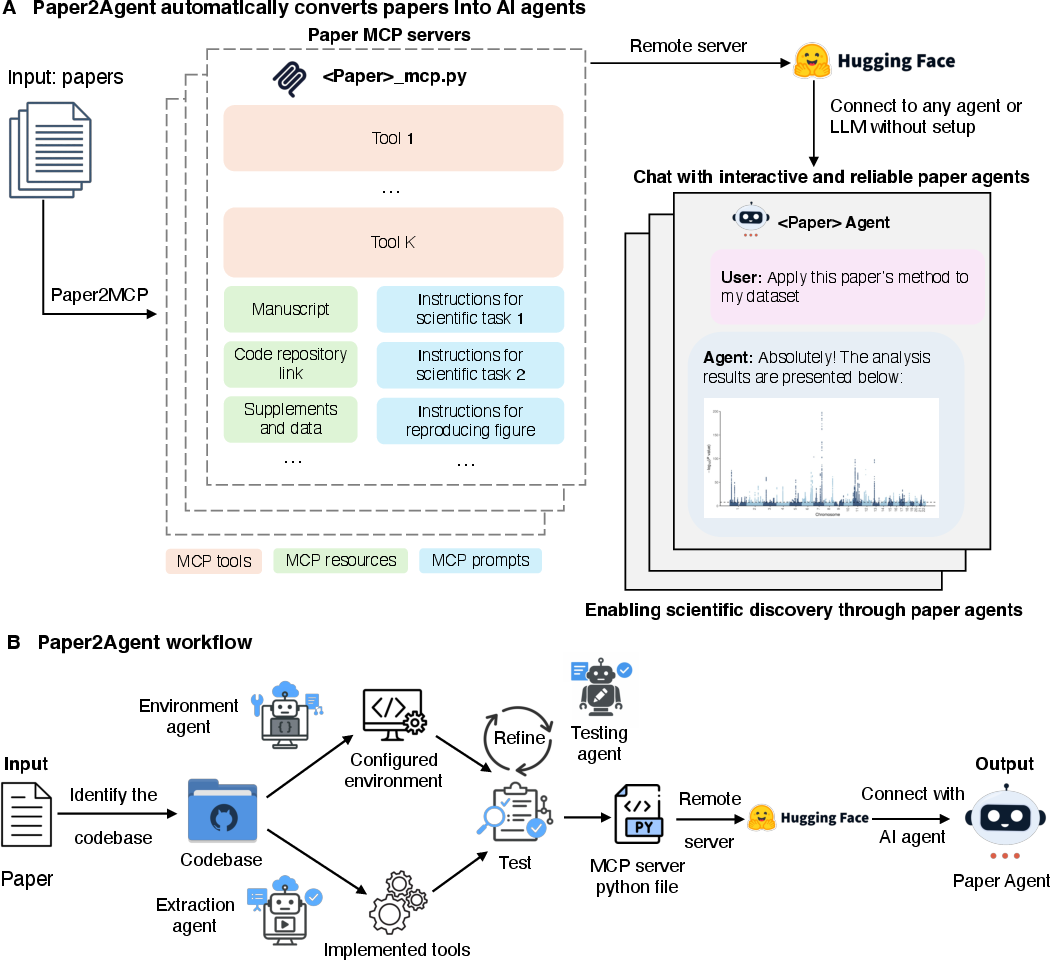
Paper2Agent: Reimagining Research Papers As Interactive and Reliable AI Agents
Abstract: We introduce Paper2Agent, an automated framework that converts research papers into AI agents. Paper2Agent transforms research output from passive artifacts into active systems that can accelerate downstream use, adoption, and discovery. Conventional research papers require readers to invest substantial effort to understand and adapt a paper's code, data, and methods to their own work, creating barriers to dissemination and reuse. Paper2Agent addresses this challenge by automatically converting a paper into an AI agent that acts as a knowledgeable research assistant. It systematically analyzes the paper and the associated codebase using multiple agents to construct a Model Context Protocol (MCP) server, then iteratively generates and runs tests to refine and robustify the resulting MCP. These paper MCPs can then be flexibly connected to a chat agent (e.g. Claude Code) to carry out complex scientific queries through natural language while invoking tools and workflows from the original paper. We demonstrate Paper2Agent's effectiveness in creating reliable and capable paper agents through in-depth case studies. Paper2Agent created an agent that leverages AlphaGenome to interpret genomic variants and agents based on ScanPy and TISSUE to carry out single-cell and spatial transcriptomics analyses. We validate that these paper agents can reproduce the original paper's results and can correctly carry out novel user queries. By turning static papers into dynamic, interactive AI agents, Paper2Agent introduces a new paradigm for knowledge dissemination and a foundation for the collaborative ecosystem of AI co-scientists.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
A simple explanation of “Reimagining Research Papers As Interactive and Reliable AI Agents”
