- The paper demonstrates that polymer-assisted condensation (PAC) enables local enrichment of Nus proteins at rrn operons, nucleating condensate formation.
- Monte Carlo simulations with harmonic and Lennard-Jones potentials quantitatively match experimental diffusion and residence time data of Nus proteins.
- Dynamic modeling incorporating active noise reconciles the transient colocalization observed in fluorescence imaging with the weak signals in Hi-C maps.
Mechanistic Modeling of RNA Polymerase and Chromosome Spatial Organization in E. coli
Introduction and Context
This work presents an in-depth mechanistic model for the spatial co-organization of RNA polymerase (RNAP), its associated anti-termination factor NusA (hereafter Nus), and the chromosome in Escherichia coli. Ribosomal RNA operons (rrn operons) are highly transcribed loci and their spatial clustering has been observed experimentally. Notably, RNAPs form dense biomolecular condensates at rrn operons, proposed to be driven by liquid–liquid phase separation (LLPS) mechanisms, with Nus playing a central role. However, disparate results from fluorescence imaging (suggesting operon colocalization) and Hi-C experiments (showing lack of stable contacts) have posed unresolved questions regarding the underlying physical dynamics. This study develops a simulation-based, physical model to probe these phenomena and reconcile apparently contradictory observations.
The chromosome is represented as a coarse-grained flexible ring polymer with 460 monomers (each corresponding to a topologically distinct 10 kbp DNA domain) confined within a cylindrical geometry emulating E. coli cellular dimensions. rrn operons are mapped onto specific monomers according to genomic location.
Nus proteins are modeled in two states: bound (attached to RNAPs occupying rrn operons, forming a corona around operon sites) and free (diffusing in the cytoplasm). RNAPs are modeled as linkers tethering Nus proteins to rrn operon sites on the chromosome, and are present in quantities consistent with experimental cell data for slow growth conditions.
Protein–protein and protein–polymer interactions are governed by harmonic and Lennard-Jones (LJ) potentials. Critically, free Nus–Nus and free–bound Nus interactions include attractive LJ terms, facilitating potential condensate formation. Monte Carlo dynamics govern the evolution, with time scales mapped to experimental diffusion constants.
The choice of model parameters (protein concentrations, interaction strengths) is informed by comprehensive empirical literature, and detailed phase diagrams are constructed in silico to ensure that simulated regimes lie outside the miscibility gap for bulk NusA (preventing spurious bulk phase separation).
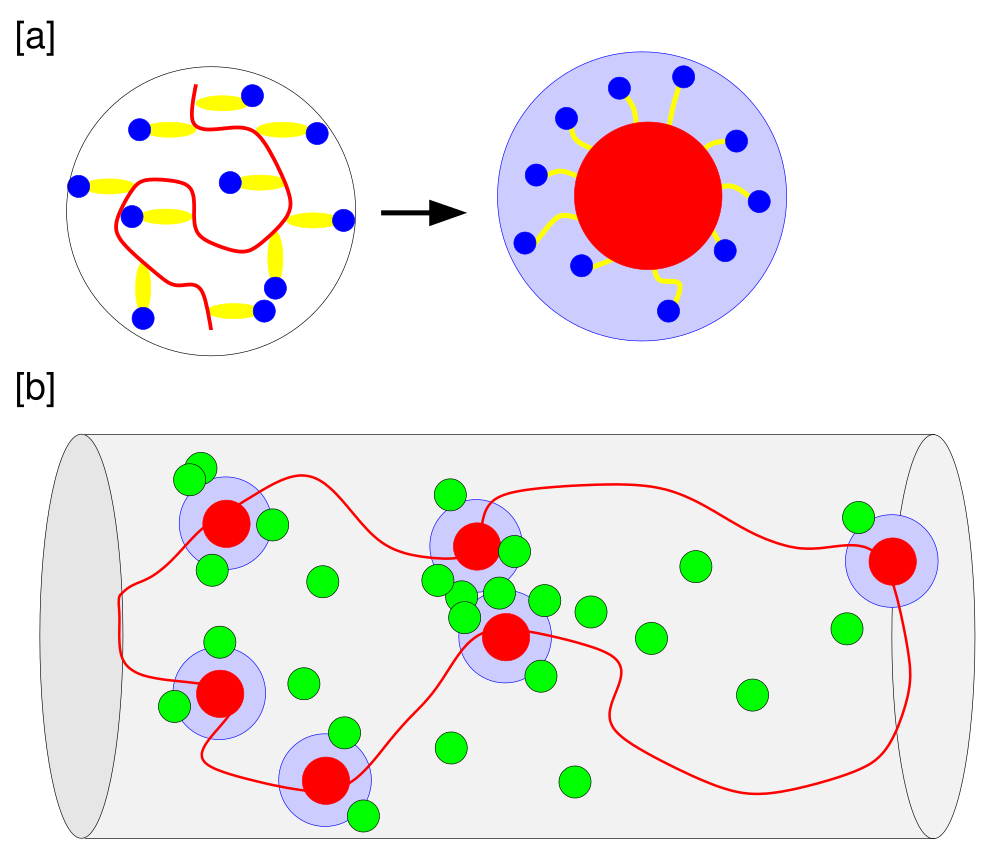
Figure 1: Schematic of the coarse-grained chromosome model and associated RNAP/Nus complexes around rrn operon loci.
