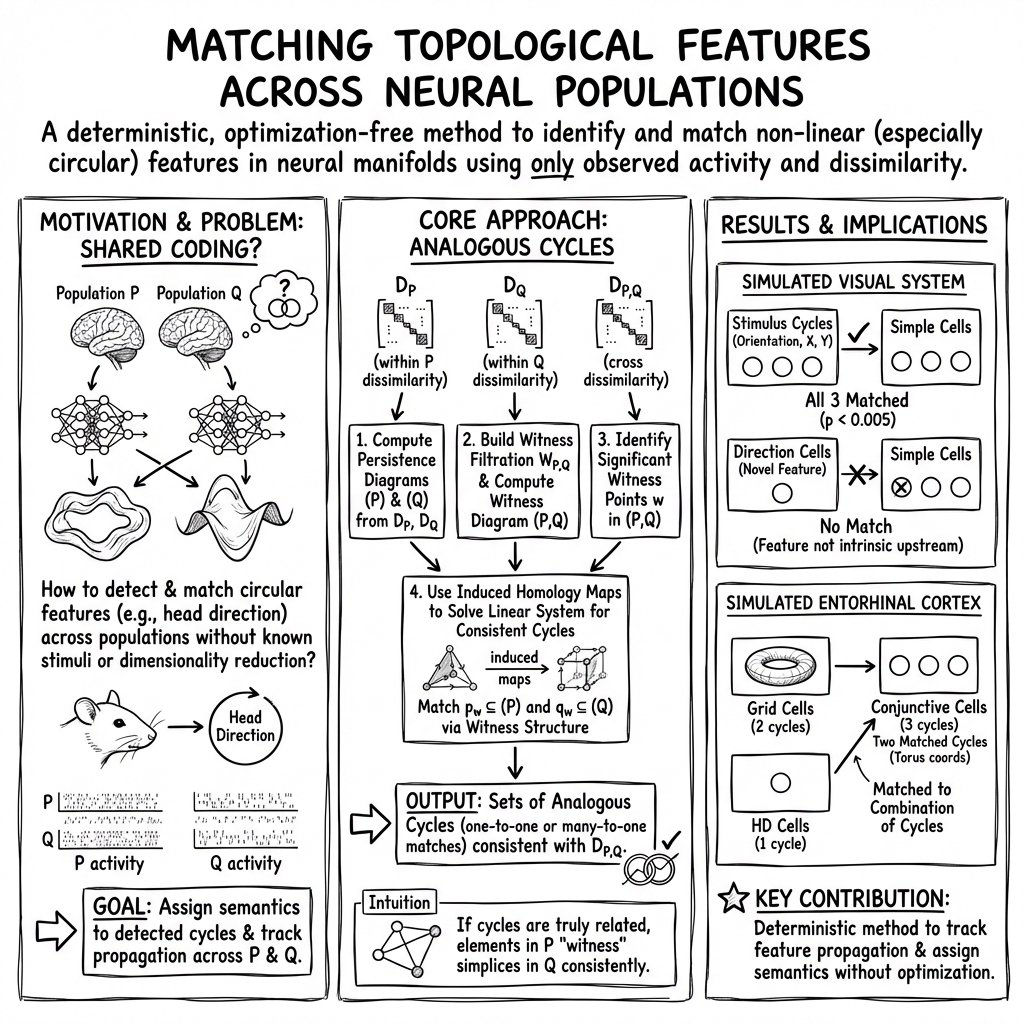
- The paper introduces a novel method of analogous cycles to track circular features in neural manifolds across populations.
- It applies persistent homology on dissimilarity matrices to match topological features, validated through simulations and in vivo data.
- The approach provides key insights into neural coding and paves the way for advanced computational neuroscience and AI models.
Exploring the Topology of Neural Manifolds and Analogous Cycles
Introduction
The paper "Tracking the topology of neural manifolds across populations" (2503.20629) introduces a comprehensive mathematical framework aimed at comparing the topological features of neural manifolds across distinct neural populations, particularly focusing on those with circular features. By leveraging the method of analogous cycles, the study provides a systematic approach to identify and match corresponding cycles within dissimilar neural manifolds without relying on common dimensionality reduction or optimization techniques.
