- The paper presents a weakly supervised self-learning framework that leverages global Gleason scores to generate patch-level pseudo-labels.
- It employs a teacher-student CNN architecture with multiple-instance learning and max-pooling, achieving a Cohen's quadratic kappa of 0.82 on patch-level grading.
- The approach reduces reliance on detailed pixel-level annotations, offering increased generalizability and robustness in computational pathology.
Self-learning for Weakly Supervised Gleason Grading of Local Patterns
Introduction
The paper "Self-learning for weakly supervised Gleason grading of local patterns" (2105.10420) introduces an innovative methodology to tackle the challenges in grading prostate cancer via histological images using deep learning techniques. The Gleason grading system remains the gold standard for prostate cancer diagnosis and prognosis. However, manual grading of prostate cancer through histology slides is highly subjective and time-consuming. Computer-aided diagnosis (CAD) systems aim to assist pathologists in this task, but they typically require labor-intensive pixel-level annotations. This paper proposes a weakly supervised self-learning framework using convolutional neural networks (CNNs) to overcome these limitations by leveraging global Gleason scores of whole slide images (WSIs).
In prostate cancer, Gleason grades (GG3, GG4, GG5) categorize cancer severity based on gland morphology observed in histology samples (Figure 1). The diagnostic process involves grading local glandular patterns, using the two most prominent grades to determine a global Gleason score, which is crucial but subject to high variability.
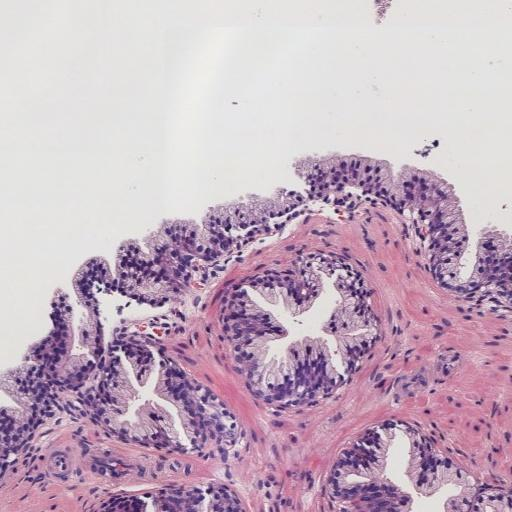
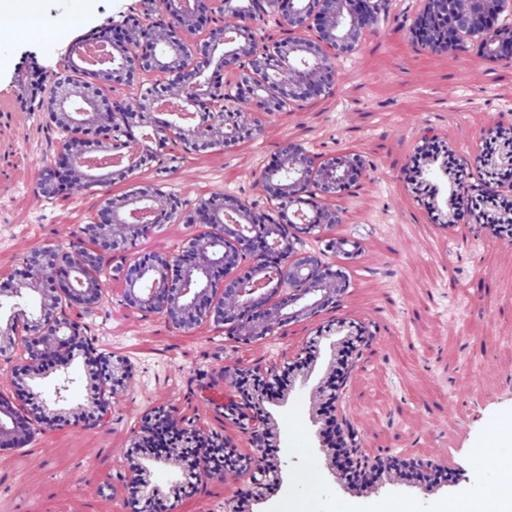
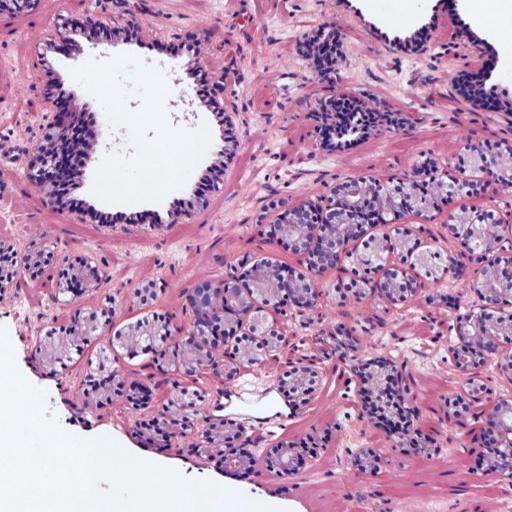
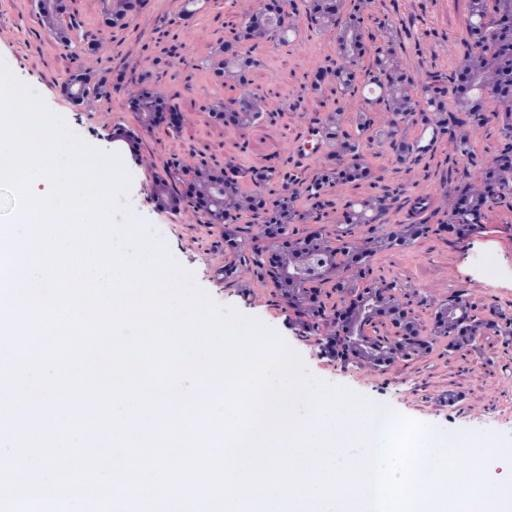
Figure 1: Histology regions of prostate biopsies. (a): region containing benign glands, (b): region containing GG3 glandular structures, (c): region containing GG4 patterns, (d): region containing GG5 patterns.
