- The paper demonstrates that integrating CPC with SNNs improves both encoding accuracy and biological plausibility in self-supervised tasks.
- It replaces conventional CNN encoders with SNN-based encoders using LIF models and STDP rules to generate robust spike-based representations for classification and autoencoding.
- The integrated model achieves validation accuracies above 95% on MNIST, highlighting its potential for neuromorphic hardware deployment.
Integration of Contrastive Predictive Coding and Spiking Neural Networks
Introduction
The paper presents a systematic investigation into the integration of Contrastive Predictive Coding (CPC) with Spiking Neural Networks (SNNs), aiming to enhance the biological plausibility of predictive coding models while leveraging the energy efficiency and temporal dynamics inherent to SNNs. CPC, a self-supervised learning paradigm, is designed to learn representations that capture the predictive structure of sequential data via contrastive loss. SNNs, in contrast, process information through discrete spike events, closely mimicking the computational mechanisms of biological neurons. The study addresses the gap in prior research by directly combining CPC and SNNs, evaluating the encoding capacity of SNNs trained for classification and autoencoding tasks, and benchmarking their integration on a sequentially paired MNIST dataset.
Model Architecture and Encoding Strategies
The proposed architecture replaces the conventional CNN encoder in CPC with an SNN-based encoder. Two SNN variants are evaluated:
- SNN-Classifier: Trained for MNIST digit classification using the leaky integrate-and-fire (LIF) model and STDP learning rules. Input images are converted to spike trains via Poisson encoding, and spike counts per neuron are aggregated into 400-dimensional vectors.
- SNN-Autoencoder: Trained explicitly for encoding, also based on the LIF model, but optimized to minimize reconstruction error over 25 time steps. The membrane potential at the final time step serves as the latent representation.
Both encoders are integrated with a CPC network, where a GRU serves as the autoregressive component. The CPC network is trained to maximize the dot product similarity between predicted and true encoded vectors for positive (sequential) samples and minimize it for negative (non-sequential) samples. The output logits are normalized via a sigmoid activation, and binary cross-entropy loss is used for optimization.
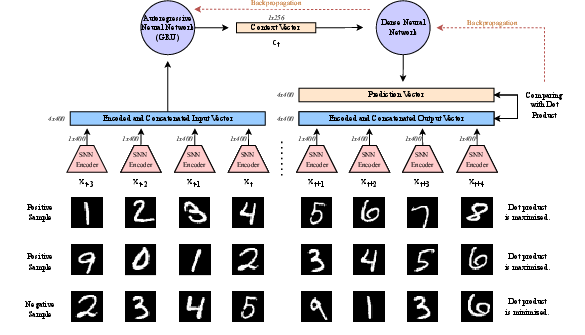
Figure 1: Structure of the CPC and SNN integration model, illustrating the flow from sequential input pairs to positive/negative sample classification.
Encoding Analysis and Biological Plausibility
The SNN-Classifier, despite being trained for classification, demonstrates the ability to generate distinct spike-based encodings for different digits. Within-class encodings exhibit high intra-class similarity, supporting their utility as input representations for CPC.


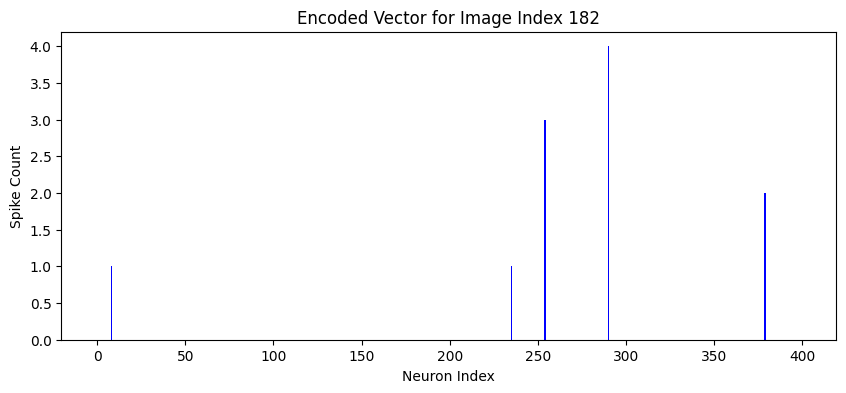
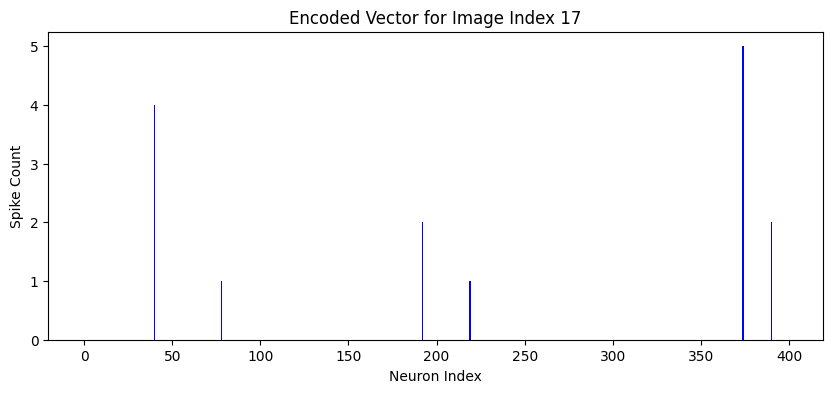
Figure 2: Visualization of 400-dimensional spike count vectors generated by the SNN for digits 5 and 7, highlighting class-specific encoding patterns.
