- The paper introduces a dual RAG architecture combining FAISS-based retrieval and a QLoRA fine-tuned Mistral-7B model for accurate biomedical Q&A.
- The methodology leverages semantic embeddings and document normalization, achieving BERTScore F1 improvements up to 0.88-0.90 for breast cancer queries.
- The system demonstrates potential for expanding multilingual and personalized applications by bridging complex biomedical literature with user-friendly AI interfaces.
Biomedical Literature Q&A System Using Retrieval-Augmented Generation (RAG)
Introduction
The paper "Biomedical Literature Q&A System Using Retrieval-Augmented Generation (RAG)" (2509.05505) explores the integration of retrieval and generation mechanisms to enhance medical question-answering systems. The RAG architecture is employed to deliver precise, evidence-based responses by leveraging a combination of neural retrieval mechanisms and generative LLMs. This system addresses the limitations inherent in traditional medical search engines and LLMs by enhancing factual accuracy and contextual grounding.
Methodology
RAG Architecture
The proposed system employs a RAG architecture that integrates retrieval processes with generative models to provide contextualized and accurate responses to medical queries. The retrieval component utilizes FAISS for vector-based similarity search driven by semantic embeddings generated via a mini-LM model, which is optimized for retrieval-centric tasks. The generative component is powered by a fine-tuned Mistral-7B-v0.3 model, employing QLoRA for parameter-efficient training. This dual approach ensures the retrieval of relevant medical documents that guide the LLM in producing responses that are both fluent and factually accurate.
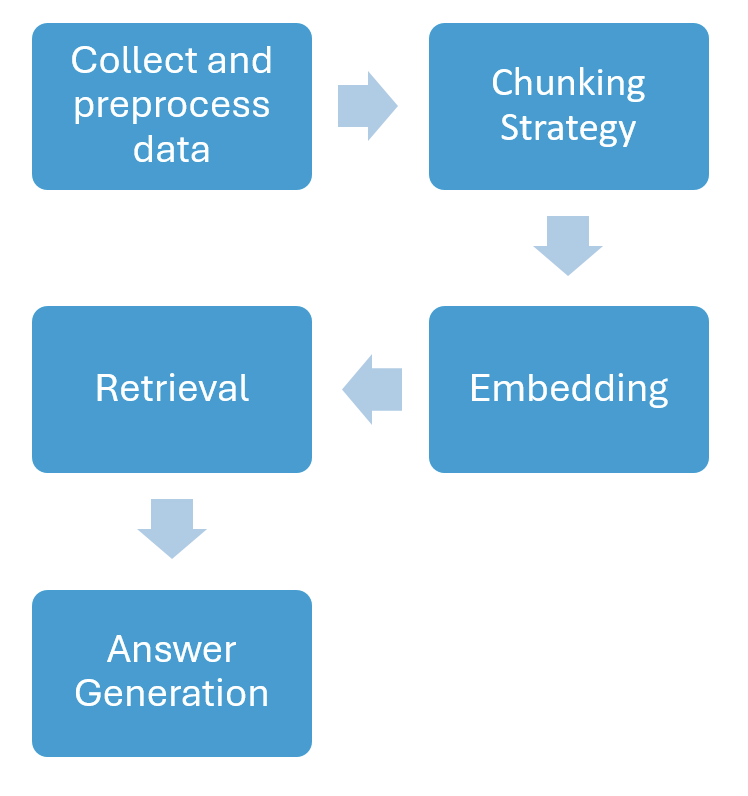
Figure 1: RAG Pipeline.
The system collects data from diverse sources such as PubMed articles, medical encyclopedias, and custom datasets. Key preprocessing steps include document normalization, chunking, and noise removal, which enhance both retrieval and generation stages.
Evaluation and Results
The system was evaluated using the BERTScore metric, which measures semantic fidelity and factual consistency through contextual embeddings. The evaluation was conducted in three configurations: a baseline zero-shot model, a retrieval-augmented model, and a retrieval-augmented model fine-tuned with QLoRA.