PAC-Driven Protein Condensation and Chromosomal Reorganization
A principal insight of the paper is the demonstration that protein condensation—despite bulk concentrations/interaction strengths insufficient for homogeneous LLPS—arises through a polymer-assisted condensation (PAC) pathway. Here, the multivalent corona of bound Nus at rrn operons locally enriches free Nus protein density, nucleating condensate formation in cis even for globally subcritical conditions. This is a robust extension of prior PAC theory applied to eukaryotic heterochromatin, now contextualized for prokaryotic chromosome organization.
Simulation results confirm the selective nucleation of Nus condensates at rrn operon sites and subsequent coalescence, yielding statistical—but not permanent—colocalization of distant genomic loci.
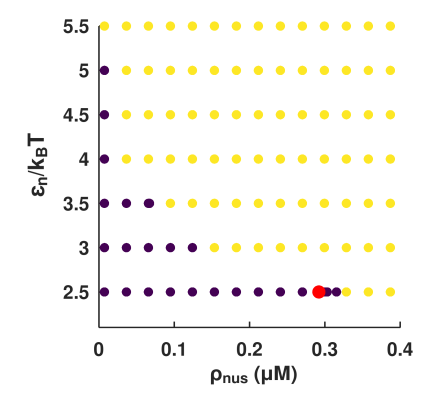
Figure 2: Simulation snapshots reveal spatial colocalization of rrn operons and bimodal neighbor distance distributions indicative of condensate formation via PAC.
