- The paper introduces a deep learning system using a U-Net architecture to automate Gleason grading, achieving a quadratic Cohen's kappa of 0.918 with consensus grade groups.
- It employs a semi-automated labeling process to segment cell-level images and compute normalized tissue volume percentages, reducing reliance on manual annotations.
- The system outperforms 10 out of 15 pathologists and demonstrates strong ROC performance (0.990 for malignancy, 0.978 for grade group stratification), enhancing diagnostic precision.
Automated Gleason Grading of Prostate Biopsies using Deep Learning
Introduction
In the domain of prostate cancer diagnostics, the Gleason score serves as a critical prognostic marker, yet its application is plagued by inter- and intra-observer variability which compromises its utility. The paper "Automated Gleason Grading of Prostate Biopsies using Deep Learning" (1907.07980) introduces a deep learning-based system designed to automate the process of Gleason grading in prostate biopsies, aiming to enhance reproducibility and accuracy across different healthcare settings. This system leverages a semi-automated labeling process to circumvent the need for exhaustive manual annotations.
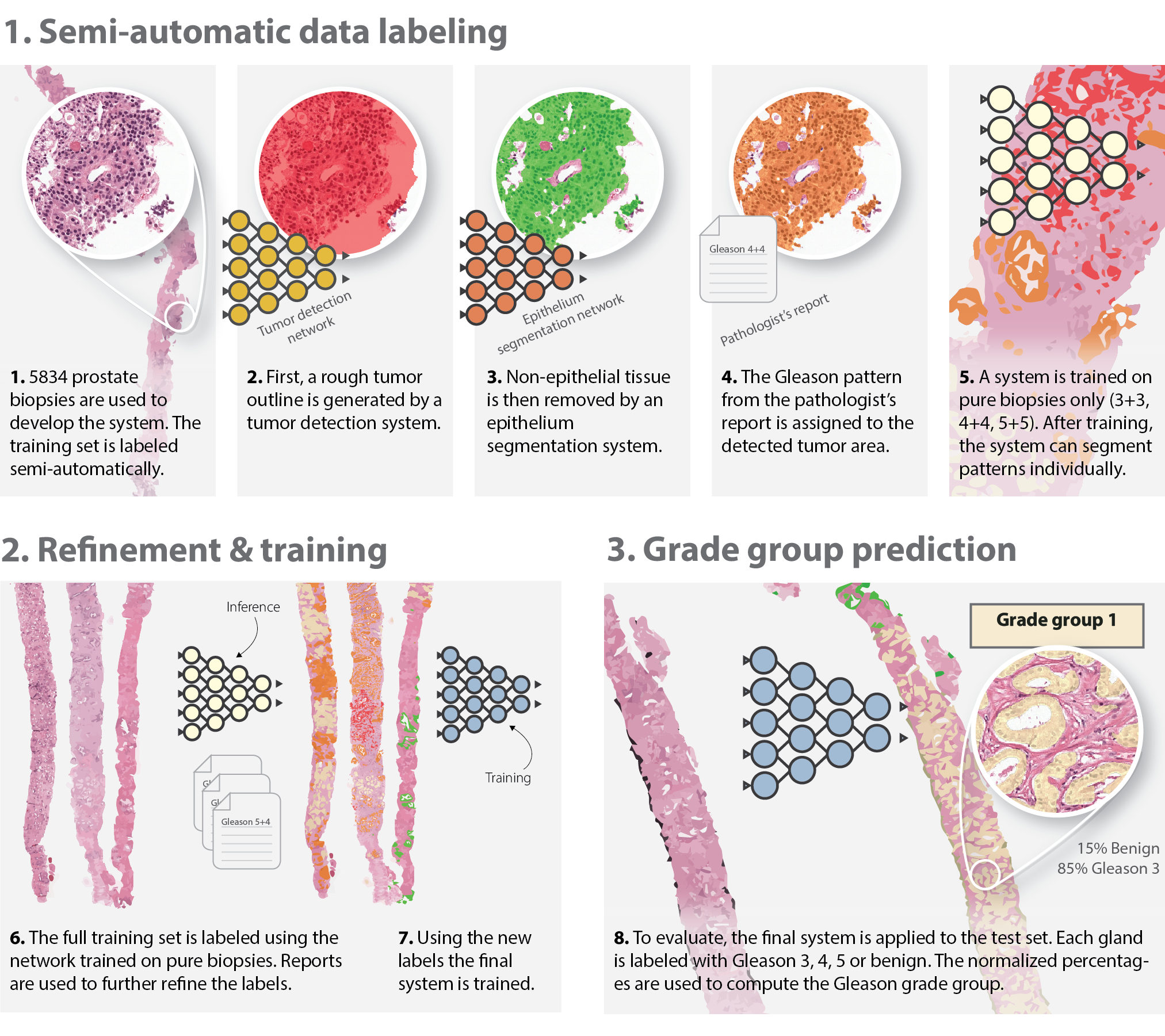