Methodology: Analogous Cycles
The primary contribution of this study is the method of analogous cycles, which utilizes observed dissimilarity matrices to match topological features across neural populations. The process involves constructing persistence diagrams via the computation of persistent homology, which summarizes the topological features, specifically focusing on 1-dimensional cycles that correspond to circular structures.
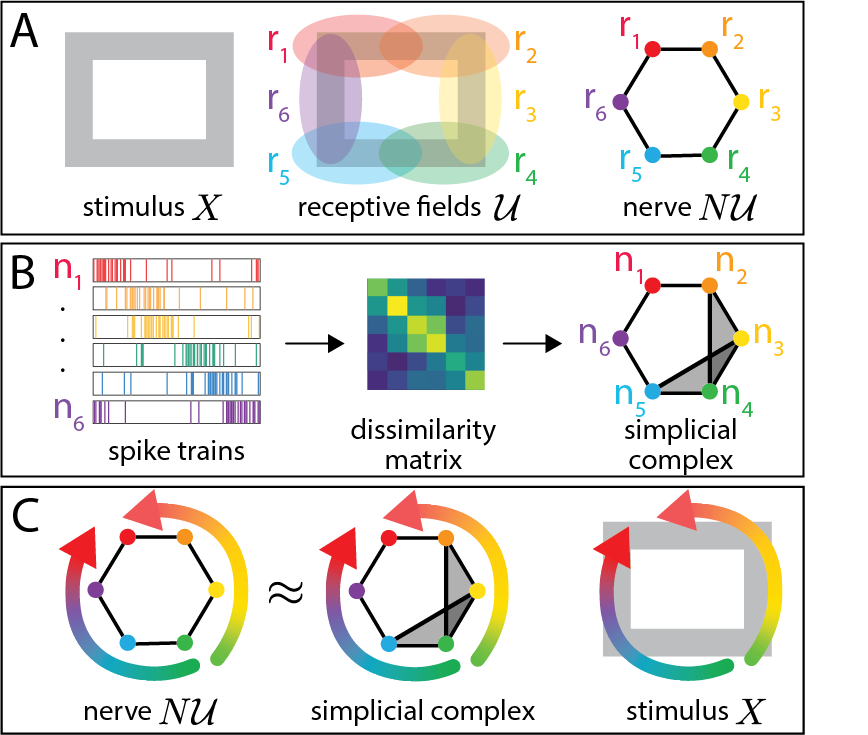

Figure 1: Illustration of the design of neural manifolds and persistent homology for circular features.
The method offers a robust mechanism for interpreting neural activity and extracting meaningful information by analyzing the interactions of receptive fields. It operates independently of the manifold's linearity and does not involve a reliance on external correlates, thereby providing an objective quantitative analysis grounded in mathematical principles.
Simulations and Validation
The study validates the analogous cycles method through a series of simulation experiments. It uses artificial neural systems simulating simple cells in visual cortex and head direction cells in the entorhinal cortex to test the framework’s ability to accurately track and match shared topological features. By constructing simulated populations with known topological features, the research verifies the accuracy of analogous cycles in identifying and matching these features across various layers of neural networks.
