Hunt Globally: Wide Search AI Agents for Drug Asset Scouting in Investing, Business Development, and Competitive Intelligence
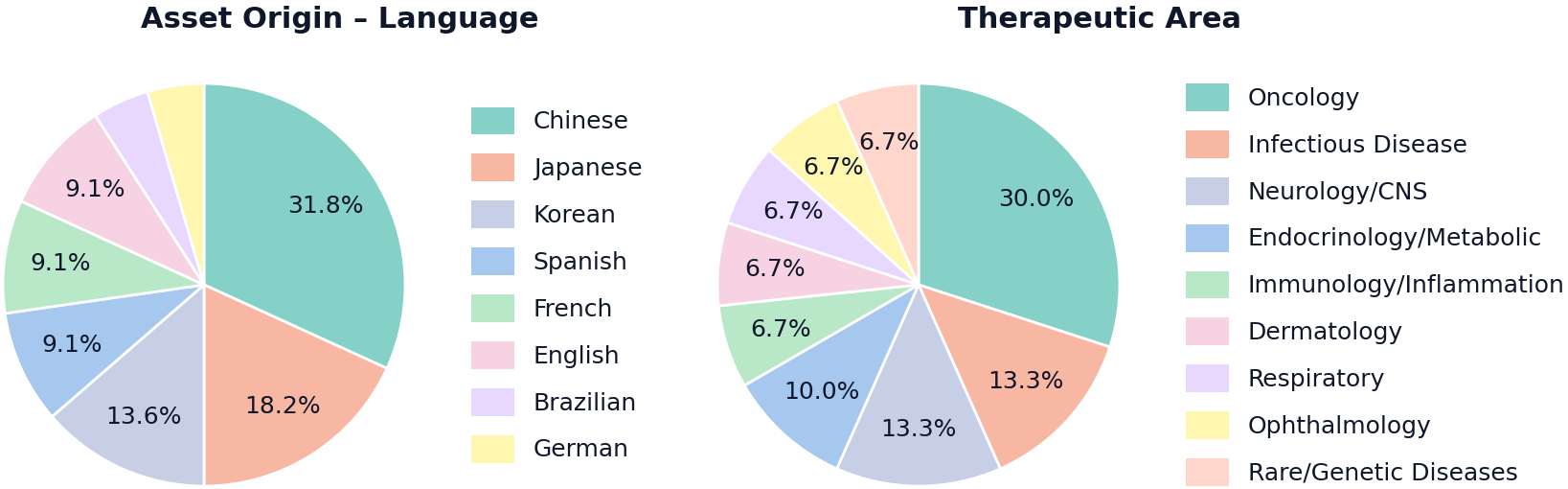
Abstract: Bio-pharmaceutical innovation has shifted: many new drug assets now originate outside the United States and are disclosed primarily via regional, non-English channels. Recent data suggests that over 85% of patent filings originate outside the U.S., with China accounting for nearly half of the global total. A growing share of scholarly output is also non-U.S. Industry estimates put China at 30% of global drug development, spanning 1,200+ novel candidates. In this high-stakes environment, failing to surface "under-the-radar" assets creates multi-billion-dollar risk for investors and business development teams, making asset scouting a coverage-critical competition where speed and completeness drive value. Yet today's Deep Research AI agents still lag human experts in achieving high recall discovery across heterogeneous, multilingual sources without hallucination. We propose a benchmarking methodology for drug asset scouting and a tuned, tree-based self-learning Bioptic Agent aimed at complete, non-hallucinated scouting. We construct a challenging completeness benchmark using a multilingual multi-agent pipeline: complex user queries paired with ground-truth assets that are largely outside U.S.-centric radar. To reflect real-deal complexity, we collected screening queries from expert investors, BD, and VC professionals and used them as priors to conditionally generate benchmark queries. For grading, we use LLM-as-judge evaluation calibrated to expert opinions. On this benchmark, our Bioptic Agent achieves 79.7% F1 score, outperforming Claude Opus 4.6 (56.2%), Gemini 3 Pro + Deep Research (50.6%), OpenAI GPT-5.2 Pro (46.6%), Perplexity Deep Research (44.2%), and Exa Websets (26.9%). Performance improves steeply with additional compute, supporting the view that more compute yields better results.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Explain it Like I'm 14
Overview
This paper is about building smarter AI helpers that can search the world for new drug projects (“assets”)—especially ones hidden in non-English sources—to help investors and business teams find valuable opportunities fast and without missing anything important. The authors introduce a way to fairly test how well such AI agents can find “all the right drugs,” and they present a new AI system, the Bioptic Agent, that does this wide, careful search better than other popular tools.