1. What is this paper about?
This paper introduces Paper2Agent, a system that turns scientific papers into helpful AI “assistants.” Instead of a paper just sitting there for you to read, Paper2Agent creates an interactive agent that can explain the paper, run its code, and apply its methods to new data—all through simple chat-style questions. The goal is to make research easier to use, more reliable, and faster to build on.
2. What questions are the researchers asking?
The paper looks at a few big questions in everyday terms:
- How can we turn a hard-to-use research paper (with complicated code and setup) into something you can talk to and use right away?
- Can an AI assistant safely and accurately run a paper’s methods, not just make up code?
- Will this approach help people reproduce results from the paper and use the method on new data?
- Can this work across very different science topics, like genetics and single-cell biology?
3. How did they do it?
Think of a research paper like a cookbook full of recipes, but the kitchen isn’t set up, and the tools are scattered. Paper2Agent sets up the kitchen, organizes the tools, and gives you a smart helper who can cook any recipe you describe in plain language.
Here’s the approach, step by step:
- They find the paper’s official code and examples.
- They automatically set up the software “kitchen” (the environment) so everything runs the same way for everyone.
- They scan the tutorials and code to extract the key actions (like “clean this data” or “run this model”) and turn them into reusable functions called tools.
- They test these tools over and over until the results match what the paper reports.
- They package everything as an MCP server and connect it to a chat-style AI agent you can talk to.
What is MCP?
- MCP (Model Context Protocol) is like a universal plug that lets AI agents use tools and data in a standardized way. It defines how to send inputs in, get outputs back, and keep things consistent and safe.
What’s inside a paper’s MCP server?
- Tools: Small, reliable functions that do the paper’s main steps (like scoring a DNA variant or clustering cells).
- Resources: Organized files, links, and data from the paper (like datasets and figures).
- Prompts: Short instructions that describe multi-step workflows (like a checklist for a full analysis), so the agent runs steps in the right order.
They used Claude Code (an AI coding assistant) to do the heavy lifting and hosted some of the MCP servers online (so users don’t need to install anything).
4. What did they find?
They tested Paper2Agent on three real research projects and showed that the agents are accurate, useful, and easy to use.
AlphaGenome (genomics: predicting DNA variant effects)
- What they built: An agent with 22 tools that can score DNA changes across tissues, visualize results, and prioritize likely disease genes.
- Results: On 15 example questions from the original tutorial and 15 brand-new questions, the agent got 100% correct answers (checked against running the original code by hand).
- Why it matters: It can reproduce the paper’s results and handle new tasks without someone needing to learn the complicated code. It even re-examined a published result and offered an alternative gene explanation, showing how agents can help revisit scientific claims.
TISSUE (spatial transcriptomics: adding uncertainty to gene predictions)
- What they built: An agent that runs the TISSUE pipeline end-to-end and explains required inputs and outputs in simple terms.
- Results: It matched human-run results on real datasets and could automatically find and download relevant datasets using a structured registry.
- Why it matters: It doubles as both a teacher (Q&A about how to use the method) and a worker (it runs the full analysis reliably).
Scanpy (single-cell RNA-seq: preprocessing and clustering)
- What they built: An agent focused on the most common workflow—cleaning, normalizing, reducing dimensions, clustering, and labeling cells.