Methods
The proposed method comprises two main stages using a teacher-student training framework. The teacher model employs a multiple-instance learning (MIL) approach to infer patch-level Gleason grades from biopsy-level annotations by aggregating patch-level inferences (Figure 2). The teacher model uses global image labels to generate pseudo-labels for patch-level learning.
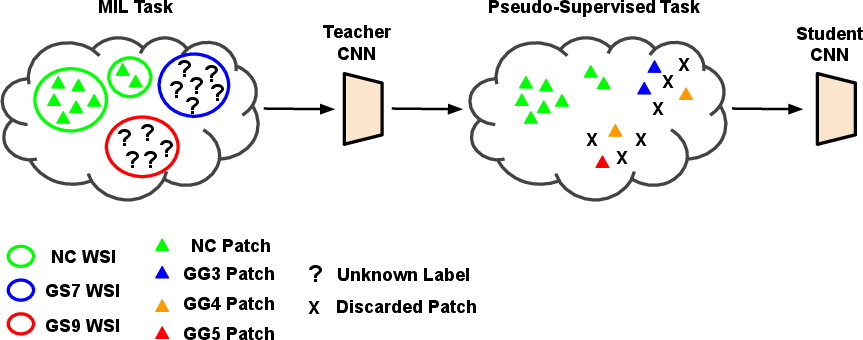
Figure 2: Self-learning CNNs pipeline for weakly supervised Gleason grading of local cancerous patterns in whole slide images. MIL: multiple-instance learning; WSI: whole slide image; NC: non-cancerous; GS: Gleason score; GG: Gleason grade.
During the prediction phase, the teacher model applies a max-pooling operation to combine the inferences of local instances into whole slide predictions, optimizing high-confidence predictions (Figure 3).
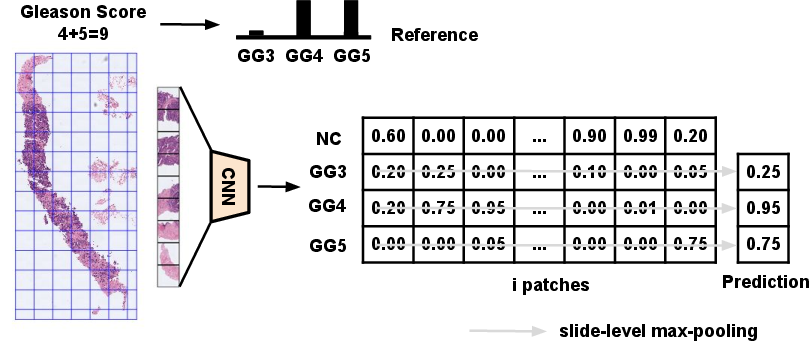
Figure 3: Teacher CNN for the prediction of local Gleason grades in a multiple-instance learning framework. GG: Gleason grade.
Subsequently, a label refinement process post-produces pseudo-labels from the teacher model, which are then used to train an equivalent student model on a pseudo-supervised dataset. This step eliminates false negative patches and enhances the learning capabilities of the student model.
Results
Experiments evaluated the methods across different datasets for patch-level grading and biopsy-level scoring. For patch-level Gleason grading, the student model using max-pooling achieved an inter-dataset average Cohen's quadratic kappa of 0.82, indicating substantial agreement, outperforming previous supervised approaches that suffered from generalization limitations to external datasets (Figure 4).