Key Objectives
The paper set out to answer three simple questions:
- Can we create a fair, tough test to see if AI can find drug projects from around the world, not just English-language or U.S.-centric sources?
- Can an AI agent be designed to find a complete set of matching drug assets for complex, realistic investor-style queries, without making things up?
- How does this new agent compare to leading AI research tools in terms of accuracy and completeness?
How They Did It (Methods, in Plain Language)
Think of this like a global treasure hunt where the clues are scattered across many countries and languages.
- Building a tough benchmark (the test):
- The team first collected real drug assets from local news sources in different regions and languages (like Chinese, Japanese, Korean, French, Spanish, etc.). This helps avoid bias toward English-only content.
- They cleaned and organized each asset with an “Attributes Enrichment Agent” that double-checks facts, resolves different names or aliases, and adds details like the drug’s stage, target, developer, and trials.
- A “Google Search Agent” compared how discoverable each asset was in English vs. its original language to favor “under-the-radar” items less likely to show up with easy English searches.
- They then generated realistic queries (search questions) based on real investor and business development requests. Importantly, they hid direct identifiers (like the drug’s exact name or code) so the AI had to reason from clues rather than just match words.
- An automated validator agent and human experts checked that each query matched its ground-truth asset and felt like a real investor question.
- Designing the Bioptic Agent (the AI scout):
- Investigator Agents: These are like multilingual explorers that search the web in parallel (e.g., English and Chinese) to find candidate assets that might match the query.
- Criteria Match Validator Agent: A “referee” that checks each candidate against the query’s rules, pulling evidence and links to prove the match or explain why it fails.
- Deduplication Agent: A cleaner that removes duplicates and merges aliases (since the same drug might have multiple names or codes, and different names across languages).
- Coach Agent: A “strategy planner” that looks at what the explorers found, notices gaps or errors, and suggests new angles to try next. The search grows like a tree with different branches representing new strategies, and the agent invests more effort into branches that look promising.
- Tree-based self-learning: Instead of just following one long search path, the system builds many branches, keeps track of all candidates and evidence, and focuses on the branches that produce the most true matches. Think of it like exploring a map: if one path leads to treasure, the agent explores similar paths more.
- Scoring and comparison:
- They measured performance using F1-score, which balances precision (how many found items were correct) and recall (how many correct items were found). F1 is the harmonic mean of these two, so higher is better.
- They compared Bioptic Agent against several leading AI research tools on this new benchmark.
Main Findings and Why They Matter
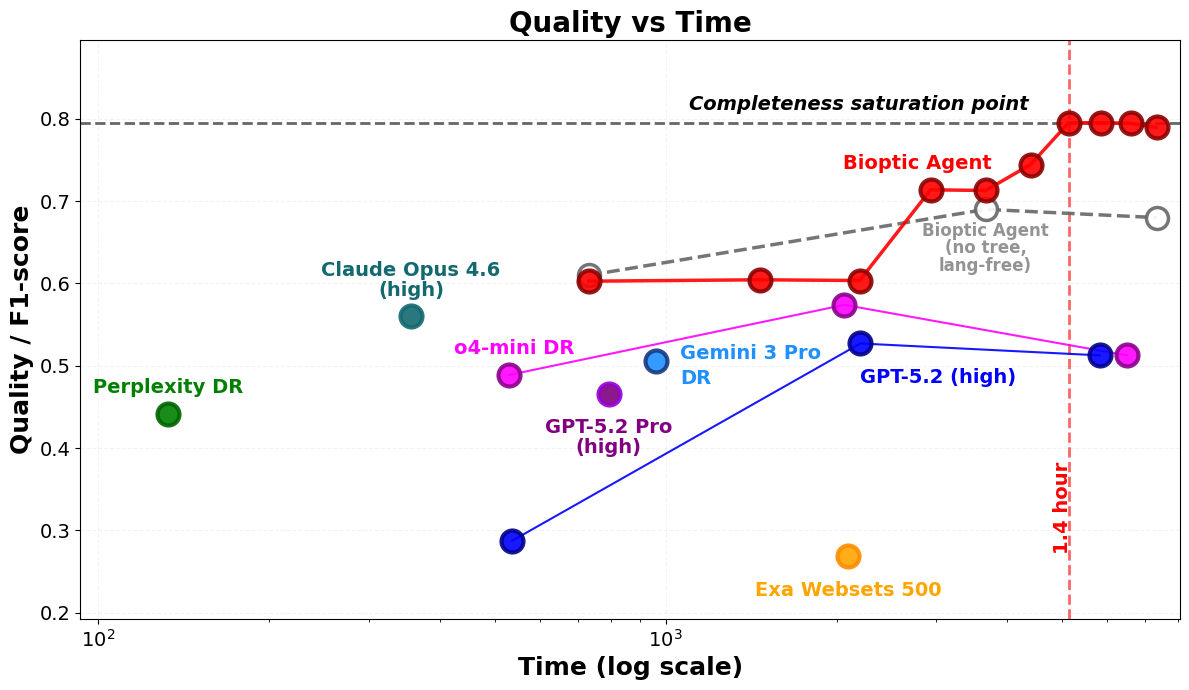
- The Bioptic Agent achieved a 79.7% F1-score on the benchmark.