The SNN-Autoencoder, while less biologically realistic due to its reliance on convolutional layers, achieves higher encoding fidelity. The trade-off between biological plausibility and encoding performance is evident: the SNN-Classifier aligns more closely with neurobiological principles, whereas the SNN-Autoencoder offers superior representational capacity.
Experiments are conducted on two MNIST subsets (2500 and 5000 samples), with both SNN-Classifier and SNN-Autoencoder serving as encoders. The CPC network is trained using batches of positive and negative pairs, with early stopping and adaptive learning rate scheduling.
Validation accuracy is tracked across epochs for each encoding strategy:
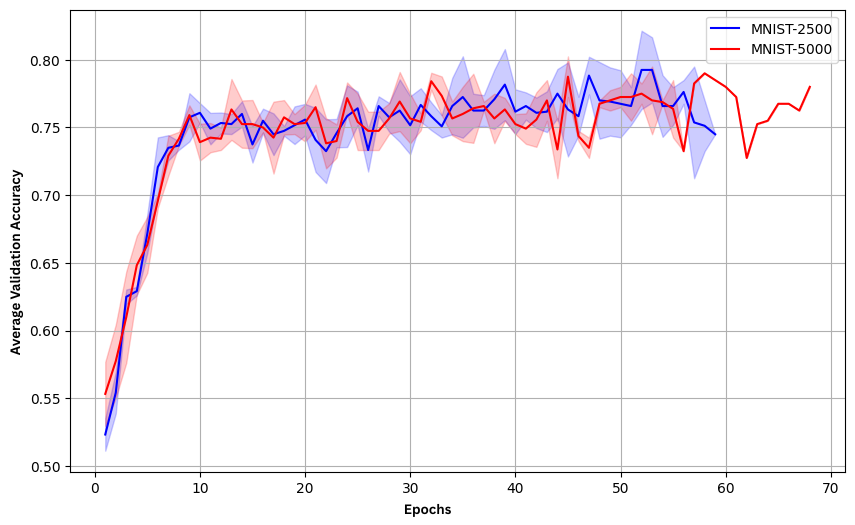
Figure 3: Validation accuracy of the CPC network using SNN-Classifier for encoding, demonstrating moderate performance.
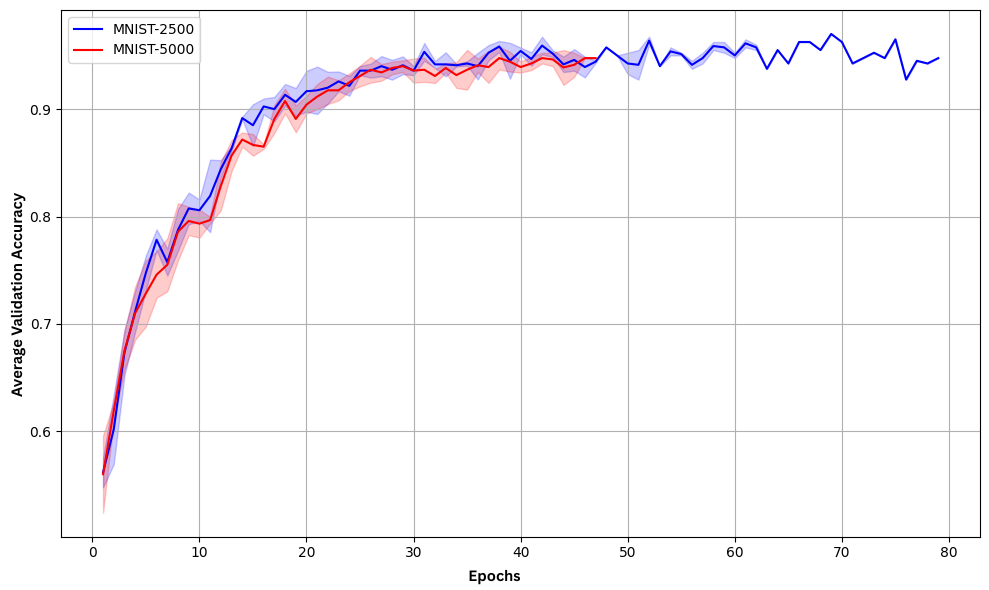
Figure 4: Validation accuracy of the CPC network using SNN-Autoencoder for encoding, showing superior performance and faster convergence.