Empirical results demonstrated significant improvements in response quality with retrieval-augmented strategies, particularly in domain-specific contexts such as breast cancer. The fine-tuned RAG model achieved a BERTScore F1 score of 0.843 on a comprehensive medical dataset, outperforming baseline models and illustrating the benefits of contextual augmentation.
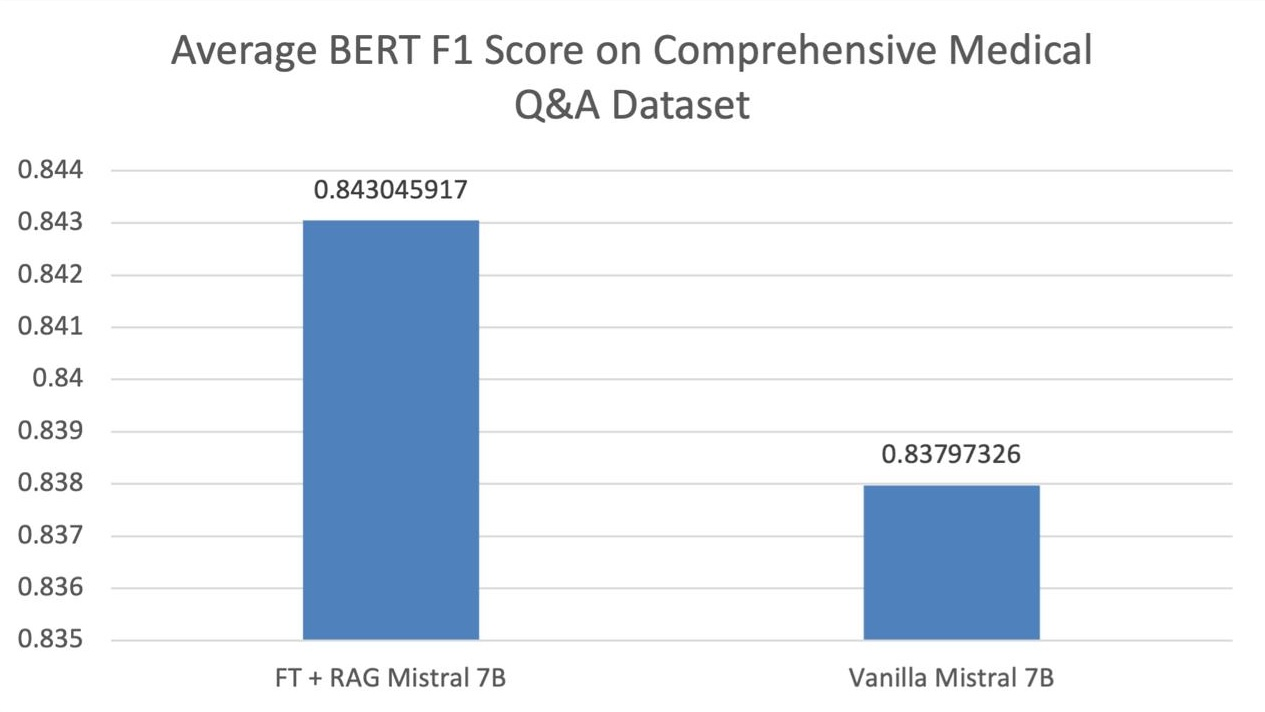
Figure 2: Performance Comparison on Comprehensive Medical Q&A Dataset.
Further experiments on a breast cancer-specific dataset corroborated these findings, with an observed BERTScore F1 improvement to 0.88-0.90. These outcomes highlight the criticality of domain-specific data in maximizing model performance and establishing reliable biomedical information systems.