- It beat several top systems:
- Claude Opus 4.6: 56.2%
- Gemini 3 Pro + Deep Research: 50.6%
- OpenAI GPT-5.2 Pro: 46.6%
- Perplexity Deep Research: 44.2%
- Exa Websets: 26.9%
- Performance improved sharply when the AI used more compute, supporting the idea that investing more computer power can help the agent find more assets and verify them better.
- The agent’s strengths come from:
- Searching across languages to catch non-English disclosures.
- Keeping a persistent record of candidates and evidence (not losing track of what’s already found).
- Using a specialized validator to check complex, multi-part queries like an expert would.
- Steering search via a “coach” that learns from mistakes and focuses on uncovered areas.
These results matter because missing a single valuable drug program can cost companies billions of dollars in lost deals. A system that finds more of the right assets earlier gives investors and business development teams a real edge.
What This Means Going Forward
- For investors, business development, and competitive intelligence teams: AI agents built for completeness and evidence-backed verification can dramatically improve global scouting, especially in regions where important information is published in local languages first.
- For the biopharma industry: Since so much innovation now happens outside the U.S., having AI that “hunts globally” reduces the risk of missing high-potential assets and helps teams move faster on partnerships, licensing, or acquisitions.
- For AI research: Wide, multilingual, “find-all” searches are different from typical web Q&A. This work suggests that:
- Benchmarks should test completeness and real-world complexity, not just short answers or nicely written summaries.
- Agents should keep detailed, traceable evidence and treat search as a branching, learning process, not just a single pass.
- More compute can be productively used to expand coverage and verify results.
In short, the paper shows a practical way to test and build AI that can scout the world’s drug landscape thoroughly and responsibly—helping decision-makers spot hidden opportunities that might otherwise be missed.
Knowledge Gaps
Knowledge gaps, limitations, and open questions
Below is a single, focused list of what remains missing, uncertain, or unexplored in the paper, framed as concrete, actionable directions for future research:
- Ground-truth completeness: The benchmark pairs each query with a single ground-truth asset; it does not require enumerating the full set of matching assets. Build multi-GT or fully enumerated ground truths to measure true recall/coverage in high-cardinality “find-all” tasks.
- Residual selection bias: Assets seeded from regional news may over-represent entities with media coverage or certain geographies/modalities. Quantify and mitigate bias via stratified sampling and inclusion of non-news sources (e.g., registries, patents, conference abstracts, local regulatory filings).
- Discoverability metric robustness: The English-vs-local “discoverability” filter relies on Google SERP page counts. Validate its stability across time, personalization, and search engines; compare against alternative metrics (e.g., domain diversity, citation graphs, language-specific index coverage).
- LLM-as-judge calibration: The validator’s “88% precision” is reported without full calibration details. Publish confusion matrices, recall, agreement with experts, and robustness to adversarial/ambiguous cases and non-English evidence; quantify error propagation to final metrics.
- Entity resolution accuracy: Deduplication and alias normalization (cross-lingual, code-names) lack quantitative evaluation. Measure false merges/splits and link assets to authoritative identifiers (INN, UNII, CAS, ChEMBL IDs, trial IDs) to improve reliability.