A control experiment with random encoding yields near-chance accuracy, confirming the necessity of structured spike-based representations.
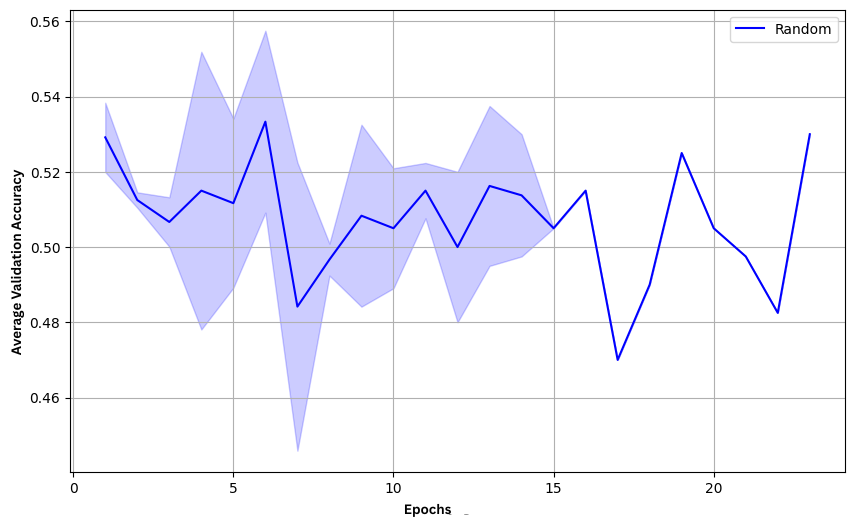
Figure 5: Validation accuracy of the CPC network with random encoding, illustrating baseline performance at chance level.
Key numerical results:
| Dataset |
Encoding Method |
Max Validation Accuracy |
| MNIST-2500 |
SNN-Autoencoder |
0.9683 |
| MNIST-5000 |
SNN-Autoencoder |
0.9583 |
| MNIST-2500 |
SNN-Classifier |
0.8033 |
| MNIST-5000 |
SNN-Classifier |
0.7992 |
| MNIST-2500 |
Random |
0.5558 |
The SNN-Autoencoder consistently outperforms the SNN-Classifier, achieving validation accuracies above 95%. The SNN-Classifier, while less performant, still demonstrates substantial encoding capacity, validating its use in biologically inspired CPC models. Increasing dataset size does not yield significant accuracy improvements but accelerates convergence.
Theoretical and Practical Implications
The integration of CPC and SNNs advances the development of predictive coding models with enhanced biological realism and energy efficiency. The demonstrated ability of SNNs trained for classification to serve as effective encoders in CPC frameworks suggests that spike-based representations are sufficiently expressive for self-supervised learning tasks. The results also highlight the trade-off between biological plausibility and representational power, with convolutional SNN-Autoencoders offering higher accuracy at the expense of neurobiological fidelity.
From a practical perspective, the model is well-suited for deployment on neuromorphic hardware, where spike-based computation can yield substantial energy savings. The approach is extensible to other time-dependent datasets, such as human action recognition, and can be adapted to more biologically realistic predictive coding architectures.
Future Directions
Potential avenues for future research include:
- Application of the integrated CPC-SNN model to complex, temporally structured datasets beyond MNIST.
- Exploration of alternative predictive coding frameworks with greater biological fidelity.
- Implementation and benchmarking on neuromorphic hardware platforms.
- Investigation of hybrid architectures that balance biological plausibility and encoding performance.
Conclusion
The paper provides a rigorous evaluation of CPC-SNN integration, demonstrating that SNNs—whether trained for classification or encoding—can serve as effective encoders in contrastive predictive coding frameworks. The SNN-Autoencoder achieves the highest classification accuracy, while the SNN-Classifier offers a biologically plausible alternative with reasonable performance. The findings support the feasibility of biologically inspired, energy-efficient predictive coding models and lay the groundwork for future research in spike-based self-supervised learning and neuromorphic computing.









