- The paper demonstrates that retrieval-augmented generation significantly boosts LLM accuracy in staging pancreatic cancer, achieving 70% overall accuracy.
- It employs a five-step prompt leveraging Japanese clinical guidelines to rigorously assess TNM classification, local invasion, and resectability.
- The study underscores the benefit of evidence-backed retrieval for transparency while noting that misinterpretations persist, necessitating expert oversight.
Retrieval-Augmented Generation Improves Pancreatic Cancer Staging Accuracy in LLMs
Introduction
This study evaluates the impact of Retrieval-Augmented Generation (RAG) on pancreatic cancer staging using LLMs. The principal experimental comparison is between NotebookLM (a RAG-LLM employing Gemini 2.0 Flash as its internal model) and Gemini 2.0 Flash deployed as a standalone LLM, focusing on the isolated effect of RAG rather than model architecture. The investigation leverages reliable external knowledge (REK) in the form of Japanese pancreatic cancer staging guidelines to assess model-driven classifications across 100 fictional CT-based cases.
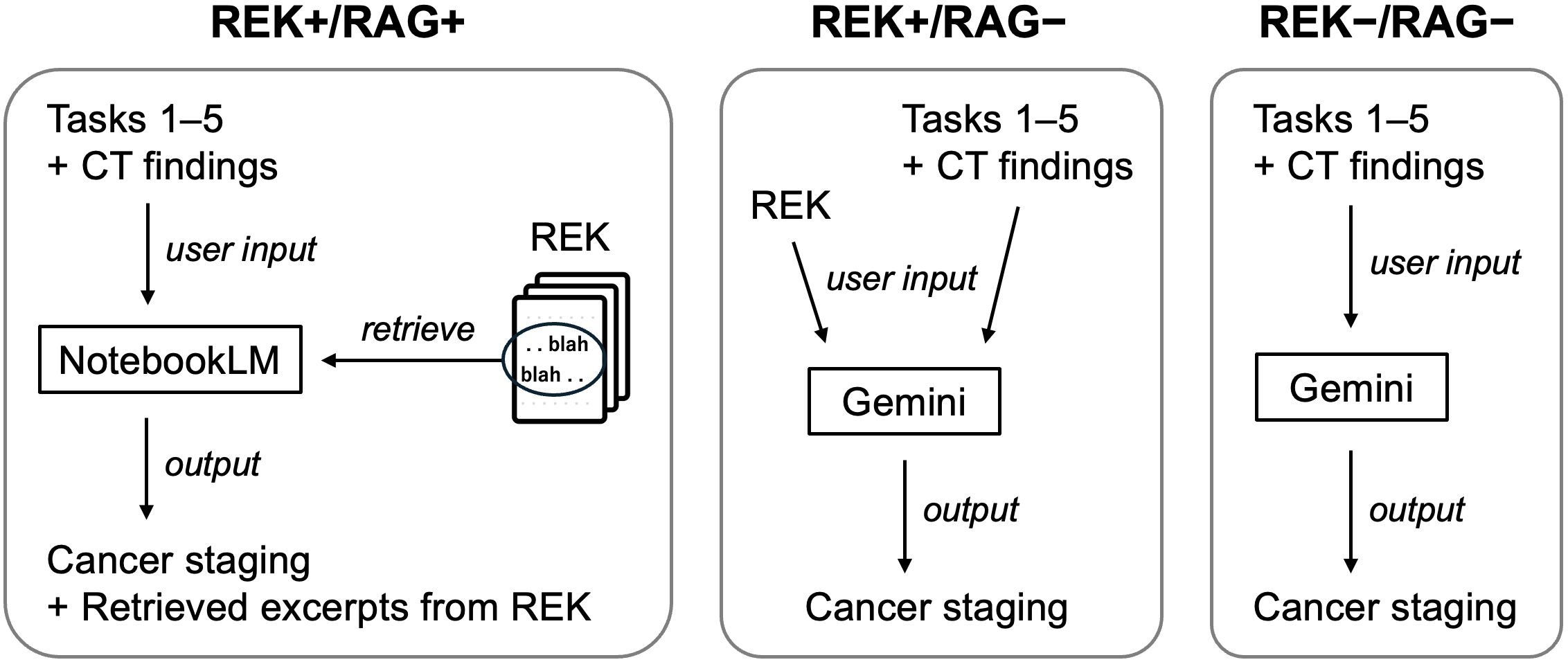
Figure 1: Overview of the experimental workflow, delineating roles of REK, RAG, and LLM configurations in cancer staging.
Experimental Design and Methodology
The dataset comprises 100 radiologist-generated fictional pancreatic cancer cases, with standardized CT findings and staging components including TNM classification, detailed local invasion factors, and resectability assessments. Staging guidelines were extracted from the eighth edition of the Japanese classification of pancreatic carcinoma, encoded as REK.