- Reward function and selection policy: The tree search’s node reward and UCB selection are referenced but not specified. Provide formal definitions, hyperparameters, and ablations to isolate their contribution to recall and precision.
- Compute-cost scaling: The claim that “performance improves steeply with additional compute” is not backed by detailed scaling curves. Quantify tokens, queries, wall-clock, and dollar/energy cost per F1 gain; identify diminishing returns and optimal budgets.
- Baseline comparability: Baseline configurations (versions, language capabilities, time/compute budgets, prompt constraints) are not detailed. Release scripts and settings to ensure fair, reproducible head-to-head comparisons, including non-English browsing enablement.
- Language coverage gaps: Evaluation emphasizes English and (at least) Chinese. Extend and report performance across Japanese, Korean, Portuguese, German, French, Spanish, Russian/Ukrainian, and other languages; analyze transliteration/script challenges and per-language failure modes.
- Small query prior: The seed corpus includes 48 investor/BD queries. Expand to a larger, more diverse corpus; test generalization to unseen intents, constraint compositions, and domains to reduce template-induced overfitting.
- Real-world impact: No prospective case studies showing improved asset capture or BD outcomes. Run live trials with BD teams to measure time-to-discovery, missed-opportunity reduction, and downstream deal value.
- Temporal robustness: Assets and disclosures evolve quickly. Introduce time-split evaluations, freshness metrics, and re-benchmarking over months to quantify drift, update cadence, and stability.
- Paywalled/grey literature: Coverage of paywalled local sources, academic proceedings, and grey literature is not assessed. Quantify recall gains from negotiated access and evaluate ethical/legal constraints.
- Complex constraint verification: Queries like “≤ N competitors ahead globally” require rigorous competitor counting. Define methodologies (stage normalization, target specificity, geography) and measure error rates on these derived constraints.
- Hallucination rates: Non-hallucination is claimed, but explicit hallucination metrics (unsupported claims, fabricated citations) are not reported. Add per-claim grounding audits and strict provenance checks.
- Evidence conflict resolution: How conflicting multi-source evidence is reconciled (recency, source credibility) is unspecified. Define aggregation policies, versioning, and uncertainty reporting.
- Benchmark release and licensing: It is unclear whether the benchmark (queries, GT assets, provenance, validator outputs) will be publicly released. Provide datasets, code, and licensing to enable independent replication and extension.
- Schema and ontology mapping: Asset attributes are extracted but not mapped to standard ontologies (MeSH, ICD-10/11, ATC, ChEMBL, DrugBank) for targets, indications, modalities. Implement cross-mapping and evaluate accuracy.
- Detailed error taxonomy: The paper lacks a systematic analysis of failure modes (language-specific, source-type gaps, alias mismatches, constraint parsing errors). Publish a taxonomy with prevalence to guide targeted improvements.
- Security, compliance, and ethics: Legal compliance for scraping regional sources, handling sensitive/regulatory documents, and respecting embargoes are not addressed. Establish policies and audit logs.
- Robust query handling: Performance on underspecified, noisy, or conflicting queries is not evaluated. Introduce tests for uncertainty handling, partial matches, and “insufficient evidence” responses.
- Model/version drift: Sensitivity to LLM provider updates is not measured. Create continuous benchmarking protocols and stability checks across model versions.
- Hybrid integrations: The paper critiques vendor databases but does not test hybrid pipelines. Quantify gains from integrating structured sources (Clarivate, GlobalData) with agentic mining.
- Human-in-the-loop control: Investigator tools for steering the directive tree, inspecting rationales, and correcting errors are not described. Evaluate the impact of expert oversight on recall and precision.
- Environmental and monetary footprint: Token/compute intensity implies non-trivial costs. Report energy and dollar costs per query and explore efficiency optimizations (caching, adaptive exploration, selective heavy deduplication).
Practical Applications
Immediate Applications
Below is a concise list of deployable, real-world uses that leverage the paper’s Bioptic Agent architecture, completeness-first benchmark, and multilingual mining pipeline.
- Healthcare/Pharma (Business Development and Corporate Strategy): Global asset scouting co-pilot
- What it does: Runs multilingual, breadth-first searches to surface “under-the-radar” drug programs that match investor-grade, multi-constraint queries; produces deduplicated, evidence-backed candidate sets with explicit provenance and validator rationales.