Figure 1: Overview of the development of the deep learning system. We employ a semi-automated method of labelling the training data (top row), removing the need for manual annotations by pathologists.
Development of the Deep Learning System
The core of the system is based on the U-Net architecture, known for its proficiency in image segmentation tasks. The deep learning model operates by segmenting images at the cell-level and assigning Gleason growth patterns to tumorous glands. This approach allows the system to compute normalized tissue volume percentages, which then inform the determination of Gleason scores and grade groups. The absence of manual pixel-level annotations is compensated by a semi-automatic labeling technique combining pre-existing segmentation algorithms and slide-level grades from pathology reports. This design not only increases efficiency but also enables training on large datasets without extensive labeling efforts.
Results and Comparison
Contrasting the system against an external panel of pathologists, the deep learning system achieved a quadratic Cohen's kappa of 0.918 with the consensus grade group on the test set. Notably, the system outperformed 10 out of 15 pathologists in an external evaluation panel showcasing its robust performance against varying levels of human expertise.
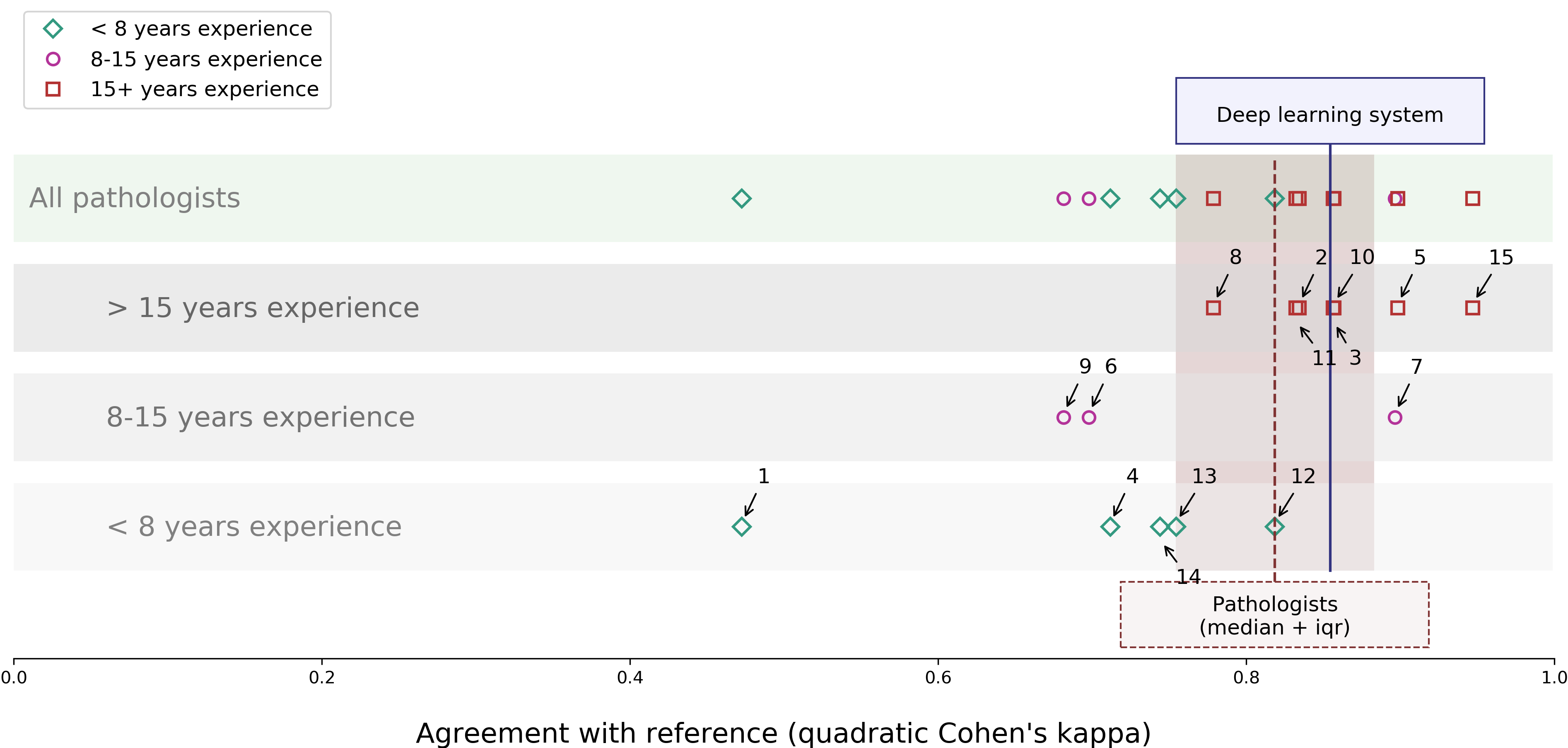

Figure 2: Agreement on Gleason grade group between each pathologist of the panel and the deep learning system with the consensus. The panel members are split out according to their experience level. Additionally, the median kappa of the pathologists is shown in brown.
Additionally, the system demonstrated strong ROC performance in both distinguishing malignant from benign biopsies, and using grade group 2 as a threshold for higher grade tumors. With an area under the curve of 0.990 for malignancy determination and 0.978 for stratification by grade group 2, it showcases effective clinical applicability in biopsy-level risk stratification.
Implications and Future Directions
The implications of this study are multifaceted, spearheading advancements in computational pathology driven by deep learning. Practically, the system could be integrated as a pre-screening or secondary review tool in pathology laboratories, heightening diagnostic precision and prioritizing high-grade biopsy assessments. Theoretically, it sets a precedent for leveraging deep learning systems in medical diagnostics where historical manual grading systems are fraught with subjectivity and variability.