- Results: With just the path to a dataset, the agent reproduced human researcher outputs on several public datasets. They used MCP “prompts” to ensure the steps happen in the right order.
- Why it matters: It gives beginners an easy, accurate pipeline and ensures consistent, repeatable results.
5. Why is this important?
Turning papers into interactive AI agents has several big benefits:
- Lowers barriers: You can use advanced methods by asking questions in plain language, without wrestling with software installs and complex code.
- Improves reliability: Each tool is tested against the original results and “locked in,” which reduces mistakes from AI “making up” code.
- Boosts reproducibility: Anyone can rerun the same steps and get the same answers.
- Speeds up discovery: Scientists can go from reading about a method to using it on their own data in minutes.
- Encourages collaboration: In the future, different paper-agents could work together—mixing methods and datasets to do more complex tasks.
6. Limits and next steps
The system works best when the original paper’s code is complete, clean, and well-documented. If the code is broken or unclear, it’s hard to make a reliable agent. The authors suggest:
- Using agent-ready practices in publishing (like clear code, datasets, and workflows).
- Benchmarking how well each agent reproduces results and handles new tasks.
- Combining multiple related papers into a single, more powerful agent when that makes sense.
- Adding an “agent availability” section to papers, similar to “data availability,” so readers can try the agent right away.
7. The takeaway
Paper2Agent turns research papers from static text into active, trustworthy helpers you can talk to. It makes advanced science tools easier to use, checks that they work, and helps people apply them to new problems. If widely adopted, this could change how science is shared and done—moving from reading about methods to using them instantly, accurately, and collaboratively.
Knowledge Gaps
Knowledge gaps, limitations, and open questions
Below is a concise, actionable list of what remains missing, uncertain, or unexplored in the paper, to guide future research and development.
- Lack of large-scale evaluation across a broad corpus of papers: no statistics on success/failure rates, conversion coverage, or typical failure modes when agentifying hundreds/thousands of heterogeneous repositories.
- Limited benchmarking depth: “100% accuracy” is reported on small, curated query sets; no stress tests with noisy inputs, adversarial prompts, randomized seeds, or distribution shifts, and no quantitative metrics for figure/visualization fidelity (e.g., image similarity thresholds).
- No standardized evaluation framework: absence of formal metrics for agent reliability (e.g., tool-level pass rates, workflow-level reproducibility, latency, determinism), generalization (to novel queries/datasets), and end-to-end task success.
- Dependence on tutorials for tool extraction: unclear how the system performs when repositories lack clear, high-quality tutorials or runnable examples; no methodology for extracting tools from narrative-only or poorly documented code.
- Handling papers without public or complete codebases is unspecified: workflows to infer tools from the manuscript text, generate missing code, and validate outputs against the paper’s claims are not developed or rigorously evaluated.
- Cross-language/codebase support is unaddressed: no strategy for wrapping R/Matlab/Julia/C++ repositories, mixed-language stacks, or complex build systems requiring compiled extensions.
- Environment and hardware variability: reproducibility under different OSes, CUDA/GPU versions, TPUs, HPC schedulers, and specialized drivers is not tested; no guidance on reproducible GPU builds or deterministic inference.
- Compute scalability and cost: missing analysis of runtime, memory, throughput, and cloud cost for large datasets or heavy models; no autoscaling, caching, or resource quota policies for multi-user MCP servers.
- Maintenance and versioning: no policy for pinning, updating, and deprecating MCP servers when upstream repos or dependencies change; lack of CI/CD pipelines for automatic rebuilds, regression tests, and semantic versioning.
- Security model for remote execution is unspecified: sandboxing, code execution isolation, least-privilege permissions, dependency supply-chain security, and protection against arbitrary code execution are not detailed.
- Secrets and API key handling: risk of credential leakage in prompts, logs, or shared sessions is not addressed; no secure secret management, rotation, or client-side encryption guidance.
- Data privacy and regulatory compliance: no procedures for handling sensitive user data (e.g., clinical genomics) under HIPAA/GDPR; policies for data minimization, PII redaction, and secure storage/transfer are missing.