- Tools/products/workflows: Scouting Console; CRM/DealCloud integration (“Query-to-Dealflow” pipeline); Landscape Monitor (by indication/target/modality); watchlist refresh and alerting using English-vs-local discoverability signals.
- Assumptions/dependencies: Access to regional sources or SERP tools; language coverage beyond English; sufficient compute (quality scales with compute); LLM-as-judge calibration (≈88% precision); human-in-the-loop review for high-stakes decisions; compliance with data-use policies.
- Finance (VC/PE/Public Markets): Dealflow accelerator and white-space finder
- What it does: Converts screening theses into exhaustive, ranked asset lists including early-stage and regionally disclosed programs; maps competitive ceilings (e.g., “≤ N competitors ahead”) and modality/target constraints.
- Tools/products/workflows: Global Dealflow Feed with evidence packs; Investor Thesis Generator using query templates; risk screens for omission and aliasing; portfolio pipeline coverage audits.
- Assumptions/dependencies: Ongoing benchmark calibration to investor reality; dedup accuracy for code-names/transliterations; domain-specific validator prompts; integration with existing diligence workflows and compliance policies.
- Competitive Intelligence (R&D, Portfolio Strategy): Landscape monitor and rapid coverage audits
- What it does: Enumerates and tracks competitor programs across languages; detects new entrants and pipeline changes; surfaces stealth assets with low English discoverability.
- Tools/products/workflows: Competitive Landscape Monitor; “Under-amplified” asset alerts; constraint-aware dashboards (e.g., pathway, indication slice, modality).
- Assumptions/dependencies: Reliable alias resolution; refresh cadence aligned to fast-moving disclosures; access to regional registries, press, and corporate PDFs.
- Software/Data Providers (Drug Databases, Knowledge Graphs): Coverage gap augmentation
- What it does: Augments curation pipelines with multilingual asset mining, validator-grade evidence pairing, and cross-lingual dedup to reduce blind spots and lag.
- Tools/products/workflows: Coverage Gap Filler API; deduplication services (light/heavy modes) with canonicalization; provenance-rich schema outputs for ingestion.
- Assumptions/dependencies: Licensing for data ingestion; harmonization with internal ontologies (targets, modalities, indications); throughput/cost controls given steep compute-response tradeoffs.
- Academia (Biomedical Informatics, Innovation Studies): Benchmarking and landscape research
- What it does: Uses the completeness benchmark to evaluate “enumerate-all” capabilities of web agents; produces reproducible, provenance-backed landscape maps for indication/target classes and geographic origins.
- Tools/products/workflows: Benchmark usage for agent evaluation; open query templates reflecting real investor intents; white-space analyses and attrition studies.
- Assumptions/dependencies: Access to benchmark assets/query pairs; compute budgets; awareness of residual selection bias (news-coverage skew) and mitigation steps described.
- Policy/Regulators (Public Health, Industrial Policy): Innovation surveillance and early signal detection
- What it does: Monitors domestic and regional pipelines, especially low-visibility programs; tracks trial activity and licensing flows; uses discoverability profiles (English vs local) to identify information asymmetries.
- Tools/products/workflows: Regulatory Radar; Innovation Heatmap by modality/indication/geography; grant/procurement triage informed by evidence density.
- Assumptions/dependencies: Data-sharing agreements; multilingual capability in monitored regions; careful interpretation to avoid over-reliance on public-comms signals.
- Daily Life (Patient Advocacy, Health Journalism, Clinical Community): Indication-focused navigator for upcoming therapies
- What it does: Produces curated lists of preclinical/clinical assets per disease with citations; highlights trial geographies and stages.
- Tools/products/workflows: Indication Navigator with guardrails; fact-checked summaries; media-ready evidence bundles for reporting.
- Assumptions/dependencies: Non-diagnostic use; clinician/expert review before patient guidance; frequent updates to reflect rapidly changing pipelines.
- Legal/IP and BD Due Diligence: Evidence packs with transparent provenance
- What it does: Generates audit-ready documentation for licensing/partnerships, including validated claims and source quotes.
- Tools/products/workflows: “Evidence Pack” generator; modality/target/geography compliance checks; alias normalization for contract clarity.