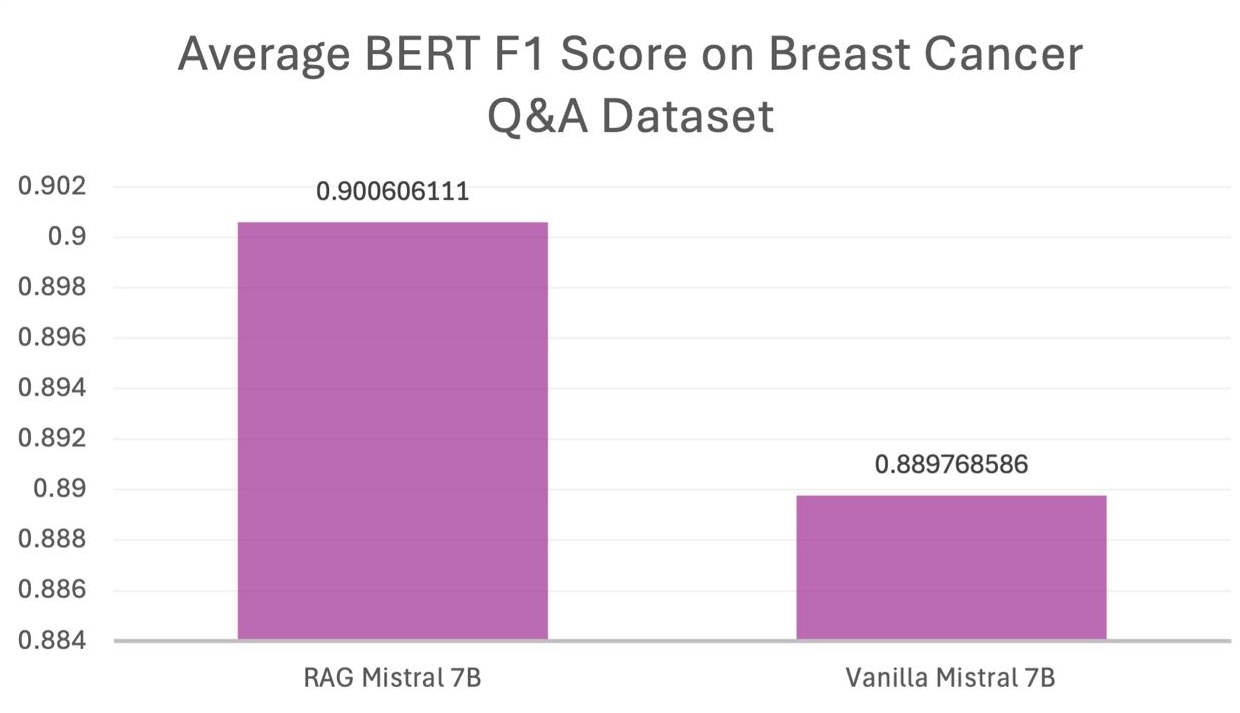
Figure 3: Performance Comparison on Breast Cancer Scraped Dataset.
Implications and Future Directions
This work underscores the potential of RAG-based systems to enhance public access to trustworthy medical knowledge. By bridging the gap between complex biomedical literature and user-friendly interfaces, the system serves as a template for future developments in medical AI. Potential extensions include multilingual adaptation, enhanced privacy-preserving techniques, and personalized user interactions to further extend the system's applicability across different linguistic and cultural domains.
Scalability considerations, particularly in contexts with language-specific medical terminologies, necessitate future research. Moreover, integrating context-aware multi-turn dialogues and user profiling could improve response personalization and engagement, laying a foundation for a more holistic medical consultation experience.
Conclusion
The deployment of a Retrieval-Augmented Generation system represents a step forward in leveraging AI to improve healthcare information delivery. Through careful integration of retrieval and generation components, the system demonstrates improved factual accuracy and response relevance, particularly in specialized medical domains. These findings lay groundwork for advancing AI-driven healthcare applications, promoting efficient and accessible medical information dissemination, and providing a robust platform for future exploration in personalized medical AI systems.


