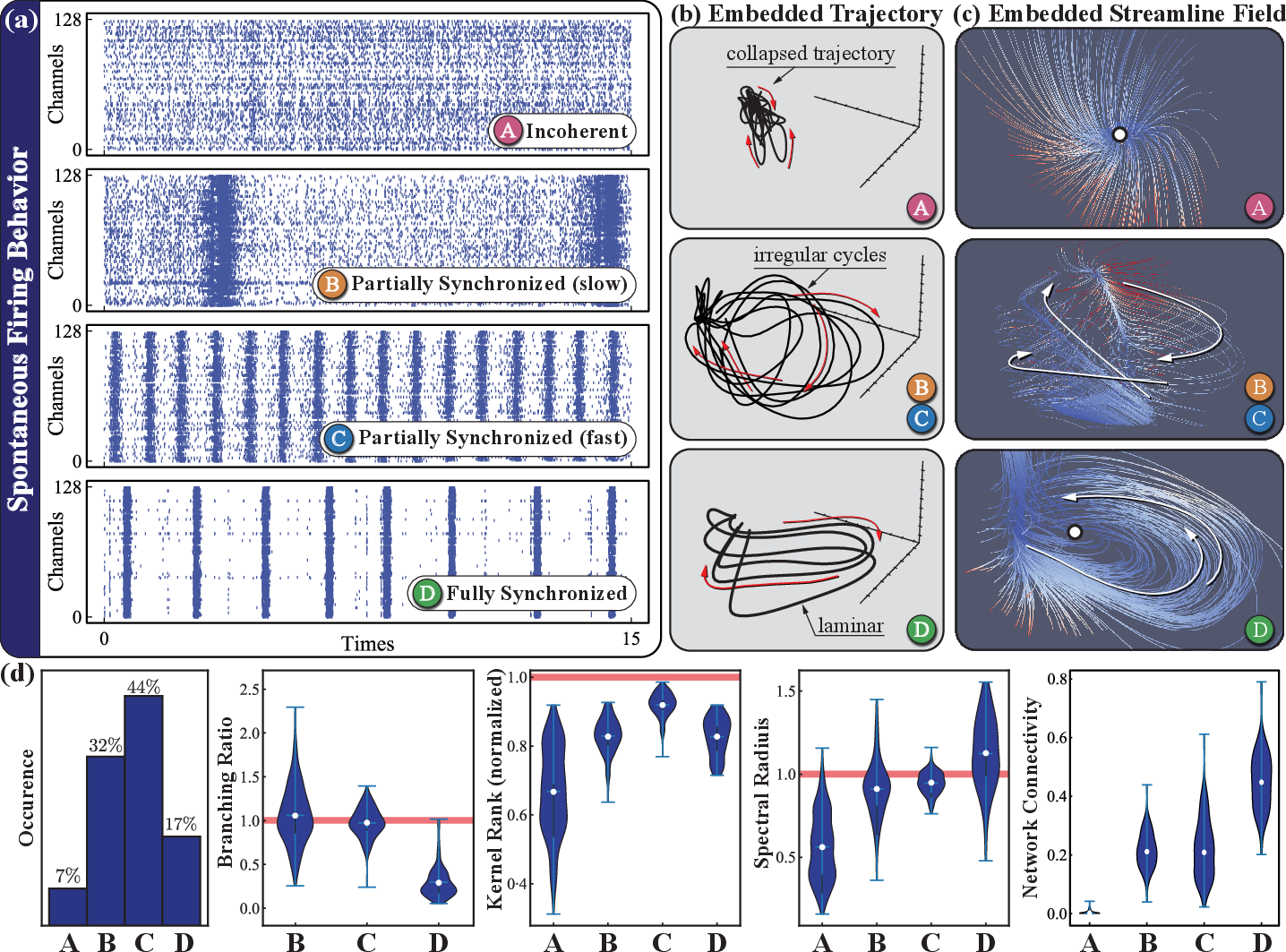
Figure 2: Spontaneous activity types, latent state trajectories, and quantitative diagnostics, illustrating diversity and computational suitability across a large ensemble of cultures.