- Assumptions/dependencies: Recency of sources; cautious handling of early-stage disclosures and transliterations; human counsel sign-off.
Long-Term Applications
These opportunities likely require further research, scaling, and ecosystem development before reliable deployment.
- Healthcare/Software: Near-real-time global drug pipeline graph
- What it could be: A continuously updated, cross-lingual knowledge graph unifying trials, patents, press, regulatory filings, and corporate PDFs; heavy-mode dedup for canonicalization.
- Potential tools/products/workflows: Open APIs; enterprise-grade data feeds; full lineage of evidence for compliance-grade consumption.
- Assumptions/dependencies: Data licensing; governance for provenance trust; robust alias and code-name evolution tracking; sustained compute.
- Finance/Partnerships: Licensing and partnering marketplace
- What it could be: Matching engine connecting scouts and asset owners across geographies using constraint-aware queries and completeness-first discovery.
- Potential tools/products/workflows: Outreach prioritization co-pilot; partner fit scoring; cross-border compliance workflows.
- Assumptions/dependencies: Legal frameworks for deal-making; standardized schemas; trust and verification mechanisms beyond LLM judgment.
- Clinical Practice: EHR-integrated trial matching and therapy radar
- What it could be: Precision matching of patients to emerging trials and therapies, updated by global scouting signals.
- Potential tools/products/workflows: Decision support embedded in EHRs; clinician-facing evidence views; automated eligibility parsing.
- Assumptions/dependencies: HIPAA/PHI compliance; clinical validation; institutional liability and workflow integration; regulator approvals.
- Government/Policy: Industrial policy analytics and funding calibration
- What it could be: Dynamic measurement of innovation flows (by origin, stage, modality) to inform grants, translational funding, and trade policy.
- Potential tools/products/workflows: National innovation dashboards; regional pipeline comparators; impact modeling.
- Assumptions/dependencies: Method transparency; bias mitigation (e.g., media/registry coverage skew); data-sharing agreements.
- Education/Workforce Development: Analyst training and certification
- What it could be: Curricula built around query templates, validator rationale interpretation, and tree-based exploration to train BD/CI analysts.
- Potential tools/products/workflows: Capstone projects using the benchmark; certification tracks; simulated deal screens.
- Assumptions/dependencies: Industry acceptance; standardized competency rubrics; maintained benchmark updates.
- Cross-Sector Generalization (Software, Robotics, Energy, Materials): Wide-search enumerative discovery
- What it could be: Adapt Bioptic-style agents for “find-all” tasks in other sectors (e.g., energy storage breakthroughs, robotics components, materials patents, edtech programs).
- Potential tools/products/workflows: Sector-specific validators and ontologies; multilingual mining across trade registries and technical literature.
- Assumptions/dependencies: Domain-specific schema design; high-precision validators; sector-relevant source coverage.
- Compliance and Risk (Finance, Supply Chain, ESG): Automated enumerate-all scanning
- What it could be: Exhaustive identification of sanctioned entities, supply-chain exposures, or ESG risks from heterogeneous disclosures.
- Potential tools/products/workflows: Compliance dashboards; audit trails with citations; periodic risk refreshes.
- Assumptions/dependencies: Precision-first validator tuning; strict access controls and logging; regulatory audit acceptance.
- Agent Orchestration Platforms (Software): Coach-directed multi-agent frameworks
- What it could be: Generalized tree-based orchestration (selection, rollout, evaluate, backpropagate, expand) for tasks where recall and provenance are paramount.
- Potential tools/products/workflows: Standardized reward models; directive libraries; language-parallel investigator pools.
- Assumptions/dependencies: Benchmarking standards for recall/completeness; compute budgets; reproducibility guarantees.
- Scientific Discovery (Academia/Pharma): Early signal synthesis for novel mechanisms
- What it could be: Rapid detection of emerging MoAs (e.g., PAPD5/7–ZCCHC14 axis, RNA-targeting strategies) across gray literature and regional disclosures to guide lab programs.
- Potential tools/products/workflows: Hypothesis generators with evidence lineage; cross-lab collaboration prompts.
- Assumptions/dependencies: Expert curation; experimental validation; careful separation of signal vs. noise in early reports.
Cross-cutting assumptions and dependencies
- Compute scaling: The paper shows steep quality gains with more compute; budgeting and latency tradeoffs will shape feasibility.