Probability distributions of nearest-neighbor free Nus distances and cluster-size analyses show sharp transitions from homogeneous to condensed states as interaction strength crosses a threshold, with the largest clusters encompassing the majority of Nus content only above a critical LJ strength.
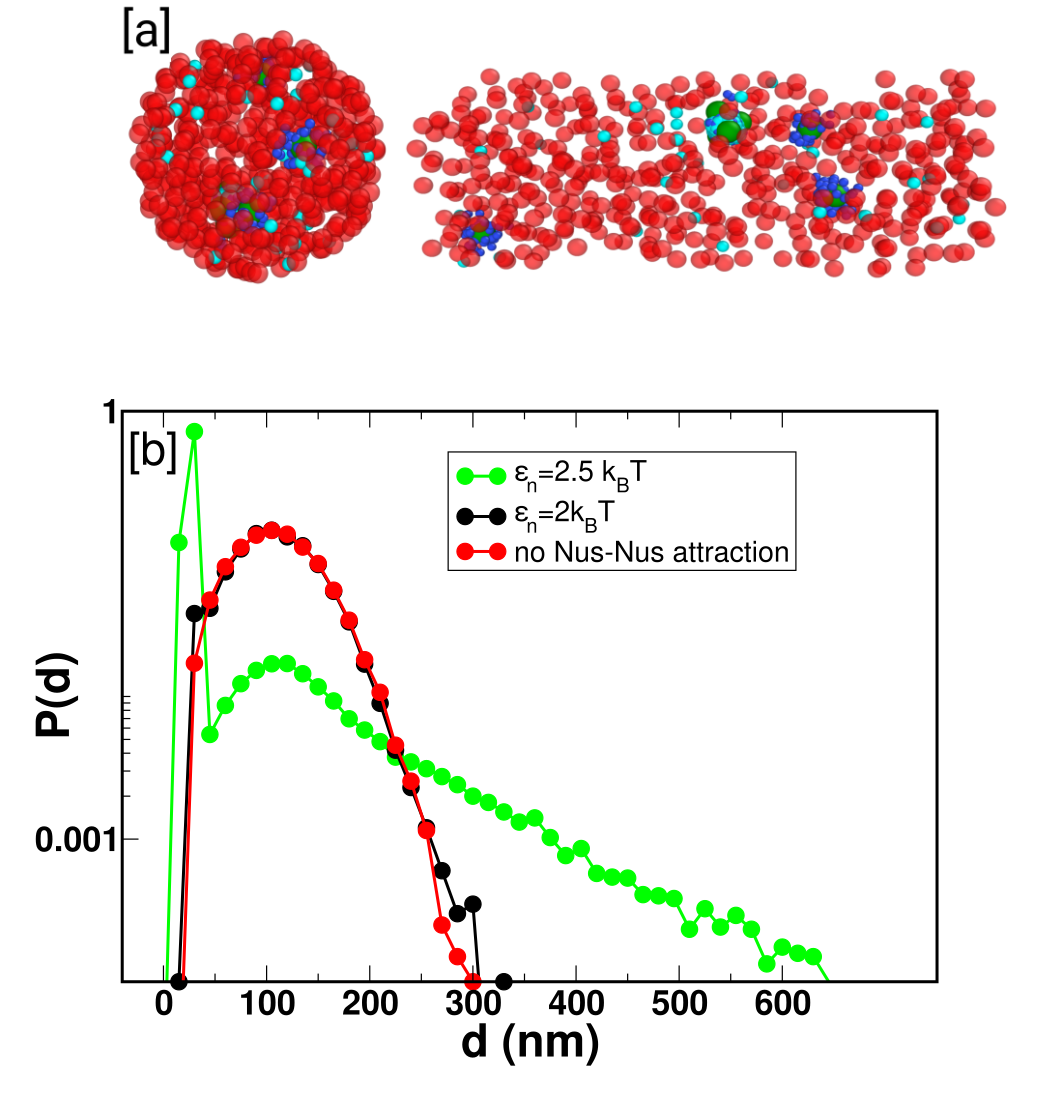
Figure 3: Cluster size distribution and radial composition analysis demonstrate peripheral localization of rrn operons, with Nus populations occupying both the core and periphery of clusters.
Dynamic Properties of Nus Populations and Correspondence with Experiment
Analysis of Nus dynamics demonstrates rapid exchange between condensed and dilute phases. The mean residence time of free Nus within a cluster is found to be short (∼11,000 MCS, corresponding via mapping to ~0.25 seconds experimentally), and the single-particle diffusion constants inside condensates are reduced by approximately two orders of magnitude compared to the cytoplasmic phase, quantitatively mirroring single-molecule experimental measurements.