- Licensing and IP compliance: unclear how third-party code/data licenses (e.g., GPL/BSD/Apache, data EULAs) propagate through MCP servers; no automated license auditing or attribution/citation enforcement.
- Validation of MCP prompt generation: automatically inferred workflow prompts may encode errors or hidden assumptions; no formal verification, differential testing, or expert review of prompt fidelity across domains.
- Orchestration across multiple MCPs: no conflict resolution, tool selection policy, ontology reconciliation, or schema harmonization when combining tools from different papers in a single session.
- Provenance, auditability, and reproducibility over time: limited discussion of standardized, exportable execution traces (inputs, versions, parameters, seeds), data lineage, and immutable artifact storage for long-term reproducibility.
- Uncertainty and calibration: outputs are presented without systematic uncertainty estimation or calibration beyond specific methods (e.g., TISSUE); no general framework to quantify and communicate confidence in agent-generated analyses.
- Error handling and recovery: no specification of graceful degradation, retries, rollbacks, or safe fallbacks when tools fail mid-pipeline or when partial outputs may mislead users.
- Robust testing beyond tutorials: absence of unit tests, integration tests, fuzz/property-based tests, and coverage metrics for generated tools; no red-team evaluations for prompt-injection or tool-misuse vulnerabilities.
- User studies and human factors: no empirical assessment of usability, learning curve, trust, and error comprehension among non-experts; lack of design guidelines for effective human-in-the-loop oversight.
- Generalization beyond computational biology: applicability to domains requiring hardware interaction (robotics), simulators, real-time constraints, or proprietary ecosystems remains untested.
- End-to-end training workflows: framework focuses on inference and analysis; support for reproducible model training, hyperparameter sweeps, large-scale data preprocessing, and checkpoint management is not addressed.
- Large data handling: no strategy for streaming, chunked processing, out-of-core computation, or distributed execution; impact of dataset size on reliability and performance is unquantified.
- Discoverability and standards: no metadata schema or registry for “agent availability,” indexing, search, or interoperability standards to enable users/journals to find, compare, and cite agents.
- Governance when agents contradict paper conclusions: no review, sign-off, or disclosure protocols for agent-generated reinterpretations that differ from published claims; no guidance on responsible communication.
- Bias, fairness, and equity: no audits for demographic or domain biases inherited from models/datasets; lack of bias mitigation techniques and transparency reporting for agent outputs.
- Offline and constrained environments: no pathway for air-gapped or low-resource deployments (e.g., hospitals, field labs); dependency mirrors, model artifact pinning, and offline LLM options are not discussed.
- Multi-tenancy and concurrency: unaddressed issues of user isolation, quota enforcement, request prioritization, and denial-of-service protections on shared MCP servers.
- Image/figure equivalence: no standardized procedures or thresholds for validating regenerated figures (e.g., structural similarity, pixel-wise tolerances, statistical equivalence of plotted data).
- Internationalization: no support for multilingual papers, code comments, or user prompts; cross-lingual extraction of tools and resources is unexplored.
- Community and sustainability: no contribution, review, and moderation workflows for community-submitted MCP servers; unclear funding and maintenance model for long-term hosting and compute costs.
- Quantifying “agentifiability”: the proposed idea lacks a formal rubric or automated scoring pipeline linking repository quality (docs, tests, modularity) to conversion success, reproducibility, and maintenance burden.
- Ethical boundaries of autonomy: policies for what actions agents may autonomously take (e.g., downloading large datasets, contacting external APIs, writing to user storage) are not defined; permissioning and consent controls are absent.
- Ontology/version drift: reliance on external ontologies (e.g., UBERON, CL) and metadata can introduce silent errors as those resources evolve; no monitoring and update strategies to prevent semantic mismatches over time.