- Multilingual coverage: Performance depends on language parallelism; expanding to additional languages and regional sources improves recall.
- Validator reliability: LLM-as-judge must be calibrated to expert opinions and task constraints; periodic re-tuning is necessary.
- Data access: SERP tools, registries, paywalled sources, and corporate PDFs may require licenses; rate limiting and robots rules must be respected.
- Provenance and audit: High-stakes use requires transparent evidence lineage and human oversight to mitigate hallucinations and alias errors.
- Bias and representativeness: Regional news-driven seeds can bias coverage; mitigation steps (non-English mining, discoverability filters) reduce but do not eliminate skew.
- Legal and compliance: IP, privacy, and regulatory constraints vary by jurisdiction; enterprise deployment needs governance, logging, and policy adherence.
Glossary
- AAV: Adeno-associated virus; a viral vector commonly used to deliver genetic material in gene therapy. "Vectorized RNAi assets (ddRNAi/shRNA, e.g., AAV-based) surfaced via national registries and local pipeline disclosures."
- alias resolution: The process of identifying and reconciling multiple names or identifiers that refer to the same entity. "LLM-as-judge components for alias resolution and up-to-date attribute extraction."
- Antisense oligonucleotide (ASO): Short, synthetic strands of nucleic acids that bind RNA to modulate gene expression. "Antisense oligonucleotide assets (RNase-H/gapmer/LNA) surfaced via national registries and local pipeline disclosures."
- asset scouting: Systematic search and identification of drug programs (assets) that match investment or BD criteria. "applying AI to drug asset scouting"
- biomarker: A measurable indicator of a biological state or condition used to assess disease or treatment response. "biomarker(s)"
- Business Development (BD): Corporate function focused on partnerships, licensing, and acquisitions to expand a pipeline. "BD/VC-style multi-constraint screening queries"
- Criteria Match Validator Agent: An AI component that checks whether candidate assets meet all query criteria, with evidence-backed rationales. "Criteria Match Validator Agent checks each candidate asset against the query criteria and outputs a match verdict plus a detailed, traceable, supported by evidence pass/fail rationale."
- ddRNAi: DNA-directed RNA interference; a vectorized approach to produce RNAi molecules inside cells. "Vectorized RNAi assets (ddRNAi/shRNA, e.g., AAV-based) surfaced via national registries and local pipeline disclosures."
- Deep Research: Agentic web-retrieval and synthesis systems optimized for multi-step, citation-backed investigations. "Deep Research Agents are tasked to use only language \ell for web queries to find news anonuncements according drugs and biotech companies written in \ell and in source s"
- Deduplication Agent: An AI component that identifies and removes duplicate assets and resolves aliases to maintain a unique set. "Deduplication Agent ensures a global set of validated assets contains only unique assets by removing duplicates and resolving aliases."
- discoverability: How easily information about an asset can be found via search; often measured across languages. "English-vs-local discoverability signal"
- F1-score: The harmonic mean of precision and recall, measuring accuracy of set retrieval. "F1-score (harmonic mean of precision and recall; higher is better)."
- GalNAc: N-acetylgalactosamine; a sugar used to target siRNA to the liver via conjugation. "GalNAc-delivered siRNA assets in Japan and Taiwan surfaced via national registries and local pipeline disclosures."
- ground truth (GT): The verified correct answer(s) used for evaluation. "ground-truth (GT) assets"
- HBV: Hepatitis B virus; a pathogen targeted by various RNA-based and small-molecule therapies. "HBV oligonucleotide programs (siRNA/ASO/ddRNAi/shRNA) in undercover APAC markets surfaced via national registries and local pipeline disclosures"
- HBsAg: Hepatitis B surface antigen; a key biomarker for HBV infection and treatment response. "HBsAg or HBV RNA reduction."
- IND: Investigational New Drug; a regulatory filing to start clinical trials in humans. "entity-agnostic templates (e.g., new clinical trial authorization, licensing agreement, IND filed, government grant, phase I initiated)"
- line of therapy: Treatment order or setting in a disease (e.g., first-line, second-line). "line of therapy"
- LLM-as-judge: Using a LLM to evaluate or grade outputs against criteria, calibrated to expert judgments. "For grading, we use LLM-as-judge evaluation calibrated to expert opinions."