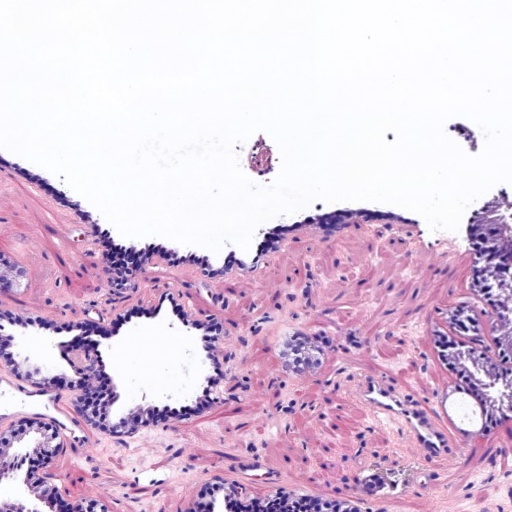
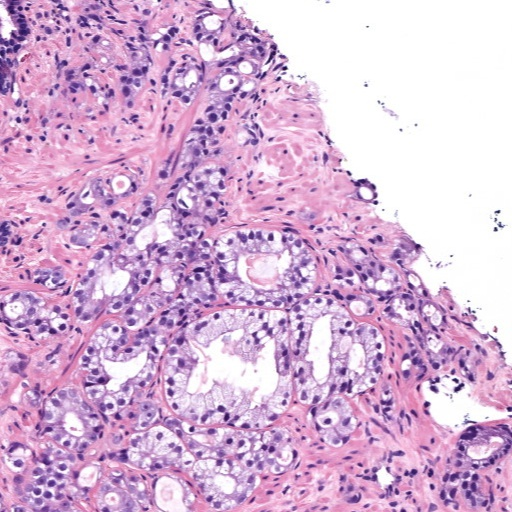
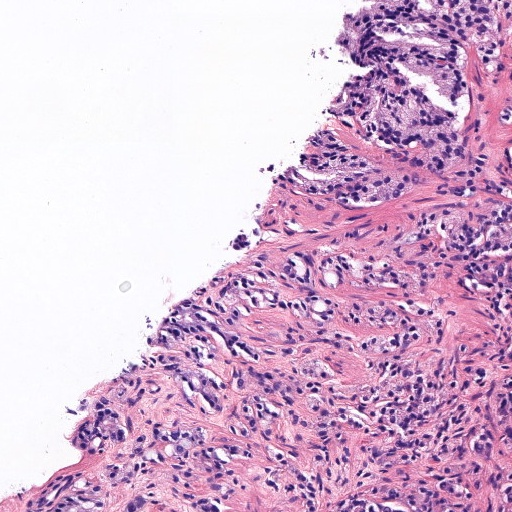
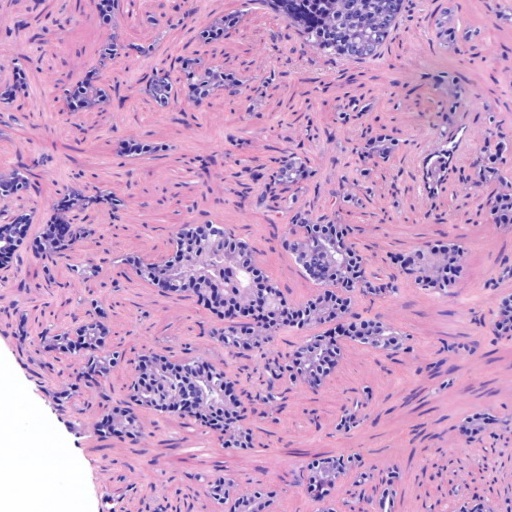
Figure 4: Histology regions with mixed Gleason grades from SICAP dataset. (a): region containing benign and GG3 glands, (b): region containing GG3 and GG4 glandular structures, (c) and (d): region containing GG4 and GG5 patterns.
Confusion matrices display the high accuracy of the student model for predicting patch-level Gleason grades on various datasets, showing adaptability to diverse cases (Figure 5).