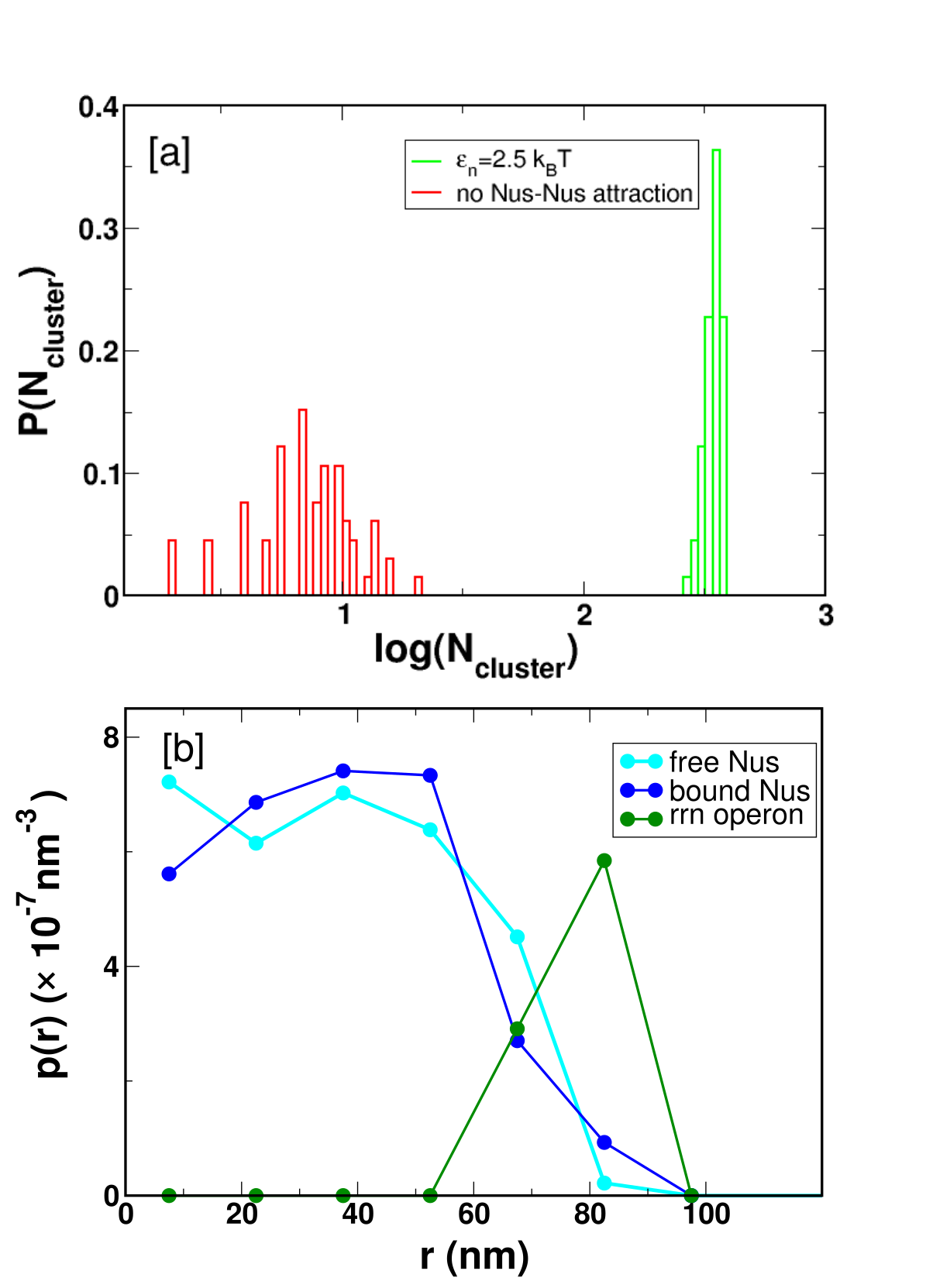
Figure 4: Residence-time statistics and diffusion constant distributions for Nus proteins highlight dynamic partitioning and long-lived yet dynamic condensates.
Simulations robustly reproduce the experimental observation that, under condensate-forming conditions, rrn operon monomers are statistically closer than expected for random pairs, with spatial distributions of pairwise operon distances showing sharply elevated short-range contact probabilities. However, these clusters, while potentially long-lived on simulation timescales, are not immortal.
Contact maps derived from the simulated ensembles—mirroring Hi-C protocols—highlight transient off-diagonal interactions among rrn operons that evolve dynamically over time, consistent with colocalization observed in fluorescence imaging but weak or washed out signals in time-averaged Hi-C data.