Neural trajectory analysis via Gaussian-process factor analysis (GPFA) yields robust, low-dimensional population dynamics embeddings, providing an operational framework for both state monitoring and substrate alignment essential for later KT procedures.
Baseline and Chaos-Controlled Reservoir Computing
Initial RC implementations demonstrate that native biological dynamics, even in optimized culture types, exhibit temporal drifts leading to performance collapse beyond several hours. This accuracy decay arises from evolving state-space attractors, with trajectory separability degrading under fixed readouts.
The core innovation, chaos-controlled RC (cc-RC), introduces low-amplitude periodic optical background modulation phase-locked to input frequency. This intervention regularizes intrinsic burst timing without overriding evoked responses, achieving sustained attractor stability and enhanced separability in latent space.
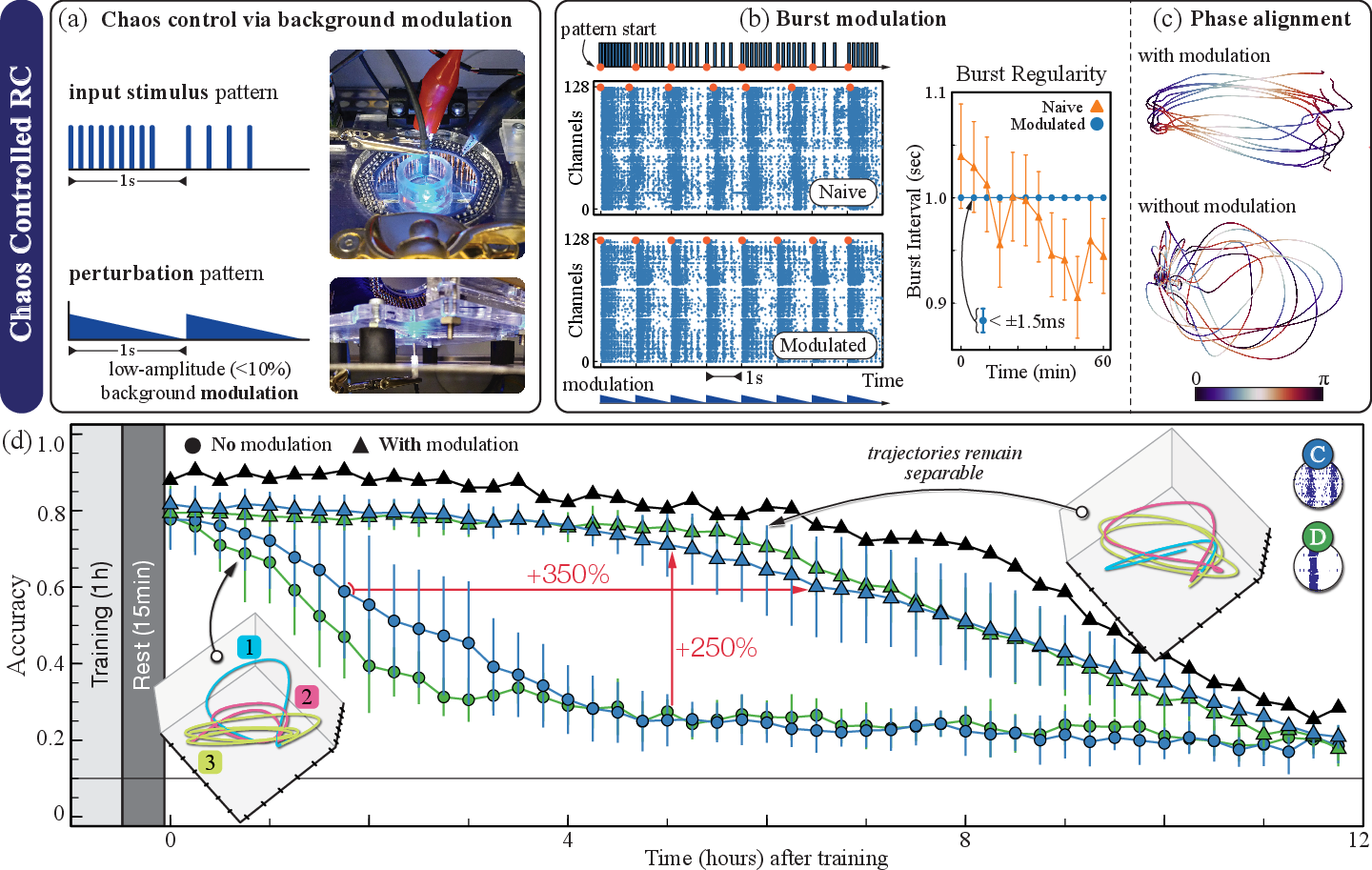
Figure 3: Effects of chaos-control modulation on burst synchrony, latent trajectory phase alignment, and task accuracy, demonstrating marked gains in stability and performance persistence.
Numerically, cc-RC extends model usability >3.5-fold above 60% accuracy thresholds, with 2.5× higher accuracy at 5 hours post-training relative to unregulated RC. The biological upper limit remains governed by the inevitable substrate degradation, but algorithmic interventions deliver substantial practical gains.
Knowledge Transplant: Geometric Alignment for Cross-Substrate Model Sharing