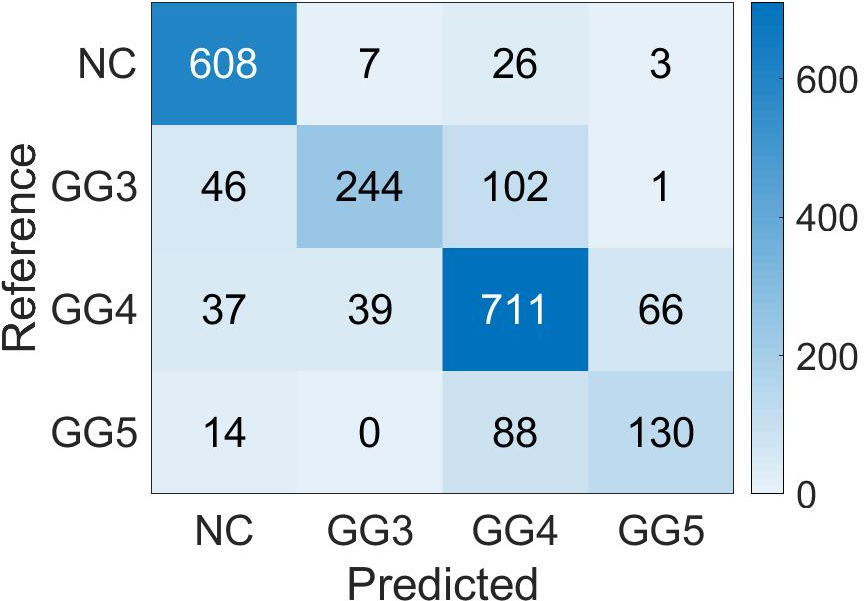
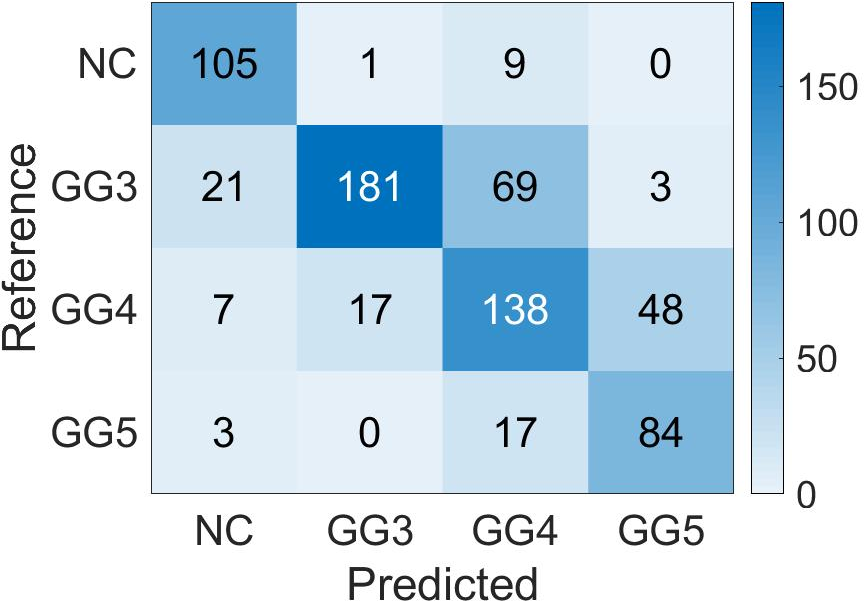
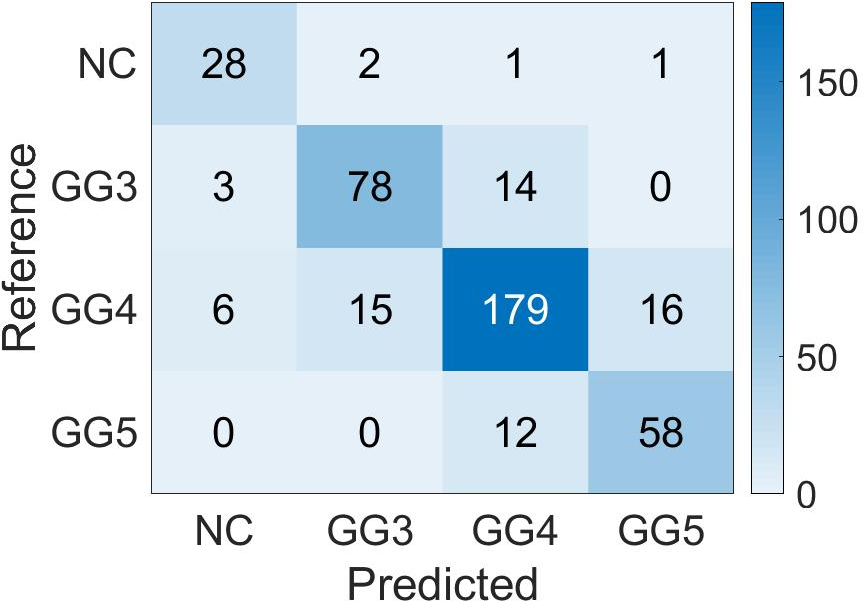
Figure 5: Confusion Matrix of the patch-level Gleason grades prediction done by Student CNN on the different test cohorts. (a): SICAP; (b): ARVANITI; (c): GERTYCH.
For global biopsy scoring, the student model's features were averaged and used with k-nearest neighbor classifiers to predict biopsy-level Grade Groups (Figure 6). This methodology proved superior in external validation tests, providing robust clustering results over conventional Gleason grading percentage approaches.