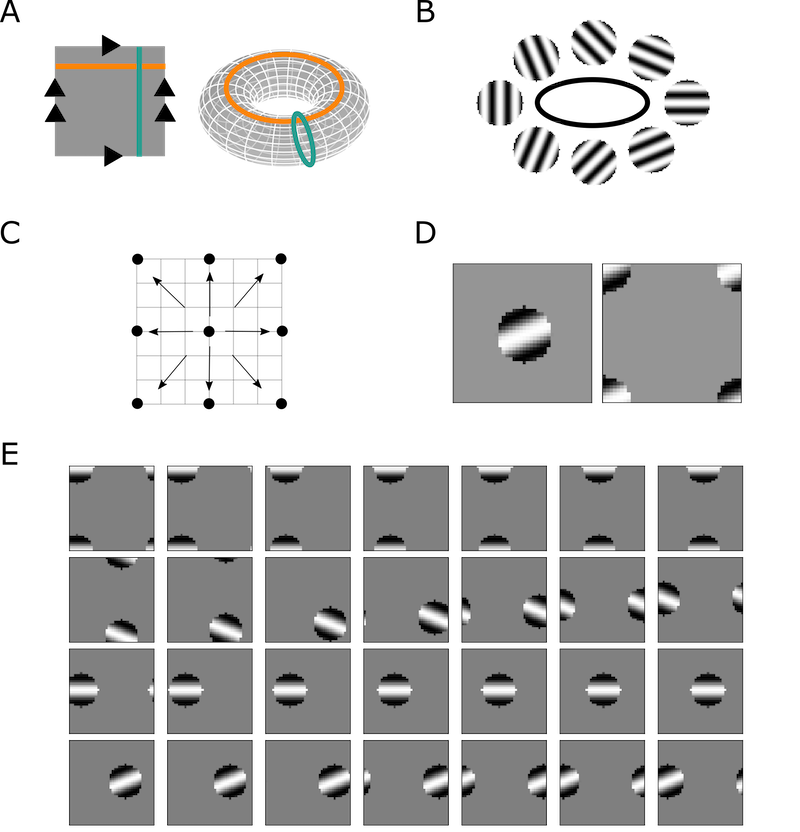
Figure 2: Results from simulation showing match of circular features between simple cells and derived neural populations.
Complex Interactions in Neural Manifolds
The study extends its validation by simulating more complex neural systems like conjunctive cells in the entorhinal cortex, highlighting the variety of non-linear geometrical relations that can exist in real neural systems. It explores how these non-linear manifolds represent stimuli and retain encoded information across different layers of neural population and successfully distinguishes between features that are intrinsic to upstream populations and those which are novel or synthesized through subsequent layers.
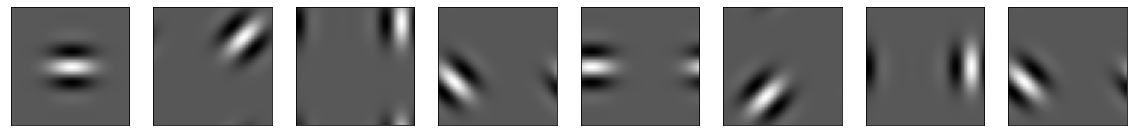

Figure 3: Visual representation of analogous cycles in the context of neural manifolds with multiple circular interactions.
In Vivo Application
Beyond simulations, the analogous cycles method is applied to in vivo data, particularly relating to mouse primary visual cortex (V1) and anterolateral (AL) visual areas. This real-world application demonstrates the framework’s practicality in identifying shared neural coding properties, even in instances of intricate, multi-directional neurological communication and interaction that extend beyond simulated models.