Practical Applications
Overview
Paper2Agent turns static research papers and their codebases into interactive, reliable AI agents by wrapping core methods as Model Context Protocol (MCP) tools, bundling associated resources, encoding workflows as prompts, validating outputs against reference results, and deploying remotely (e.g., Hugging Face Spaces). The framework reduces adoption friction, raises reproducibility, and enables natural-language execution of complex methods. Below are practical, real-world applications derived from its findings, methods, and innovations, grouped by immediacy and sector, with dependencies noted.
Immediate Applications
The following applications are deployable now using the released framework, MCP servers, and demonstrated case studies (AlphaGenome, TISSUE, Scanpy).
Industry
- Variant interpretation copilot for R&D
- Sector(s): Biotech, Pharma, Clinical Genomics (research use)
- Use case: Prioritize causal variants/genes in GWAS loci, interpret predicted regulatory effects across modalities, generate figures/reports (e.g., LDL SORT1/CELSR2/PSRC1 example).
- Tools/products/workflows: AlphaGenome MCP agent embedded in LIMS/ELN; “one-prompt” variant-to-report workflow; API endpoints for batch scoring.
- Assumptions/dependencies: Valid API keys/quotas; data governance/IRB for human data; compute capacity; method license; non-diagnostic usage labeling.
- Contract bioinformatics delivery with standardized outputs
- Sector(s): CROs, Core Facilities
- Use case: Offer reproducible single-cell preprocessing/clustering (Scanpy agent) and uncertainty-aware ST analyses (TISSUE agent) with consistent, audited outputs.
- Tools/products/workflows: Agent-backed service templates; auto-generated QC reports; CI logs showing tool and dataset hashes.
- Assumptions/dependencies: Sufficient input metadata; containerized execution; dataset size fits available hardware; customer approvals for pipeline defaults.
- Agent marketplace integrations
- Sector(s): AI platforms, Developer tooling
- Use case: Add “Run the paper” buttons to code/model hubs; bundle MCP servers as installable extensions in IDEs or chat-based coding agents.
- Tools/products/workflows: Paper MCP registry; agent connectors for Claude Code, OpenAI, local LLMs; MCP discovery APIs.
- Assumptions/dependencies: MCP compatibility; rate limiting; sandboxing; per-paper licensing and attribution.
- Figure regeneration and method demos embedded in publications
- Sector(s): Scientific publishing platforms, Preprint servers
- Use case: Readers click “Recreate Figure X” or “Apply to your data” next to a paper; demos running in hosted spaces.
- Tools/products/workflows: Hosted MCP servers per paper; “Try this method” interactive panels; auto-provisioned environments.
- Assumptions/dependencies: Publisher-hosted compute; author-provided code and minimal example data; content moderation.
Academia
- Agent-enabled reproducibility checks in peer review
- Sector(s): Journals, Conferences, Program Committees
- Use case: Reviewers run agentized tests that reproduce tutorial results, verify numerical outputs/figures, and flag non-runnable tools.
- Tools/products/workflows: “Agent-enabled review” plugin; standardized logs; pass/fail badges in submission portals.
- Assumptions/dependencies: Review-time compute; permissive code/data licenses; dataset stubs for tests; clear timeouts.
- “Living methods” for labs and consortia
- Sector(s): Research labs, Consortia
- Use case: Convert lab methods into reusable MCP tools; onboard new members; scale method reuse across projects/sites.
- Tools/products/workflows: Internal MCP hub; method prompts encoding canonical workflows; versioned agent releases with changelogs.
- Assumptions/dependencies: Code modularity; docstrings/examples; data availability or synthetic fixtures; maintainer bandwidth.
- Teaching and training with interactive agents
- Sector(s): Higher education, Professional training
- Use case: Assignments and workshops where students run Scanpy/TISSUE pipelines, query method inputs/assumptions, and compare outputs to ground truth.
- Tools/products/workflows: Classroom Spaces; graded notebooks auto-calling MCP tools; rubric tied to agent outputs.
- Assumptions/dependencies: Stable hosting during class; small example datasets; institutional sign-ins; rate limits.
- Rapid reanalysis of public datasets
- Sector(s): Bioinformatics, Systems biology
- Use case: Pull ST datasets via structured MCP resources (e.g., Zenodo), apply pipelines, and export reports without manual setup.