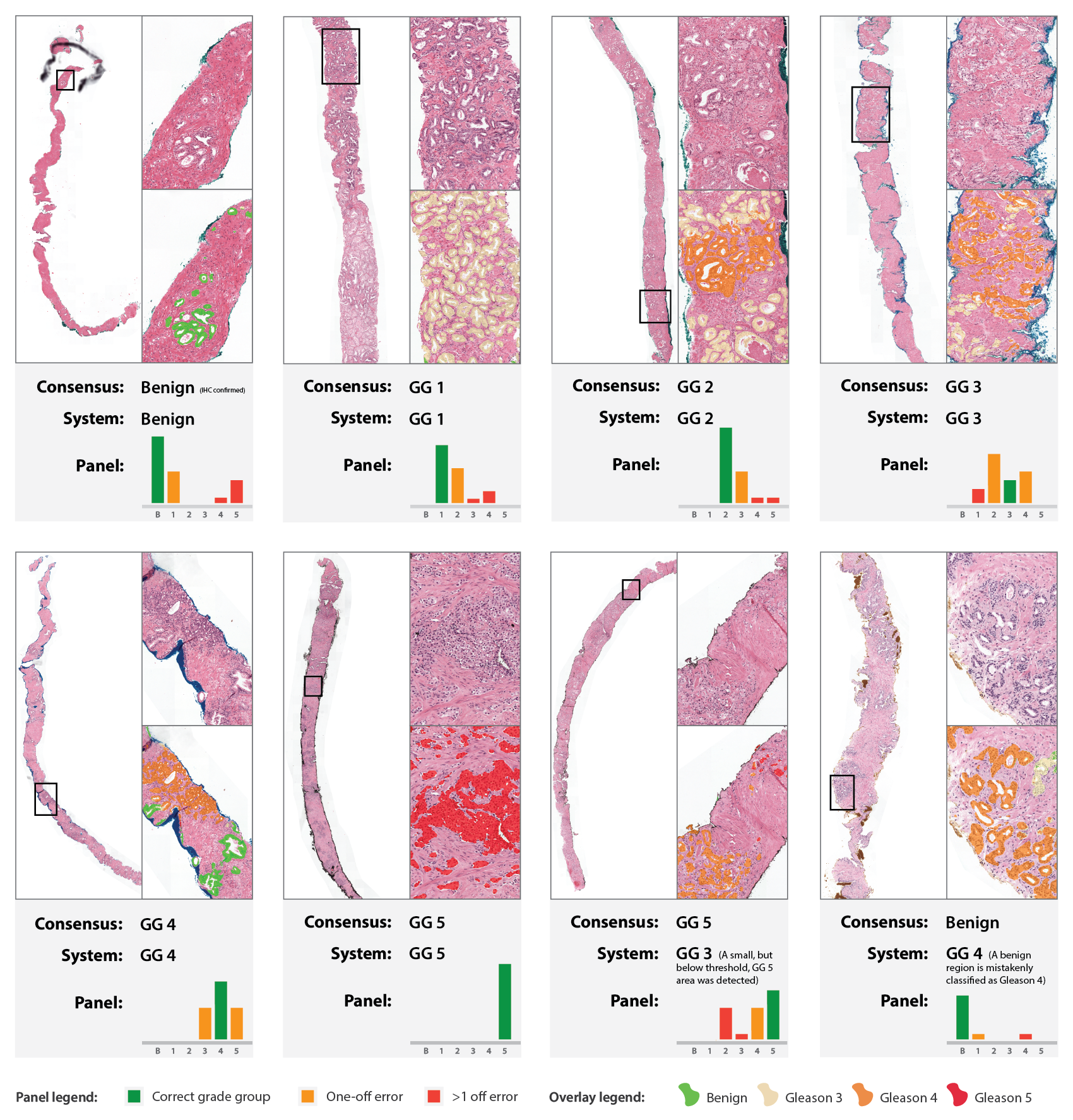
Figure 3: Examples from the observer set. For each case, the grade group of the reference standard, the predicted grade group by the deep learning system, and the distribution of grade groups from the panel is shown.
Future research could expand on this foundation by incorporating data from diverse geographic and clinical settings, thus further validating and potentially improving model robustness across varying histopathological parameters. Additionally, extending the system's capabilities to recognize other tumor types beyond acinar adenocarcinoma in prostate biopsies could lead to comprehensive diagnostic tools applicable across a broader spectrum of oncological pathology.
Conclusion
The study illustrates the significant potential of deep learning for enhancing the accuracy and efficiency of Gleason grading in prostate cancer diagnosis. The automated system not only matches but in many instances surpasses clinician performance across varying contexts, promising a reliable computational alternative that is both scalable and interpretable. As the field of AI-driven medical diagnostics continues to evolve, systems like this could pave the way for more consistent, expert-level healthcare solutions in prostate cancer prognostics.