Three experimental groups were defined:
- REK+/RAG+: NotebookLM with REK using integrated RAG.
- REK+/RAG-: Gemini 2.0 Flash with REK loaded manually (no retrieval).
- REK-/RAG-: Gemini 2.0 Flash without REK (model-only).
LLMs were tasked via a five-step prompt sequence covering local invasion, TNM assignment (T, N, M), and downstream resectability assessment. Staging was correct only if every component matched ground truth.
Results
NotebookLM (REK+/RAG+) yielded a staging accuracy of 70%, compared to Gemini 2.0 Flash with REK at 38%, and Gemini 2.0 Flash without REK at 35%. TNM classification accuracy for NotebookLM reached 80%; Gemini 2.0 Flash with REK achieved 55%, and without REK 50%. Notably, NotebookLM provided explicit supporting excerpts from REK for its decisions, attaining a 92% retrieval accuracy.
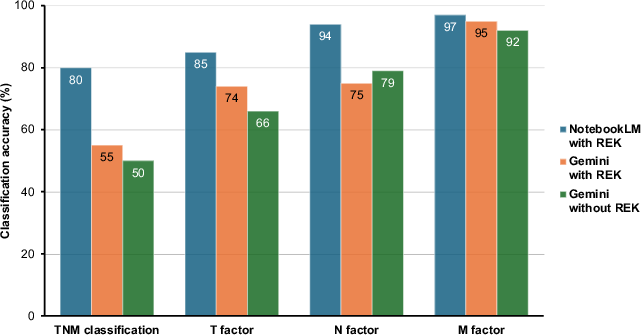
Figure 2: TNM classification performance, with NotebookLM (REK+/RAG+) showing marked gains in overall and factorwise accuracy.
Classification of local invasion factors mirrored the same trend. However, improvement in resectability category accuracy was less prominent; detailed breakdowns indicate that model misinterpretation, even in the presence of accurate retrieval, can still occur.
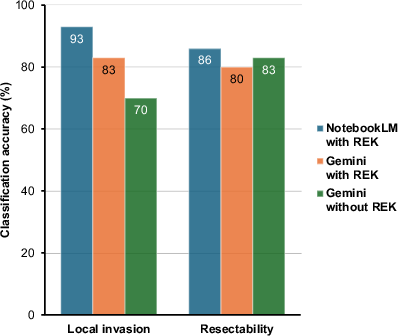
Figure 3: Comparative performance for local invasion factor determination and resectability classification across model groups.
Representative case analyses (see Figure 4 and Figure 5 in the manuscript) illustrate correct staging and retrieval (Case 98) and misclassification despite correct retrieval (Case 48), demonstrating both the strengths and residual weaknesses of retrieval-augmented workflows.
Discussion
The findings reinforce RAG's direct contribution to LLM reliability and accuracy in clinical guideline-driven classification. Prior studies using GPT-3.5/4 for pancreatic cancer-related tasks similarly reported insufficient clinical-grade accuracy without structured knowledge injection, while RAG-LLMs showed substantial performance improvement and the ability to ground output in evidence (2503.15664).
Nevertheless, hallucinations and misinterpretation of clinical text persist, even with high retrieval accuracy. For instance, model confusion between ‘splenic vein’ and ‘portal vein’ produced resectability classification errors, underscoring the persistent need for human expert oversight.
Practical deployment in radiology confronts data privacy concerns for internet-based LLMs; as a result, there is an evident shift toward open-source, locally operating LLMs with RAG capabilities, which can ensure information security and regulatory compliance.
The robustness of RAG improvements may depend on model capacity and REK complexity. Direct prompt injection of complete guidelines is insufficient for optimal performance if limits on context size or physician efficiency are reached. Extraction of concise, context-relevant excerpts via RAG remains crucial.
Implications and Future Directions
RAG-LLMs can support radiologists by not only improving classification accuracy but also by providing transparent, evidence-backed rationale for every classification. As guideline complexity and medical document length grow, the need for partial, targeted retrieval will intensify.
Immediate future work should include:
- Testing on real clinical datasets
- Evaluating cost-benefit of RAG-based partial knowledge delivery versus full guideline ingestion
- Exploration of differential diagnosis and broader clinical tasks
- Deployment of high-security, on-premises RAG-LLMs
Conclusion
NotebookLM, powered by Gemini 2.0 Flash with RAG, significantly outperforms unretrieved Gemini 2.0 Flash in pancreatic cancer staging, demonstrating the functional merit of retrieval-augmented workflows for guideline-driven medical tasks. However, persistent model misinterpretation issues dictate that these systems should remain adjunctive rather than standalone in clinical decision-making. For medical use cases, migration toward locally operable RAG-LLMs appears essential due to privacy and regulatory concerns.