- Tools/products/workflows: Dataset registries in MCP resources; “data-to-report” prompts; provenance stamps (inputs, hashes).
- Assumptions/dependencies: Dataset URLs live and accessible; schema consistency; controlled vocabularies (ONT, UBERON, CL).
Policy and Funding
- Pilot “agent availability” alongside code/data availability
- Sector(s): Journals, Funders
- Use case: Encourage or require an MCP server and minimal tests for computational papers to enhance reuse.
- Tools/products/workflows: Submission checklist item; MCP manifest archive; DOI linking to hosted agent/demo.
- Assumptions/dependencies: Exceptions for sensitive/regulated data; hosting credits; template tests.
- Lightweight, metrics-based reproducibility audits
- Sector(s): Funders, Research integrity offices
- Use case: Track “agentification score” (e.g., tools passed/total, coverage of figures) in grant reports and portfolios.
- Tools/products/workflows: Audit dashboards; automated CI using Paper2Agent; periodic snapshots.
- Assumptions/dependencies: Community benchmarks; shared schema for metrics; opt-in from PIs.
Daily Life and Science Communication
- Citizen-science exploration of open data (non-clinical)
- Sector(s): Public, Science journalism
- Use case: Run published pipelines on open datasets, generate interpretable summaries/figures, and understand method assumptions via Q&A.
- Tools/products/workflows: Browser-based demos with “explain this step” prompts; shareable reports.
- Assumptions/dependencies: Strong disclaimers (not for diagnosis); curated, safe example datasets; usage caps.
- Library and research office support
- Sector(s): University libraries, Research support services
- Use case: Staff use agents to answer “what inputs are required?” and to triage environment/setup issues for researchers.
- Tools/products/workflows: Knowledge-base integration; common-issue playbooks distilled from agent logs.
- Assumptions/dependencies: Staff access to hosted agents; institutional SSO; data privacy policies.
Long-Term Applications
These require further validation, scaling, standardization, governance, or regulatory approvals.
Industry
- Regulatory-grade genomic decision support
- Sector(s): Healthcare, Diagnostics
- Use case: Integrate validated, agentified genomic models into EMRs for variant interpretation and report generation in clinical workflows.
- Tools/products/workflows: FDA-cleared pipelines; locked MCP versions; audit trails and performance monitoring.
- Assumptions/dependencies: Clinical validation; post-market surveillance; PHI-safe deployments; model drift controls.
- Autonomous multi-agent R&D pipelines
- Sector(s): Pharma, Synthetic biology, Materials science
- Use case: End-to-end agent teams that plan experiments, run coupled method MCPs, interpret results, and propose next steps.
- Tools/products/workflows: Cross-agent orchestration; lab automation APIs; experiment schedulers; data/metadata harmonization.
- Assumptions/dependencies: Reliable tool composition; safety rails; cost-effective compute; IP tracking.
- Cross-paper ensemble modeling and continuous synthesis
- Sector(s): AI/ML platforms, Model hubs
- Use case: Agents coordinate across multiple paper MCPs to assemble “best-of-breed” ensembles and living benchmarks.
- Tools/products/workflows: Inter-agent protocols; standardized metrics; auto-curated leaderboards.
- Assumptions/dependencies: Compatible licenses; unified interfaces; reproducible version pinning.
- Commercial agent marketplaces and licensing
- Sector(s): Software, IP management
- Use case: Monetize agentized methods with usage-based pricing, attribution, and license compliance tracking.
- Tools/products/workflows: Billing, metering; license policy engines; revenue sharing with authors/institutions.
- Assumptions/dependencies: Clear IP ownership; standardized licenses for MCP artifacts; dispute resolution.
Academia
- Field-wide MCP registries and “research-as-agents” infrastructure
- Sector(s): Research infrastructure, Consortia
- Use case: Curated registries per domain (e.g., single-cell, proteomics) with quality ratings, tests, and interoperability guarantees.
- Tools/products/workflows: Community governance; metadata standards; artifact signing.