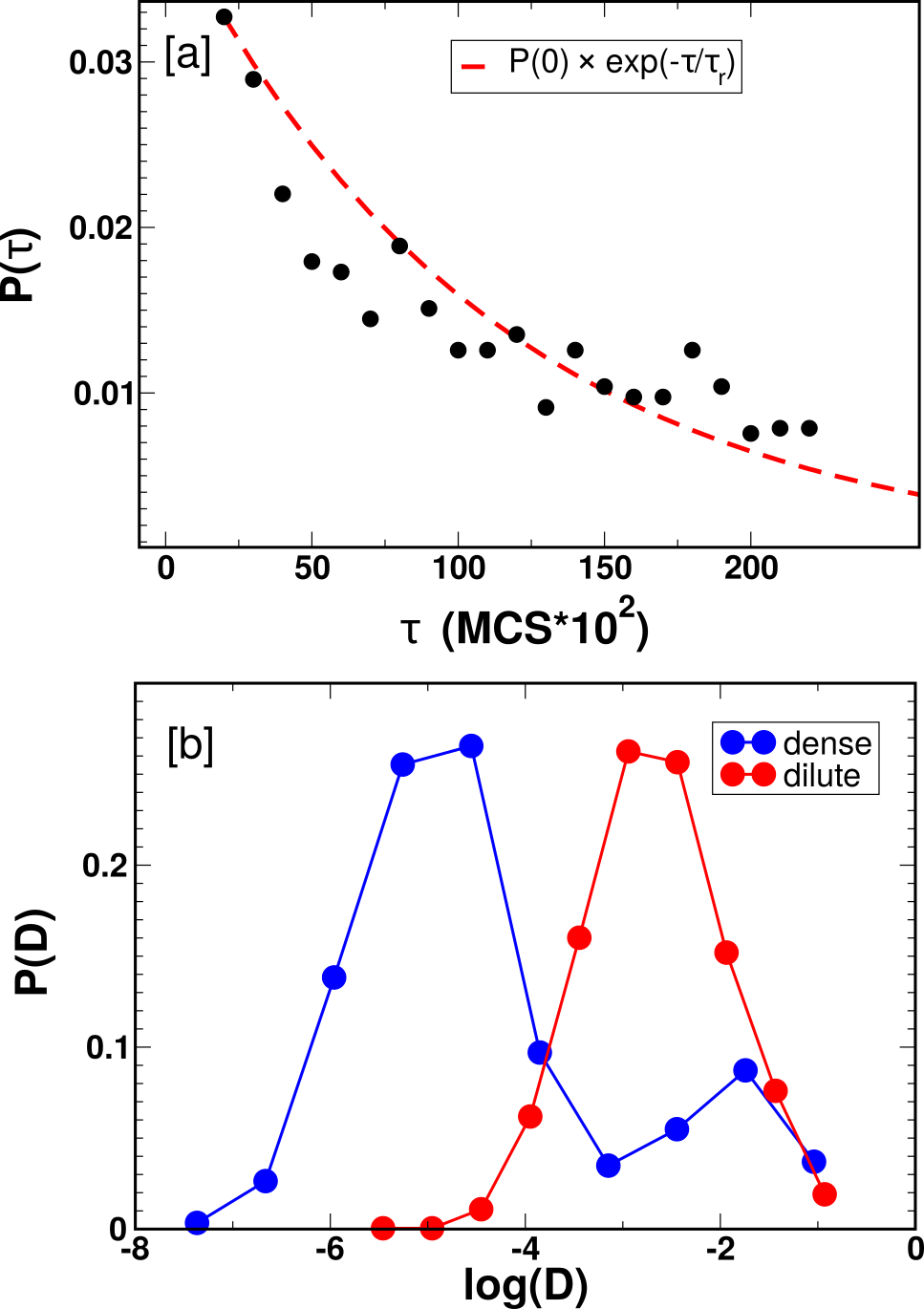
Figure 5: Pairwise rrn operon distance distributions and dynamic contact maps across simulation timescales, showing transient inter-operon proximity and the decorrelation of contacts during cluster coarsening.
Active Noise and Temporal Fluctuations: Reconciling Imaging and Hi-C Contradictions
To resolve the Hi-C/fluorescence imaging paradox, the model introduces temporal fluctuations (active noise) in Nus–Nus interaction strength, phenomenologically capturing the stochastic association/dissociation of RNAP from rrn operons during transcription. This is implemented as a time-dependent LJ parameter with Gaussian fluctuations, tuned to match empirically observed detachment times (~1 s real time).
Under these conditions, condensates form and dissolve transiently, resulting in ephemeral rrn operon colocalization. While pairwise distance distributions remain skewed toward shorter values (explaining statistical colocalization in imaging), time-averaged contact maps flatten (as in Hi-C), and the residence time of any two operons in proximity is short compared to Hi-C crosslinking timescales (∼10 min).