- LNA: Locked nucleic acid; a chemical modification in oligonucleotides to increase stability and affinity. "RNase-H gapmer/LNA ASO HBV assets surfaced via registries and local pipeline disclosures."
- LNP: Lipid nanoparticle; a delivery system for nucleic acid therapeutics like siRNA/mRNA. "LNP-delivered siRNA assets in Japan, Taiwan, South Korea, Singapore, Hong Kong, and Australia/New Zealand surfaced via national registries and local pipeline disclosures."
- mechanism of action (MoA): The specific biochemical interaction through which a drug produces its effect. "rewrite constraints (like MoA/target class, modality family, indication with population/line-of-therapy slice, maturity window, geography/origin, ownership/licensing, and evidence signals)"
- modality: The therapeutic form or technology class of a drug (e.g., small molecule, biologic, siRNA). "Program descriptors: developer(s), modality, target(s), short mechanism of action, detailed mechanism of action, indication(s), and patent(s)"
- molecular glues: Small molecules that induce interactions between proteins to degrade or modulate targets. "ZCCHC14 targeted degraders or molecular glues (non-PAPD5/7 enzymatic) with HBV RNA/antigen reduction evidence."
- multi-hop reasoning: Integrating evidence across multiple sources and steps to satisfy complex constraints. "requires evidence aggregation and multi-hop reasoning rather than lexical matching."
- PAPD5/7: Enzymes involved in HBV RNA processing; therapeutic targets for small-molecule inhibition. "PAPD5/7 (TENT4A/B) enzymatic inhibitors with HBsAg or HBV RNA reduction evidence."
- Precision oncology: Cancer treatment tailored to specific molecular features (e.g., mutations, biomarkers). "Precision oncology sub-landscapes"
- provenance: Documented source information supporting each claim to ensure traceability. "every atomic claim is paired with explicit provenance (a list of source URL and verbatim supporting quote pairs)."
- readouts: Upcoming or reported results from trials or studies that serve as catalysts. "Catalysts and upcoming readouts"
- regimen: The specific dosing schedule and combination of therapies used in a trial. "regimen"
- RNase-H gapmer: An ASO design that recruits RNase H to degrade target RNA via a chimeric structure. "RNase-H gapmer/LNA ASO HBV assets surfaced via registries and local pipeline disclosures."
- Search Engine Results Page (SERP): The page displayed by a search engine in response to a query, often quantified for coverage. "Using a SERP tool, we count the maximum number of google search result pages for (A) English queries..."
- shRNA: Short hairpin RNA; RNA molecules expressed in cells to induce RNA interference. "Vectorized RNAi assets (ddRNAi/shRNA, e.g., AAV-based) surfaced via national registries and local pipeline disclosures."
- siRNA: Small interfering RNA; short double-stranded RNA that silences gene expression via RNA interference. "SiRNA assets (GalNAc or LNP) surfaced via national registries and local pipeline disclosures."
- TENT4A/B: Alternative names for PAPD5/7 enzymes implicated in HBV RNA stabilization. "PAPD5/7 (TENT4A/B) enzymatic inhibitors with HBsAg or HBV RNA reduction evidence."
- transliteration: Converting text from one writing system to another, often causing multiple asset name variants. "aliases through code-name changes, transliterations, and subsidiary disclosures"
- trial registry: Official databases of clinical trials used for regulatory and transparency purposes. "statutory trial registries"
- Upper Confidence Bound (UCB): An exploration-exploitation selection strategy in bandit/tree search algorithms. "Select m nodes {n_i}_{i=1}{m} via Upper Confidence Bound (UCB) rule"
- vectorized RNAi: Delivery of RNAi constructs via vectors (e.g., viral) to generate interference molecules in vivo. "Vectorized RNAi assets (ddRNAi/shRNA, e.g., AAV-based) surfaced via national registries and local pipeline disclosures."
- White-space: Areas of low competition or unmet need used to identify promising, less crowded targets. "White-space and low-competition target hunting"
- ZCCHC14: A protein involved in HBV RNA machinery; target for novel inhibitors or complex disruptors. "ZCCHC14-engagers or PAPD5/7--ZCCHC14 complex disruptors, excluding PAPD5/7 enzymatic inhibitors, with HBsAg or HBV RNA reduction evidence."
Collections
Sign up for free to add this paper to one or more collections.