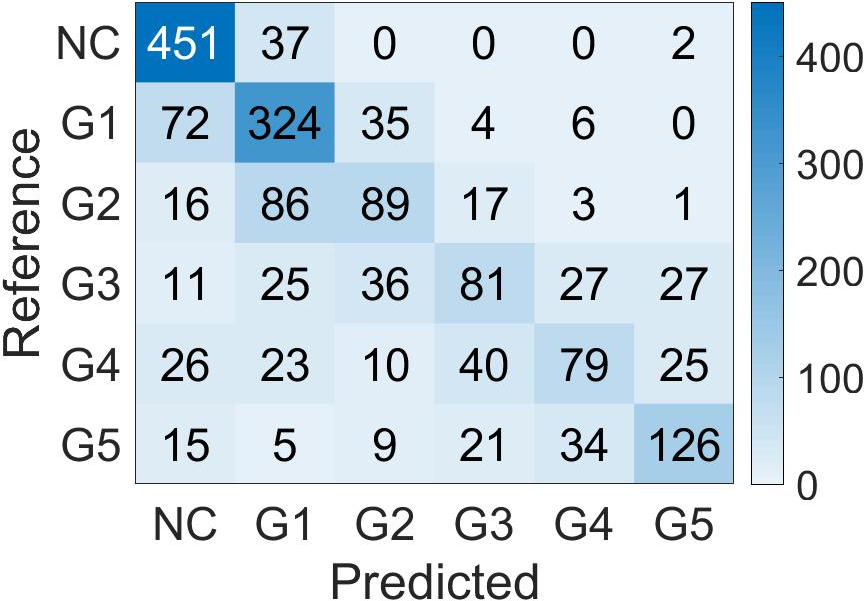
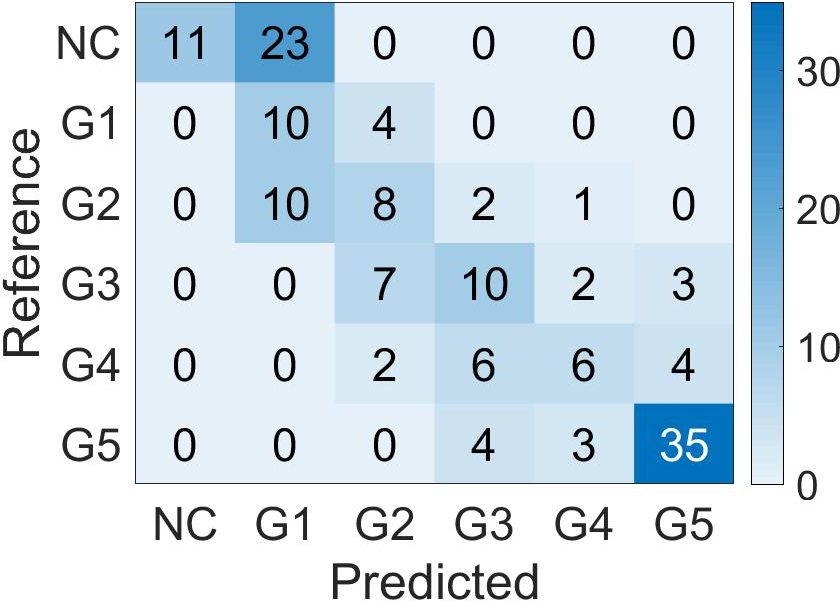
Figure 6: Confusion Matrix of the biopsy-level Grade Group prediction done by Student CNN features and k-Nearest Neighbor classifier, on the two test cohorts. (a): PANDA; (b): SICAP.
Discussion
The implications highlight the potential of weakly supervised learning frameworks to improve the reliability and robustness of Gleason grading systems. The removal of annotator bias and leveraging vast datasets enables the development of more generalizable models. Furthermore, this strategy could lead to reduced computational constraints since biopsies can be processed completely without pixel-level annotations.
Future research may explore optimizing normalization techniques to enhance generalization further and refine the framework for even larger biopsy datasets, which might provide more extensive coverage of heterogeneous cancer patterns.
Conclusion
The paper presents a significant contribution to the field of histopathology with its innovative use of self-learning and CNNs for prostate cancer grading. The system's superior performance in patch-level grading and biopsy-level scoring, the capacity to generalize without pixel-level annotations, and the focus on leveraging global Gleason scores mark a substantial advance. This approach not only validates the promise of weakly supervised learning but also paves the way for further developments in computational pathology.