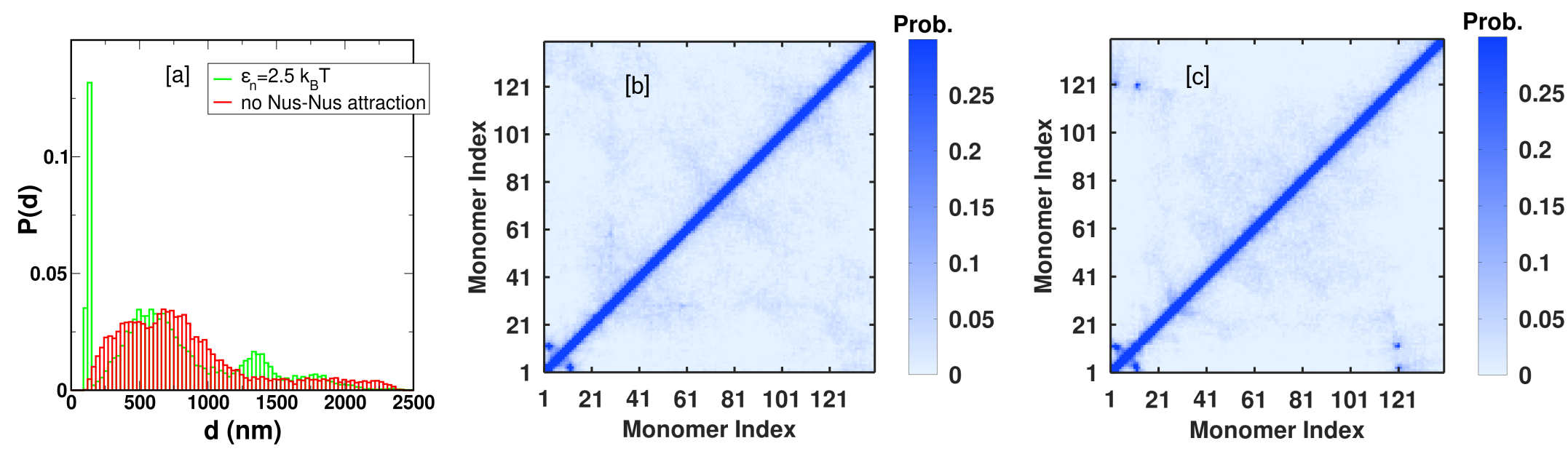
Figure 6: Incorporation of temporally fluctuating Nus interaction strengths leads to transient operon colocalization and rapid turnover, reconciling absence of persistent signals in Hi-C-derived contact maps.
Implications, Theoretical Advances, and Future Directions
This study provides a semi-quantitative physical framework for interpreting the mesoscale chromosome organization of bacteria in terms of generic polymer–protein interaction principles. The PAC mechanism, extended here to explain LLPS-mediated clustering by locally increasing protein concentration at specific genomic loci, generalizes the context-dependent nature of biomolecular condensation in confined environments.
The results bridge experimental findings from orthogonal techniques, with powerful implications for understanding how bacterial cells may exploit such mechanisms to regulate efficient ribosome production and spatial genome compartmentalization under varying physiochemical environments. The analogy to eukaryotic nucleolar assembly—where ribosome biogenesis loci and proteins coalesce into multilayered condensates—is especially pertinent, hinting at conserved physical strategies across domains of life.
Limitations include the neglect of rRNA extrusion, ongoing biochemical processing, explicit energetic costs of loop formation, and the coarse-graining of dynamic transcriptional activity. Incorporation of persistent rRNA outflux, additional protein–protein/ protein–nucleic acid effectors, and explicit modeling of supercoiling/topological constraints are promising avenues for future modeling work. Simulation-based approaches integrating polymer physics with single-molecule biochemistry will be instrumental in further dissecting the principles underlying genome organization.
Conclusion
Through a detailed simulation-based analysis, this work demonstrates that polymer-assisted condensation of Nus proteins at rrn operons can generate spatially colocalized clusters of RNA polymerase complexes in E. coli. The model reconciles disparate experimental observations by accounting for both equilibrium and non-equilibrium effects (active noise) in the dynamics of condensate formation. The results underscore the utility of physical polymer–protein frameworks for dissecting the spatial organization of bacterial chromosomes and suggest new questions regarding the functional consequences of mesoscale condensation for ribosome biogenesis and global transcriptional regulation.