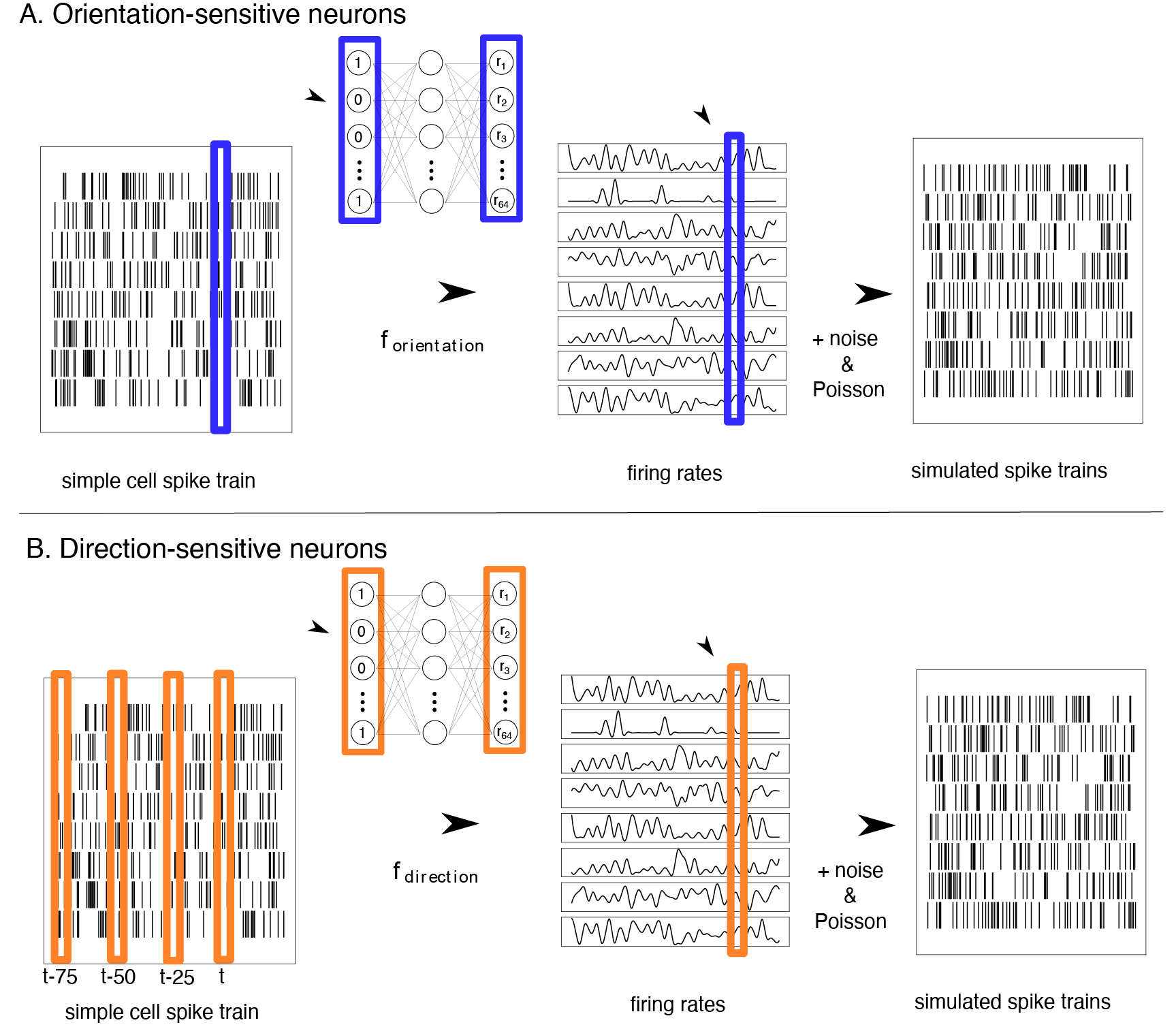
Figure 4: Application of analogous cycles on in vivo data, showing shared features between visual cortex regions.
Discussion and Implications
This study emphasizes that the method of analogous cycles is suitable for exploring and understanding complex neuronal population interactions where traditional methods may fall short. It offers an algebraically grounded framework for matching neural features across populations, facilitating a deeper understanding of how complex neural representations correspond to stimuli without relying on traditional signal correlation methods.
The implications of this research extend to multiple domains, including neural network analysis, the development of more sophisticated neural-manifold models, and broader applications in artificial intelligence that require the interpretation of high-dimensional data. This work paves the way for future investigations into the interplay of neural manifolds across populations, suggesting that neural encoding of information can be mathematically decomposed and interpreted in a manner that assists in the unfolding journeys into cognitive neuroscience and computational biology.
Conclusion
The method of analogous cycles as detailed in this paper provides a mathematically rigorous approach to compare and analyze the intrinsic topological structures within neuroscience. This approach has a significant impact on how researchers might explore neural data, offering a fresh perspective grounded in algebraic topology and providing robust mechanisms for understanding the continuity and evolution of neural encodings beyond observed data into theoretical realms of neuron-manifold interactions.