- Assumptions/dependencies: Funding for maintainers; consensus on schemas; persistent hosting.
- Agent-driven living reviews and meta-analyses
- Sector(s): Methodology, Evidence synthesis
- Use case: Agents re-run analyses as new datasets/papers appear, updating figures/conclusions with provenance and uncertainty quantification.
- Tools/products/workflows: Watchers for new releases; change-detection alerts; versioned review artifacts.
- Assumptions/dependencies: Robust evaluation pipelines; bias detection; community oversight.
- Automated benchmarking and LLM-as-judge evaluation
- Sector(s): ML research, Computational sciences
- Use case: Continuous, agent-run testbeds that score new methods across standardized tasks using human-in-the-loop and LLM-based assessments.
- Tools/products/workflows: Benchmark suites; adjudication protocols; drift and regression testing.
- Assumptions/dependencies: Validation of LLM judges; gold-standard datasets; contestable scoring.
Policy and Funding
- Standards for provenance, auditability, and safety of research agents
- Sector(s): Standards bodies, Regulators, Funders
- Use case: Require cryptographic signing, immutable logs, reproducibility badges, and safety checklists for published agents.
- Tools/products/workflows: Provenance schemas (e.g., W3C PROV extensions); certification programs; compliance tooling.
- Assumptions/dependencies: Community adoption; cross-publisher alignment; enforcement mechanisms.
- National or regional hosting for public-interest research agents
- Sector(s): Public sector, e-Infrastructure
- Use case: Sustainably host high-value agents (e.g., pandemic modeling, climate analyses) with SLAs and equitable access.
- Tools/products/workflows: Federated hosting; data access controls; cost-sharing models.
- Assumptions/dependencies: Budget commitments; data-sharing agreements; cybersecurity posture.
- Privacy, security, and dual-use governance for agentic science
- Sector(s): Regulators, Ethics boards
- Use case: Policies for sensitive data handling, export controls, and misuse mitigation when agents compose tools autonomously.
- Tools/products/workflows: Risk assessments; gated capabilities; red-teaming protocols.
- Assumptions/dependencies: Ongoing threat modeling; alignment with existing frameworks (HIPAA, GDPR, ITAR).
Daily Life and Science Communication
- Personal research companions linked to public science agents
- Sector(s): Education technology, Lifelong learning
- Use case: Learners build custom curricula and hands-on projects by orchestrating trusted paper agents across domains.
- Tools/products/workflows: Curriculum builders; scaffolded prompts; skill assessments tied to agent outputs.
- Assumptions/dependencies: Curation for quality/safety; age-appropriate defaults; affordability.
- Agent-assisted science journalism and fact-checking
- Sector(s): Media
- Use case: Rapidly reproduce key analyses/figures from papers, test alternative hypotheses, and surface method caveats.
- Tools/products/workflows: Newsroom dashboards; one-click “recreate figure” workflows; uncertainty explainers.
- Assumptions/dependencies: Editorial training; access to hosted agents; strict disclaimers on preliminary results.
Notes on Feasibility and Dependencies (Cross-Cutting)
- Code and data quality: Agentification depends on accessible, runnable, and licensed codebases plus minimally sufficient example data.
- Compute and cost: Hosting and running MCP tools require provisioned CPU/GPU, storage, and budgeting for LLM/API calls.
- Licensing and IP: Clear licenses for code, models, and datasets; policies for redistribution via MCP servers.
- Security and privacy: Sandboxing, secrets management (API keys), PHI/PII safeguards, and audit logs are essential for sensitive use.
- Standardization: Broad uptake of MCP and shared schemas for inputs/outputs/resources improves interoperability and reuse.
- Human oversight: Even with high reproducibility, expert review remains necessary for high-stakes decisions (e.g., clinical use).
These applications leverage the core innovations demonstrated by Paper2Agent: automated tool extraction and validation from papers, MCP-based standardization, hosted and composable agents, and workflow prompts that encode best practices for reproducible scientific analysis.
Collections
Sign up for free to add this paper to one or more collections.