KT addresses the RC paradigm's inability to retain or generalize learned representations across individual, mortal neural substrates. Leveraging pre-flight similarity via dynamical classification and chaos-control-induced attractor stability, the method geometrically aligns the evoked latent state-spaces of an expert (fully trained) and a new student culture of matching type. The essential RC readout map is then transformed via ridge-regularized regression, enabling the transplanted model to operate immediately on the new culture.
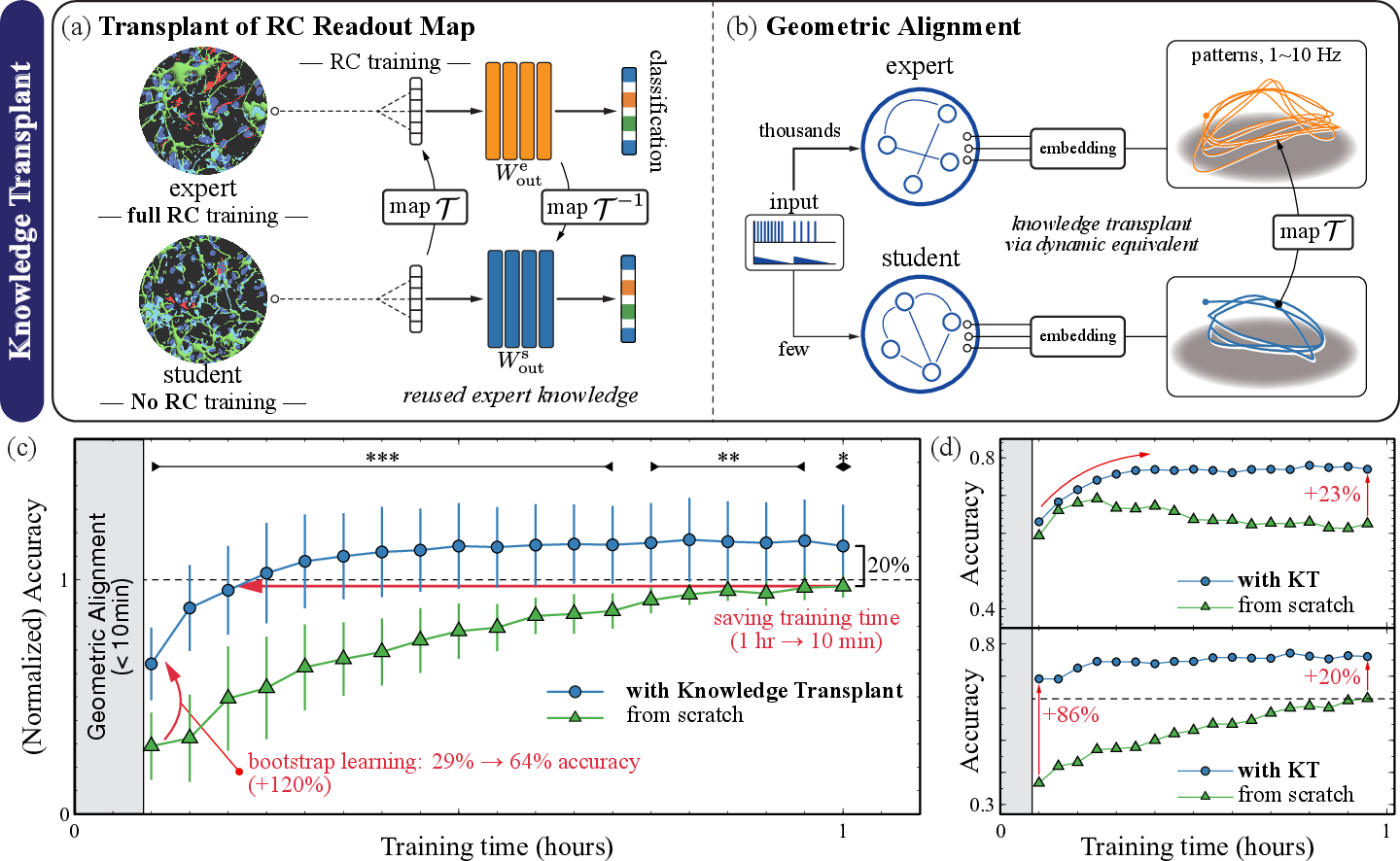
Figure 4: Schematic and empirical outcomes of Knowledge Transplant—mapping high-dimensional attractors, transforming readout mappings, and benchmarking transfer learning curves against from-scratch baselines.
Empirically, KT delivers up to 80% reduction in student training time and persistent accuracy gains (20–90%) beyond from-scratch models, including drift resistance and higher final performance ceilings. Unlike conventional actor-critic transfer or weight initialization in artificial neural networks, KT exploits substrate-independent attractor geometry, rendering the approach robust to biological variability and compatible with further online adaptation.
Implications and Future Directions
This work has several profound implications. The demonstration that biological substrate variability and mortality can be mitigated through dynamical diagnostics and optical control reframes in vitro neural cultures as reusable, shareable computational nodes. The KT approach, in particular, suggests frameworks where models and "memories" can be accumulated, shared, and refined across generations of living neural devices and potentially across different hardware substrates (neuromorphic, simulated, or hybrid).
Practically, this enables persistent biohybrid robotics, neuromorphic prosthetics, or closed-loop experimental neuroscience platforms where information is not lost with culture turnover. Theoretically, the findings reinforce the RC universality hypothesis—computational function is an emergent property of sufficiently rich dynamical systems, not their microscopic substrate—which opens the possibility of unifying artificial and biological computing models by focusing on shared attractor geometries and embedding transformations.
Future research trajectories include: scaling task complexity, investigating closed-loop or reinforcement paradigms, exploring attractor alignment across more heterogeneous systems, and dissecting the role of online synaptic plasticity in alignment and adaptation.
Conclusion
"Computing with Living Neurons: Chaos-Controlled Reservoir Computing with Knowledge Transplant" establishes a robust, empirically validated framework for living neural computation. By systematically mitigating biological variability and leveraging attractor alignment for model portability, the study lays a foundation for scalable, persistent, and transferable learning in biohybrid systems, with broad ramifications for the synthesis of artificial and living intelligence.





